Abstract
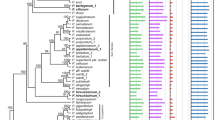
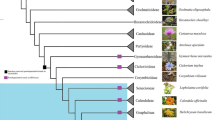
The complete sequence of the chloroplast genome of cassava (Manihot esculenta, Euphorbiaceae) has been determined. The genome is 161,453 bp in length and includes a pair of inverted repeats (IR) of 26,954 bp. The genome includes 128 genes; 96 are single copy and 16 are duplicated in the IR. There are four rRNA genes and 30 distinct tRNAs, seven of which are duplicated in the IR. The infA gene is absent; expansion of IRb has duplicated 62 amino acids at the 3′ end of rps19 and a number of coding regions have large insertions or deletions, including insertions within the 23S rRNA gene. There are 17 intron-containing genes in cassava, 15 of which have a single intron while two (clpP, ycf3) have two introns. The usually conserved atpF group II intron is absent and this is the first report of its loss from land plant chloroplast genomes. The phylogenetic distribution of the atpF intron loss was determined by a PCR survey of 251 taxa representing 34 families of Malpighiales and 16 taxa from closely related rosids. The atpF intron is not only missing in cassava but also from closely related Euphorbiaceae and other Malpighiales, suggesting that there have been at least seven independent losses. In cassava and all other sequenced Malphigiales, atpF gene sequences showed a strong association between C-to-T substitutions at nucleotide position 92 and the loss of the intron, suggesting that recombination between an edited mRNA and the atpF gene may be a possible mechanism for the intron loss.




Similar content being viewed by others
References
APG (2003) An update of the Angiosperm Phylogeny Group classification for the orders and families of flowering plants: APG II. Bot J Linn Soc 141:399–436
Asakura Y, Barkan A (2006) Arabidopsis orthologs of maize chloroplast splicing factors promote splicing of orthologous and species-specific group II introns. Plant Physiol 142:1656–1663
Barkan A (2004) Intron splicing in plant organelles. In: Daniell H, Chase CD (eds) Molecular biology and biotechnology of plant organelles: chloroplast and mitochondria. Springer, Netherlands, pp 295–322
Bausher MG, Singh ND, Lee SB, Jansen RK, Daniell H (2006) The complete chloroplast genome sequence of Citrus sinensis (L.) Osbeck var ‘Ridge Pineapple’: organization and phylogenetic relationships to other angiosperms. BMC Plant Biol 6:21
Berry PE, Hipp AL, Wurdack KJ, van Ee B, Riina R (2005) Molecular phylogenetics of the giant genus Croton and tribe Crotoneae (Euphorbiaceae sensu stricto) using ITS and trnL-trnF DNA sequence data. Am J Bot 92:1520–1534
Bonen L, Vogel J (2001) The ins and outs of group II introns. Trends Genet 17:322–331
Bowe LM, dePamphilis CW (1996) Effects of RNA editing and gene processing on phylogenetic reconstruction. Mol Biol Evol 13:1159–1166
Corneille S, Lutz K, Maliga P (2000) Conservation of RNA editing between rice and maize plastids: are most editing events dispensable? Mol Gen Genet 264:419–424
Daniell H (2002) Molecular strategies for gene containment in transgenic crops. Nat Biotechnol 20:581–586
Daniell H (2006) Production of biopharmaceuticals and vaccines in plants via the chloroplast genome. Biotechnol J 1:1071–1079
Daniell H (2007) Transgene containment by maternal inheritance: effective or elusive? Proc Natl Acad Sci USA 104:6879–6880
Daniell H, Datta R, Varma S, Gray S, Lee S-B (1998) Containment of herbicide resistance through genetic engineering of the chloroplast genome. Nat Biotechnol 16:345–348
Daniell H, Kumar S, Dufourmantel N (2005) Breakthrough in chloroplast genetic engineering of agronomically important crops. Trends Biotechnol 5:238–245
Daniell H, Lee S-B, Grevich J, Saski C, Quesada-Vargas T, Guda C, Tomkins J, Jansen RK (2006) Complete chloroplast genome sequences of Solanum bulbocastanum, Solanum lycopersicum and comparative analyses with other Solanaceae genomes. Theor Appl Genet 112:1503–1518
Davis CC, Chase MW (2004) Elatinaceae are sister to Malpighiaceae; Peridiscaceae belong to Saxifragales. Am J Bot 91:262–273
Davis CC, Webb CO, Wurdack KJ, Jaramillo CA, Donoghue MJ (2005) Explosive radiation of Malpighiales supports a mid-cretaceous origin of modern tropical rain forests. Am Nat 165:E36–E65
Davis CC, Wurdack KJ (2004) Host-to-parasite gene transfer in flowering plants: phylogenetic evidence from Malpighiales. Science 305:676–678
DeCosa B, Moar W, Lee S-B, Miller M, Daniell H (2001) Overexpression of the Bt cry2Aa2 operon in chloroplasts leads to formation of insecticidal crystals. Nat Biotechnol 9:71–74
DeGray G, Rajasekaran K, Smith F, Sanford J, Daniell H (2001) Expression of an antimicrobial peptide via the chloroplast genome to control phytopathogenic bacteria and fungi. Plant Physiol 127:852–862
Deshpande NN, Hollingsworth M, Herrin DL (1995) The atpF group-II intron-containing gene from spinach chloroplasts is not spliced in transgenic Chlamydomonas chloroplasts. Curr Genet 28:122–127
Dickau R, Ranere AJ, Cooke RG (2007) Starch grain evidence for the preceramic dispersals of maize and root crops into tropical dry and humid forests of Panama. Proc Natl Acad Sci USA 104:3651–3656
Downie SR, Katz-Downie DS, Wolfe KH, Calie PJ, Palmer JD (1994a) Structure and evolution of the largest chloroplast gene (ORF2280): internal plasticity and multiple gene loss during angiosperm evolution. Curr Genet 25:367–378
Downie SR, Llanas E, Katz-Downie DS (1994b) Multiple independent losses of the rpoC1 intron in angiosperm chloroplast DNAs. Syst Bot 21:135–151
Downie SR, Olmstead RG, Zurawski G, Soltis DE, Soltis PS, Watson JC, Palmer JD (1991) Six independent losses of the chloroplast DNA rpl2 intron in dicotyledons: molecular and phylogenetic implications. Evolution 45:1245–1259
Downie SR, Palmer JD (1992) Use of chloroplast DNA rearrangements in reconstructing plant phylogeny. In: Soltis PS, Soltis DE, Doyle JJ (eds) Molecular systematics of plants. Chapman and Hall, New York, pp 14–35
Dufourmantel N, Dubald M, Matringe M, Canard H, Garcon F, Job C, Kay E, Wisniewski JP, Ferullo JM, Pelissier B, Sailland A, Tissot G (2007) Generation and characterization of soybean and marker-free tobacco plastid transformants over-expressing a bacterial 4-hydroxyphenylpyruvate dioxygenase which provides strong herbicide tolerance. Plant Biotechnol J 5:118–133
El-Sharkawy MA (2004) Cassava biology and physiology. Plant Mol Biol 56:481–501
Ewing B, Green P (1998) Base-calling of automated sequencer traces using phred. II. Error probabilities. Gen Res 8:186–194
Fregene M (2000) Marking progress: collaboration to improve cassava. In: Kinley D (ed) Synergies in science. Consultative Group on International Agricultural Research, Washington, pp 6–7
Fregene M, Angel F, Gómez R, Rodríguez F, Chavarriaga P, Roca W, Tohme J, Bonierbale M (1997) A molecular genetic map of cassava (Manihot esculenta Crantz). Theor Appl Genet 95:431–441
Grevich JJ, Daniell H (2005) Chloroplast genetic engineering: recent advances and future perspectives. Crit Rev Plant Sci 24:83–108
Hagemann R (2004) The Sexual inheritance of plant organelles. In: Daniell H, Chase C (eds) Molecular biology and biotechnology of plant organelles: chloroplasts and mitochondria. Springer, Netherlands, pp 93–113
Hansen AK, Gilbert LE, Simpson BB, Downie SR, Cervi AC, Jansen RK (2006) Phylogenetic relationships and chromosome number evolution in Passiflora. Syst Bot 31:138–150
Hillocks RJ (2002) Cassava in Africa. In: Hillocks RJ, Thresh JM, Bellotti AC (eds) Cassava: biology, production and utilization. CABI Publishing, New York, pp 41–54
Hoffmann P, Kathriarachchi H, Wurdack KJ (2006) A phylogenetic classification of Phyllanthaceae (Malpighiales; Euphorbiaceae sensu lato). Kew Bull 61:37–53
Huelsenbeck JP, Ronquist FR (2001) MrBayes: Bayesian inference of phylogenetic trees. Bioinformatics 17:754–755 [MrBayes available at http://mrbayes.scs.fsu.edu/]
Itchoda N, Nishizawa S, Nagano H, Kubo T, Mikami T (2002) The sugar beet mitochondrial nad4 gene: an intron loss and its phylogenetic implication in the Caryophyllales. Theor Appl Genet 104:209–213
Jansen RK, Cai Z, Raubeson LA, Daniell H, dePamphilis CW, Leebens-Mack J, Müller KF, Guisinger-Bellian M, Haberle RC, Hansen AK, Chumley TW, Lee S-B, Peery R, McNeal J, Kuehl JV, Boore JL (2007) Analysis of 81 genes from 64 plastid genomes resolves relationships in angiosperms and identifies genome-scale evolutionary patterns. Proc Natl Acad Sci USA 104:19369–19374
Jenkins BD, Kulhanek DJ, Barkan A (1997) Nuclear mutations that block group II RNA splicing in maize chloroplasts reveal several intron classes with distinct requirements for splicing factors. Plant Cell 9:283–296
Kahlau S, Aspinall S, Gray JC, Bock R (2006) Sequence of the tomato chloroplast DNA and evolutionary comparison of solanaceous plastid genomes. J Mol Evol 63:194–207
Kanamoto H, Yamashita A, Asao H, Okumura S, Takase H, Hattori M, Yokota A, Tomizawa K (2006) Efficient and stable transformation of Lactuca sativa L. cv. Cisco (lettuce) plastids. Transgenic Res 15:205–217
Kathriarachchi H, Hoffmann P, Samuel R, Wurdack KJ, Chase MW (2005) Molecular phylogenetics of Phyllanthaceae inferred from five genes (plastid atpB, matK, 3′ ndhF, and nuclear PHYC). Mol Phylog Evol 36:112–134
Knauf U, Hachtel W (2002) The genes encoding subunits of ATP synthase are conserved in the reduced plastid genome of the heterotrophic alga Prototheca wickerhamii. Mol Genet Genomics 267:492–497
Kota M, Daniell H, Varma S, Garczynski SF, Gould F, Moar WJ (1999) Overexpression of the Bacillus thuringiensis (Bt) Cry2Aa2 protein in chloroplasts confers resistance to plants against susceptible and Bt-resistant insects. Proc Natl Acad Sci USA 96:1840–1845
Kumar S, Dhingra A, Daniell H (2004a) Plastid expressed betaine aldehyde dehydrogenase gene in carrot cultured cells, roots, and leaves confer enhanced salt tolerance. Plant Physiol 136:2843–2854
Kumar S, Dhingra A, Daniell H (2004b) Stable transformation of the cotton plastid genome and maternal inheritance of transgenes. Plant Mol Biol 56:203–216
Lee SB, Kaittanis C, Jansen RK, Hostetler JB, Tallon LJ, Town CD, Daniell H (2006) The complete chloroplast genome sequence of Gossypium hirsutum: organization and phylogenetic relationships to other angiosperms. BMC Genomics 7:61
Lee S-B, Kwon HB, Kwon SJ, Park SC, Jeong MJ, Han SE, Byun MO, Daniell H (2003) Accumulation of trehalose within transgenic chloroplasts confers drought tolerance. Mol Breed 11:1–13
Maliga P (2004) Plastid transformation in higher plants. Annu Rev Plant Biol 55:289–313
Mba REC, Stephenson P, Edwards K, Melzer S, Nkumbira J, Gullberg U, Apel K, Gale M, Tohme J, Fregene M (2001) Simple sequence repeat (SSR) markers survey of the cassava (Manihot esculenta Crantz) genome: towards a SSR-based molecular genetic map of cassava. Theor Appl Genet 102:21–31
McPherson MA, Fay MF, Chase MW, Graham SW (2004) Parallel loss of a slowly evolving intron from two closely related families in Asparagales. Syst Bot 29:296–307
Millen RS, Olmstead RG, Adams KL, Palmer JD, Lao NT, Heggie L, Kavanagh TA, Hibberd JM, Gray JC, Morden CW, Calie PJ, Jermiin LS, Wolfe KH (2001) Many parallel losses of infA from chloroplast DNA during angiosperm evolution with multiple independent transfers to the nucleus. Plant Cell 13:645–658
Miyagi T, Kapoor S, Sugita M, Sugiura M (1998) Transcript analysis of the tobacco plastid operon rps2/atpI/H/F/A reveals the existence of a non-consensus type II (NCII) promoter upstream of the atpI coding sequence. Mol Gen Genet 257:299–307
Okezie BO, Kosikowski FV (1982) Cassava as a food. Crit Rev Food Sci Nutr 17:259–275
Okumura S, Sawada M, Park YW, Hayashi T, Shimamura M, Takase H, Tomizawa K (2006) Transformation of poplar (Populus alba) plastids and expression of foreign proteins in tree chloroplasts. Transgenic Res 15:637–646
Olsen KM, Schaal BA (1999) Evidence on the origin of cassava: phylogeography of Manihot esculenta. Proc Natl Acad Sci USA 96:5586–5591
Olsen KM, Schaal BA (2001) Microsatellite variation in cassava (Manihot esculenta, Euphorbiaceae) and its wild relatives: further evidence for a southern Amazonian origin of domestication. Am J Bot 88:131–142
Ostersetzer O, Cooke AM, Watkins KP, Barkan A (2005) CRS1, a chloroplast group II intron splicing factor, promotes intron folding through specific interactions with two intron domains. Plant Cell 17:241–255
Pagel M, Meade A (2006) Bayesian analysis of correlated evolution of discrete characters by reversible-jump markov chain Monte Carlo. Am Nat 167:808–825 [BayesTraits available at http://www.evolution.reading.ac.uk/BayesTraits.html]
Plader W, Yukawa Y, Sugiura M, Malepszy S (2007) The complete structure of the cucumber (Cucumis sativus L.) chloroplast genome: its composition and comparative analysis. Cell Mol Biol Lett 12:584–594
Posada D, Crandall KA (1998) MODELTEST: testing the model of DNA substitution. Bioinformatics 14:817–818 [MODELTEST available at http://darwin.uvigo.es/software/modeltest.html]
Quesada-Vargas T, Ruiz ON, Daniell H (2005) Characterization of heterologous multigene operons in transgenic chloroplasts: transcription, processing, and translation. Plant Physiol 138:1746–1762
Raubeson LA, Peery R, Chumley TW, Dziubek C, Fourcade HM, Boore JL, Jansen RK (2007) Comparative chloroplast genomics: analyses including new sequences from the angiosperms Nuphar advena and Ranunculus macranthus. BMC Genomics 8:174
Raven P, Fauquet C, Swaminathan MS, Borlaug N, Samper C (2006) Where next for genome sequencing? Science 311:468
Ruf S, Hermann M, Berger I, Carrer H, Bock R (2001) Stable genetic transformation of tomato plastids and expression of a foreign protein in fruit. Nat Biotechnol 19:870–875
Ruhlman T, Lee SB, Jansen RK, Hostetler JB, Tallon LJ, Town CD, Daniell H (2006) Complete plastid genome sequence of Daucus carota: implications for biotechnology and phylogeny of angiosperms. BMC Genomics 7:222
Ruhlman T, Ahangari R, Devine A, Samsam M, Daniell H (2007) Expression of cholera toxin B-proinsulin fusion protein in lettuce and tobacco chloroplasts—oral administration protects against development of insulitis in non-obese diabetic mice. Plant Biotech J 5: 495–510
Ruiz ON, Daniell H (2005) Engineering the cytoplasmic male sterility via the chloroplast genome. Plant Physiol 138:1232–1246
Samson N, Bausher MG, Lee S-B, Jansen RK, Daniell H (2007) The complete nucleotide sequence of the coffee (Coffea arabica L.) chloroplast genome: organization and implications for biotechnology and phylogenetic relationships among angiosperms. Plant Biotech J 5:339–353
Saski C, Lee S-B, Daniell H, Wood TC, Tomkins J, Kim HG, Jansen RK (2005) Complete chloroplast genome sequence of Glycine max and comparative analyses with other legume genomes. Plant Mol Biol 59:309–322
Sato S, Nakamura Y, Kaneko T, Asamizu E, Tabata S (1999) Complete structure of the chloroplast genome of Arabidopsis thaliana. DNA Res 6:283–290
Schmitz-Linneweber C, Maier RM, Alcaraz JP, Cottet A, Herrmann RG, Mache R (2001) The plastid chromosome of spinach (Spinacia oleracea) complete nucleotide sequence and gene organization. Plant Mol Biol 45:307–315
Schmitz-Linneweber C, Regel R, Du TG, Hupfer H, Herrmann RG, Maier RM (2002) The plastid chromosome of Atropa belladonna and its comparison with that of Nicotiana tabacum: the role of RNA editing in generating divergence in the process of plant speciation. Mol Biol Evol 19:1602–1612
Shinozaki K, Ohme M, Tanaka M, Wakasugi T, Hayashida N, Matsubayashi T, Zaita N, Chunwongse J, Obokata J, Yamaguchi-Shinozaki K, Ohto C, Torazawa K, Meng BY, Sugita M, Deno H, Kamogashira T, Yamada K, Kusuda J, Takaiwa F, Kato A, Tohdoh N, Shimada H, Sugiura M (1986) The complete nucleotide sequence of the tobacco chloroplast genome: its gene organization and expression. EMBO J 5:2043–2049
Sidorov VA, Kasten D, Pang SZ, Hajdukiewicz PT, Staub JM, Nehra NS (1999) Stable chloroplast transformation in potato: use of green fluorescent protein as a plastid marker. Plant J 19:209–216
Sikdar SR, Serino G, Chaudhuri S, Maliga P (1998) Plastid transformation in Arabidopsis thaliana. Plant Cell Rep 18:245–247
Sikes DS, Lewis PO (2001) PAUPRat: PAUP* implementation of the parsimony ratchet. [PAUPRat available at http://www.ucalgary.ca/∼dsikes/software2.htm]
Simon D, Fewer D, Friedl T, Bhattacharya D (2003) Phylogeny and self-splicing ability of the plastid tRNA-Leu group I intron. J Mol Evol 57:710–720
Stahl DJ, Rodermel SR, Bogorad L, Subramanian AR (1993) Co-transcription pattern of an introgressed operon in the maize chloroplast genome comprising four ATP synthase subunit genes and the ribosomal rps2. Plant Mol Biol 21:1069–1076
Swofford DL (2003) PAUP*. Phylogenetic analysis using parsimony (*and other methods), vers. 4. Sinauer Associates, Sunderland, MA
Tillich M, Lehwark P, Morton BR, Maier UG (2006) The evolution of chloroplast RNA editing. Mol Biol Evol 23:1912–1921
Tokuoka T (2007) Molecular phylogenetic analysis of Euphorbiaceae sensu stricto based on plastid and nuclear DNA sequences and ovule and seed character evolution. J Plant Res 120:511–522
Turmel M, Lemieux C, Burger G, Lang BF, Otis C, Plante I, Gray MW (1999) The complete mitochondrial DNA sequences of Nephroselmis olivacea and Pedinomonas minor: two radically different evolutionary patterns within green algae. Plant Cell 11:1717–1730
Turmel M, Otis C, Lemieux C (2005) The complete chloroplast DNA sequences of the charophycean green algae Staurastrum and Zygnema reveal that the chloroplast genome underwent extensive changes during the evolution of the Zygnematales. BMC Biol 3:22
Turmel M, Otis C, Lemieux C (2006) The chloroplast genome sequence of Chara vulgaris sheds new light into the closest green algal relatives of land plants. Mol Biol Evol 23:1324–1338
Ugent D, Pozorski S, Pozorski T (1986) Archaeological manioc (Manihot) from coastal Peru. Econ Bot 40:78–102
Vogel J, Börner T (2002) Lariat formation and a hydrolytic pathway in plant chloroplast group II intron splicing. EMBO J 21:3794–3803
Vogel J, Börner T, Hess WR (1999) Comparative analysis of splicing of the complete set of chloroplast group II introns in three higher plant mutants. Nucleic Acids Res 27:3866–3874
Wurdack KJ, Hoffmann P, Chase MW (2005) Molecular phylogenetic analysis of uniovulate Euphorbiaceae (Euphorbiaceae sensu stricto) using plastid rbcL and trnLF sequences. Am J Bot 92:1397–1420
Wurdack KJ, Hoffmann P, Samuel R, De Bruijn A, Van der Bank M, Chase MW (2004) Molecular phylogenetic analysis of Phyllanthaceae (Phyllanthoideae pro parte, Euphorbiaceae sensu lato) using plastid rbcL DNA sequences. Amer J Bot 91:1882–1900
Wyman SK, Jansen RK, Boore JL (2004) Automatic annotation of organellar genomes with DOGMA. Bioinformatics 20:3252–3255 [DOGMA available at http://dogma.ccbb.utexas.edu/]
Xu MQ, Kaathe S, Goodrich BH, Nierwicki BS, Shub D (1990) Bacterial origin of a chloroplast intron: conserved self splicing group I introns in cyanobacteria. Science 250:1566–1569
Zubkot MK, Zubkot EI, van Zuilen K, Meyer P, Day A (2004) Stable transformation of petunia plastids. Transgenic Res 13:523–530
Zwickl DJ (2006) Genetic algorithm approaches for the phylogenetic analysis of large biological sequence datasets under the maximum likelihood criterion. Dissertation, the university of Texas at Austin. [GARLI available at http://www.bio.utexas.edu/faculty/antisense/garli/Garli.html]
Acknowledgments
Investigations reported in this article were supported in part by grants from the USDA (3611-21000-017-00D) and NIH (R01 GM 63879) to Henry Daniell. Research by Kenneth J. Wurdack and Robert K. Jansen was supported, in part, by NSF AToL grants EF 0431242 and DEB 0120709, respectively. The authors thank Kenneth Olsen, the Royal Botanic Gardens, Kew, and the herbaria cited for tissue or DNA samples.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by R. Hagemann.
Appendix
Appendix
Taxa and voucher information for 250 accessions (251 taxa) of Malpighiales and related families examined for atpF intron status. Taxa sequenced for atpF are followed by the GenBank accession number. Herbarium acronyms follow Index Herbariorum (http://sciweb.nybg.org/science2/IndexHerbariorum.asp).
Intron present
(i) Malpighiales
Achariaceae: Acharia tragodes Thunb., Cloete s.n. (BOL); Erythrospermum phytolaccoides Gardn., Chase 1277 (K); Hydnocarpus sp., Chase 1301 (K); Kiggelaria africana L., Chase 5607 (K). Balanopaceae: Balanops vieillardii Baill., Chase 1816 (K). Bonnetiaceae: Bonnetia sessilis Benth., Berry s.n. 25.7.98 (MO). Caryocaraceae: Anthodiscus amazonicus Gleason & A. C. Sm., van der Werff 13889 (MO); Caryocar glabrum Pers., Mori 22997 (NY). Clusiaceae s.l.: Mammea siamensis T. Anders., Chase 1216 (K); “Mesua” sp. (probably Kayea sp.), Coode 7884 (K); Montrouziera cauliflora Planch. & Triana, Lowry 5601 (MO); Pentaphalangium latissimum Lauterb., Chase 2100 (K); Rheedia macrophylla Planch. & Triana, Chase 1219 (K). Ctenolophonaceae: Ctenolophon englerianus Mildbr., McPherson 16911 (MO). Dichapetalaceae: Dichapetalum sp., Chase 624 (K?). Euphorbiaceae s.s.: Acalypha californica Benth., Levin 2192 (SD) GenBank EU293932; Adriana tomentosa (Thunb.) Gaudich. var. tomentosa, Cameron s.n. (US); Aleurites moluccana (L.) Willd., Wurdack s.n. (US); Amperea xiphoclada (Sieber ex Spreng.) Druce, Chase 1936 (K); Anthostema madagascariense Baill., Hoffmann 291 (K); Aparisthmium cordatum (A. Juss.) Baill., Bell et al. 93-60 (US); Astraea lobatum Klotzsch, Steinmann 2024 (RSA); Baloghia inophylla (G. Forst.) P. S. Green, Chase 3062 (K); Bernardia myricifolia (Scheele) S. Watson, McGill & Sundell 5459 (K); Bertya rosmarinifolia Planch., Chase 1938 (K); Beyeria leschenaultii (DC.) Baill., Chase 1939 (K); Blumeodendron sp., Chase 1252 (K); Calycopeplus casuarinoides L.S. Sm., Steinmann 1407 (RSA); Caperonia palustris (L.) A. St.-Hil., Wurdack D073 (US); Cavacoa aurea (Cavaco) J. Léonard, Luke & Robertson 1922 (US); Cephalomappa malloticarpa J.J. Sm., M. Chase 1256 (K); Chaetocarpus africanus Pax, Gabon, White 1319 (MO); Cladogynos orientalis Zipp., Chamchanroon 2010 (L); Claoxylon australe Baill. ex Müll. Arg., Chase 1937 (K); Clutia pulchella L., Chase 5876 (K); Cnesmone javanica Blume, Huq 10189 (US); Codiaeum variegatum (L.) Blume, Wurdack D033 (US), GenBank EU293933; Colliguaja integerrima Gillies & Hook., Landrum 8234 (MO); Conceveiba martiana Baill., Bell 93-176 (US); Croton alabamensis E.A. Sm. ex Chapm. var. alabamensis, Wurdack D008 (US), GenBankEU293934 Croton alabamensis E.A. Smith ex Chapman var. texensis Ginzbarg, Nesom 7850 (NY); Croton betulinus Vahl, Steinmann 2026 (RSA); Croton chilensis Müll. Arg., Wurdack D646 (US); Croton conduplicatus Kunth, Berry 5542 (MO); Croton cuneatus Klotzsch, Berry 7589 (PORT); Croton elliottii Chapm., Wurdack D455 (US); Croton flavens L., Steinmann 2015 (RSA); Croton guianensis Aubl., Berry 5535 (MO); Croton martinianus V.W.Steinm., Steinmann 606 (RSA); Croton nanus Gagnep., Pooma 4256 (BKF); Croton repens Schltdl., Steinmann 1062 (RSA); Croton setigerus Hook., Hughey s.n. (US); Croton socotranus Balf. f., Wurdack D644 (US); Croton speciosus Müll. Arg., Berry 7590 (MO); Croton suaveolens Torr., Devender 96-257 (RSA); Croton trinitatis Millsp., Berry 7586 (MO); Crotonogynopsis usambarica Pax, Tanzania, Polhill et al. 5129 (K); Dalechampia spathulata (Scheidw.) Baill., Wurdack D010 (US); Dichostemma glaucescens Pierre, Reitsma 1285 (NY); Ditaxis argothamnoides (Bertol. ex Spreng.) Radcl.-Sm. & Govaerts, Wurdack D105 (US); Ditrysinia fruticosa (Bartram) Govaerts & Frodin, Wurdack D149 (US); Ditta myricoides Griseb., Cedeño s.n. (US); Dodecastigma amazonicum Ducke, Mori 22811 (US); Endospermum moluccanum (Teijsm. & Binn.) Kurz, Chase 1258 (K); Enriquebeltrania crenatifolia (Miranda) Rzed., Cabrera 10768 (NY); Erythrococca sp., Cameroon, Cheek s.n. (K); Euphorbia epithymoides L., Chase 102 (NCU); Euphorbia mesembryanthemifolia Jacq., Wurdack D102 (US); Euphorbia pulcherrima Willd. ex Klotzsch cv. ‘Brilliant Diamonds’, Wurdack D084 (US); Excoecaria agallocha L., Motley & Cameron 2053 (NY); Excoecaria cochinchinensis Lour., Chase 1260 (K); Garcia nutans Vahl ex Rohr, Wurdack D051 (US); Gitara venezolana Pax & K. Hoffm., Bell et al. 94-312 (US); Gymnanthes lucida Sw., Wurdack D055 (US); Hippomane mancinella L., Wurdack D053 (US); Homalanthus populneus (Geiseler) Pax, Chase 1266 (K); Hura crepitans L., Wurdack D089 (US) GenBankEU293935; Jatropha gossypifolia L., Tayor 11763 (MO); Jatropha integerrima Jacq., Wurdack D047 (US); Joannesia princeps Vell., Chase 1262 (K); Klaineanthus gaboniae Pierre, White (ser. 2) 334 (MO); Koilodepas bantamense Hassk., Chase 1263 (K); Lasiocroton bahamensis Pax & K. Hoffm., Wurdack D058 (US); Leucocroton microphyllus (A. Rich.) Pax & K. Hoffm., HAJB 81915 (HAJB); Mabea sp., Bell et al. 94-30 (US); Macaranga grandifolia (Blanco) Merr., Wurdack D046 (US); Mallotus japonicus (L. f.) Müll. Arg., Wurdack D004 (US); Mallotus philippensis (Lam.) Müll. Arg., Wurdack D822 (US); Manniophyton africanum Müll. Arg., White 3366 (MO); Maprounea guianensis Aubl., Wurdack D332 (US); Melanolepis multiglandulosa Reichb. & Zoll., Motley 2493 (NY); Micrococca capensis (Baill.) Prain, Kurzweil 463/89 (K). Monadenium guentheri Pax, Chase 1007 (K); Monotaxis grandiflora Endl., Horn 2386 (DUKE); Moultonianthus leembruggianus (Boerl. & Koord.) Steenis, Challen et al. 3 (K); Nealchornea yapurensis Huber, Fine s.n. (US); Neoboutonia mannii Benth. & Hook. f., Fay 6701 (MO); Neoscortechinia kingii (Hook. f.) Pax & K. Hoffm., Chase 1265 (K) GenBank EU293936; Neoshirakia japonica (Siebold & Zucc.) Esser, Wurdack D005 (US); Omphalea diandra L., Chase 570 (K); Ostodes paniculata Blume, Chase 1267 (K); Pachystroma longifolium (Nees) I. M. Johnst., Prince s.n. (US); Pausandra martinii Baill., Berry 7466 (MO); Pedilanthus tithymaloides (L.) Poit., Wurdack D034 (US); Pera bicolor (Klotzsch) Müll. Arg., Gillespie 4300 (US), GenBank EU293937 Pimelodendron zoanthogyne J.J. Sm., Chase 1268 (K); Plukenetia volubilis L., Armbruster 38 (?); Pogonophora schomburgkiana Miers ex Benth., Larpin 1022 (US); Pseudosenefeldera inclinata (Müll. Arg.) Esser, Wurdack D385 (US); Ricinocarpos tuberculatus Müll. Arg., Chase 2164 (K); Ricinodendron heudelotii (Baill.) Heckel, Chase 1269 (K); Ricinus communis L., Wurdack D009 (US); Seidelia triandra (E. Mey.) Pax, Giess 13381 (MO); Senefelderopsis croizatii Steyerm., Berry s.n. (MO); Spathiostemon javensis Blume, Chase 1261 (K); Stillingia sylvatica L. subsp. tenuis (Small) D. J. Rogers, Wurdack D117 (US); Strophioblachia fimbricalyx Boerl., Chase 1270 (K); Sumbaviopsis albicans (Blume) J.J. Sm., Chase 1271 (K); Suregada glomerulata (Blume) Baill., Chase 1272 (K); Synadenium grantii Hook. f., Wurdack D259 (US); Syndyophyllum occidentale (Airy Shaw) Welzen, Aik et al. SAN-111913 (L); Tannodia cordifolia (Baill.) Baill., Ralimanana 293 (K); Tetrorchidium cf. macrophyllum Müll. Arg., Bell et al. 93-204 (US), GenBank EU293938; Tragia fallax Müll. Arg., Bell et al. 94-378 (US); Tragia urticifolia Michx., Wurdack D074 (US); Tragiella anomala (Prain) Pax & K. Hoffm., Mwasumbi et al. 16202 (MO); Trewia nudiflora L., Chase 1273 (K); Triadica sebifera (L.) Small, Wurdack D059 (US); Trigonostemon verrucosus J. J. Sm., Chase 1274 (K); Vernicia montana Lour., Chase 2105 (K). Euphroniaceae: Euphronia guianensis (R. H. Schomb.) H. Hallier, Mori 23699 (NY). Goupiaceae: Goupia glabra Aubl., Prevost 3031 (CAY). Humiriaceae: Humiria wurdackii Cuatrec., Wurdack s.n. (US); Sacoglottis gabonensis Urb., Stone 3283 (MO); Vantanea guianensis Aubl., Pennington 13855 (K). Hypericaceae: Cratoxylum formosum (Jack) Benth & Hook. f. ex Dyer, Chase 1218 (K). Irvingiaceae: Irvingia malayana Oliv., Simpson 2638 (K?); Klainedoxa gabonensis Pierre, Bradley et al. 1092 (MO). Ixonanthaceae: Cyrillopsis paraensis Kuhlm., Hentrich 68 (NY); Ochthocosmus longipedicellatus Steyerm. & Luteyn, Berry 6561 (MO). Lacistemataceae: Lacistema aggregatum Rusby, Pennington et al. 583 (K); Lozania pittieri (Blake) L. B. Sm., Pennington et al. 584 (K). Linaceae: Durandea pentagyna K. Schum., Takeuchi 7103 (MO); Linum perenne L., Chase 111 (NCU); Reinwardtia indica Dumort., Chase 230 (NCU). Malpighiaceae: Dicella nucifera Chodat, Anderson 13607 (MICH); Galphimia gracilis Bartl., Adelson s.n. (MICH), GenBank EU293939. Medusagynaceae: Medusagyne oppositifolia Baker, Fay s.n. (K) [Kew 1981-2059]. Ochnaceae s.s.: Elvasia calophyllea DC., Amaral s.n. (K?); Lophira lanceolata Tiegh., Schmidt 1902 (MO); Ochna multiflora DC., Chase 229 (NCU). Pandaceae: Galearia filiformis (Blume) Boerl., Chase 1334 (K); Microdesmis pierlotiana J. Léonard, Gereau et al. 5654 (MO); Panda oleosa Pierre., Schmidt et al. 2048 (MO). Phyllanthaceae: Amanoa strobilacea Müll. Arg., McPherson 16826 (MO); Apodiscus chevalieri Hutch., Schmidt et al. 2094 (MO); Aporusa frutescens Blume, Chase 1251 (K); Astrocasia neurocarpa (Müll. Arg.) I. M. Johnst. ex Standl., Wurdack D743 (US); Bischofia javanica Blume, Levin 2200 (SD), GenBank EU293940; Cleistanthus oblongifolius (Roxb.) Müll. Arg., Chase 1257 (K); Croizatia brevipetiolata (Secco) Dorr, Dorr et al. 8555 (US), GenBank EU293941; Didymocistus chrysadenius Kuhlm., Gillespie et al. 4805 (US); Discocarpus essequeboensis Klotzsch, Hoffman 996 (US); Gonatogyne brasiliensis (Baill.) Müll. Arg., Cordeiro & Esteves 1384 (K); Heywoodia lucens Sim, Saufferer & Muchai SS-1544 (US); Hieronyma oblonga (Tul.) Müll. Arg., Bell 94-252 (US); Hymenocardia acida Tul., Walters et al. 897 (MO); Lachnostylis bilocularis R. A. Dyer, Kurzweil 83/88 (K); Leptonema glabrum (Leandri) Leandri, McPherson & Rabenantoandro 18389 (MO); Maesobotrya vermeulenii (de Wild.) J. Léonard, Bradley et al. 1032 (MO); Poranthera huegelii Klotzsch, Spjut 7369 (US); Richeria grandis Vahl, Merello et al. 1714 (MO); Sauropus racemosus Beille, Soejarto et al. 10648 (NY); Spondianthus preussii Engl., Merello et al. 1661 (MO); Zimmermania ovata E.A. Bruce, Faden s.n. (US). Picrodendraceae: Androstachys johnsonii Prain, Chase 1904 (K); Hyaenanche globosa (Gaertn.) Lamb. & Vahl, Chase 1445 (K); Picrodendron baccatum Krug & Urb. ex Urban, Wurdack D050; Podocalyx loranthoides Klotzsch, Berry & Aymard 7226 (MO), GenBank EU293955 (partial sequence not used in phylogenetic analyses); Tetracoccus dioicus Parry, Levin 2202 (DUKE), GenBank EU293942. Podostemaceae: Marathrum c.f. oxycarpum Tul., Gabriel da Cachoeirc 9/1996 (AM); Podostemum ceratophyllum Michx., Horn & Wurdack s.n. (DUKE). Putranjivaceae: Drypetes capillipes Pax & K. Hoffm., Harris 4884 (K); Drypetes fallax Pax & K. Hoffm., Harris 4892 (K); Drypetes macrostigma J.J. Sm., Chase 1259 (K); Putranjiva roxburghii Wall., Wurdack D057 (US), GenBank EU293956 (partial sequence not used in phylogenetic analyses). Quiinaceae: Quiina pteridophylla (Radlk.) Pires, Pires s.n. (CPATU) Touroulia guianensis Aubl., Pires (CPATU). Rhizophoraceae s.s.: Bruguiera gymnorhiza Lam., Chase 12838 (K); Carallia brachiata (Lour.) Merr., Chase 2151 (K); Crossostylis multiflora Brongn. & Gris ex Pancher & Sebert, Lowry 5686 (MO); Paradrypetes subintegrifolia G. A. Levin, Acevedo-Rdgz. & Cedeño 7560 (US); Cassipourea lanceolata Tul., Schatz 3689 (MO). Salicaceae s.l.: Abatia parviflora Ruiz & Pav., Pennington 676 (K); Casearia nitida Jacq., FTG 72496; Flacourtia jangomas Steud., Chase 2150 (K); Idesia polycarpa Maxim., Wurdack D22 (US); Poliothyrsis sinensis Oliver, Wurdack D029 (US); Scyphostegia borneensis Stapf, Beaman 911 (BH). Trigoniaceae: Trigoniastrum hypoleucum Miq., Og B128 (L). Violaceae: Hybanthus concolor Spreng., Wurdack D148 (US); Hymenanthera alpina Oliv., Chase 501 (K); Rinorea bengalensis Kuntze, Chase 2148 (K).
(ii) Other related rosids (Celastrales, Oxalidales + Huaceae)
Brunelliaceae: Brunellia acutangula Humb. & Bonpl., Stergios 20646 (US), GenBank EU293943. Celastraceae: Brexia madagascariensis Thouars, Wurdack D764 (US); Maytenus senegalensis (Lam.) Exell, Collenette 4/93 (K); Plagiopteron suaveolens Griff., Chase 1335 (K); Siphonodon celastrineus Griff., Chase 2097 (K); Stackhousia minima Hook. f., Molloy s.n. (CHR); Tripterygium regelii Sprague & Takeda, Chase 1003 (K). Cephalotaceae: Cephalotus folicularis Labill., Chase 147 (NCU). Connaraceae: Rourea minor Leenh., Chase 1221 (K). Cunoniaceae: Eucryphia milliganii Hook. f., Chase 2528 (K). Elaeocarpaceae: Crinodendron hookerianum Gay, Chase 909 (K); Sloanea sp., Chase 343 (NCU). Huaceae: Afrostyrax sp., Cheek 5007 (K). Lepidobotryaceae: Ruptiliocarpon caracolito Hammel & Zamora, Hammel 19102 (MO). Oxalidaceae: Averrhoa carambola L., Chase 214 (NCU). Parnassiaceae: Parnassia grandifolia DC., Wurdack D795 (US), GenBank EU293944.
Intron absent
(i) Malpighiales
Elatinaceae: Elatine hexandra DC., Chase 2978 (K), GenBank EU293945. Euphorbiaceae s.s.: Cnidoscolus urens (L.) Arthur var. stimulosus (Michx.) Govaerts, Wurdack D002 (US); Elateriospermum tapos Blume, Soepadmo & Suhaimi s193 (NY), GenBank EU293946; Glycydendron amazonicum Ducke, Gillespie et al. 4546 (US); Glycydendron cf. amazonicum Ducke, Mori et al. 23265 (US); Hevea sp. [cf. pauciflora (Spruce ex Benth.) Müll. Arg.], Gillespie 4272 (US), GenBankEU293947; Manihot esculenta subsp. esculenta Crantz, EU117376; Manihot foetida Pohl, Olsen s.n.; Manihot grahamii Hook., Wurdack s.n. (US); Manihot pauciflora Brandegee, Villasenor 1248 (NY); Manihot walkerae Croizat, Olsen s.n.; Micrandra inundata P. E. Berry & A. Wiedenhoeft, Berry 6350 (MO). Lophopyxidaceae: Lophopyxis maingayi Hook. f., Adelbai P-10203 (US), GenBank EU293948. Malesherbiaceae: Malesherbia weberbaueri Gilg, Weigend 2000/365 (NY), GenBank EU293949. Passifloraceae s.s.: Paropsia madagascariensis (Baill.) H. Perrier, Zyhra 949 (WIS); Passiflora coccinea Aubl., Chase 2475 (K), GenBank EU293950. Phyllanthaceae: Breynia disticha J. R. Forst. & G. Forst. cv. ‘Rosea Picta’, Wurdack D012 (US); Flueggea virosa (Roxb. ex. Willd.) Voigt subsp. virosa, Wurdack D101 (US); Glochidion puberum (L.) Hutch., Wurdack D003 (US); Leptopus colchicus (Fisch. & C. A. Mey. ex Boiss.) Pojark., Wurdack D778 (US), GenBank EU293951; Leptopus esquirolii (H. Lév.) P. T. Li, 1980 Sino-American Bot. Exped. 1678 (US); Margaritaria tetracocca (Baill.) G. L. Webster, Wurdack D044 (US); Phyllanthus epiphyllanthus L., Wurdack D056 (US), GenBank EU293952; Phyllanthus fluitans Benth. ex Müll. Arg., Wurdack D761 (US); Phyllanthus juglandifolius Willd., Wurdack D759 (US); Phyllanthus nutans Sw., Wurdack D762 (US); Reverchonia arenaria A. Gray, Atwood 17245 (NY); Savia bahamensis Britton, Wurdack D048 (US). Picrodendraceae: Austrobuxus megacarpus P. I. Forster, Forster 21239 (BRI), GenBank EU293953; Choriceras majus Airy Shaw, Forster 21230 (BRI); Dissiliaria muelleri Baill., Forster 14629 (BRI); Whyanbeelia terrae-reginae Airy Shaw & B. Hyland, Forster 21231 (BRI). Turneraceae: Tricliceras longipedunculatum (Mast.) R. Fern., Kanji et al. 304 (NY); Turnera ulmifolia L., Wurdack s.n. (US), GenBank EU293954.
(ii) Other related rosids (Celastrales, Oxalidales + Huaceae): none noted.
Rights and permissions
About this article
Cite this article
Daniell, H., Wurdack, K.J., Kanagaraj, A. et al. The complete nucleotide sequence of the cassava (Manihot esculenta) chloroplast genome and the evolution of atpF in Malpighiales: RNA editing and multiple losses of a group II intron. Theor Appl Genet 116, 723–737 (2008). https://doi.org/10.1007/s00122-007-0706-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00122-007-0706-y