Abstract
Background
Haemoproteus is a sister genus to malaria parasites (Plasmodium), which both belong to the order Haemosporida (Apicomplexa). Parasites of both genera are flourishing in birds, however, Haemoproteus species are noticeably less investigated. This is unfortunate because knowledge about close relatives of malaria pathogens is important for better understanding the evolutionary origin and basic biological features of the entire group of haemosporidian infections. Moreover, recent findings show that Haemoproteus species can cause severe damage of various bird organs due to megalomeronts and other exo-erythrocytic stages. These haemosporidians are remarkably diverse, but remain neglected partly due to difficulties in species identification. Hundreds of Haemoproteus genetic lineages have been reported in birds, and numerous new lineages are found each year, but most remain unidentified to the species level. Numerous new Haemoproteus pathogens were described during the past 20 years. However, keys for their identification are absent. Identification of Haemoproteus species remains a difficult task and is an obstacle for better understanding of the distribution and epidemiology of these parasites. This study aimed to develop comprehensive keys for the identification of described avian Haemoproteus species using morphological features of their blood stages (gametocytes).
Methods
Type and voucher preparations of avian Haemoproteus species were accessed in museums in Europe, Australia and the USA. Gametocytes of most described species were examined, and these data formed a background for this study. The data also were considered from published articles containing parasite species descriptions. The method of dichotomous keys was applied. The most difficult steps in the keys were accompanied with references to the corresponding parasite pictures.
Results
In all, 201 published articles were included in this review. Morphological diagnostic features of gametocytes of all described Haemoproteus species were analysed and compared. Illustrated keys for identification of these parasite species were developed. Available information about the molecular characterization of Haemoproteus parasites was provided.
Conclusion
This review shows that 177 described species of avian Haemoproteus can be distinguished and identified in blood films using morphological characters of their gametocytes and host cells. These species were incorporated in the keys. Information about possible morphologically cryptic parasites was provided. Molecular markers are available for only 42% of the described Haemoproteus parasites, calling for researchers to fill this gap.
Similar content being viewed by others
Background
Order Haemosporida (Alveolata, Apicomplexa, Sporozoa) unites a diverse group of closely related obligate heteroxenous protists, whose currently are classified in four families—Plasmodiidae, Haemoproteidae, Leucocytozoidae and Garniidae [1,2,3,4,5]. These pathogens have many similar basic features in their life cycles. Mainly, they are transmitted exclusively by dipteran blood-sucking insect (Diptera), in which the sporozoites (invasive stage for vertebrates) develop. Sporozoites initiate the exo-erythrocytic development (exo-erythrocytic merogony or schizogony) in various tissues of vertebrate hosts. Exoerythrocytic merozoites are capable of infecting blood cells, in which gametocytes (the invasive stage for vectors) finally inhabit. Asexual dividing stages (erythrocytic meronts or schizonts) do not develop in Haemoproteus species. Gametocytes are characterized by sexually dimorphic characters. Development in vectors is similar in all haemosporidians; it consists of exflagellation, which is followed by the sexual process of the oogamy, development of motile ookinetes, sporogony, and finally the formation of sporozoites, which complete their maturation in the salivary glands of vectors. Infection of vertebrate hosts occurs actively during the blood meal of vectors, who inject sporozoites into the blood stream. Similarities in life cycles and other features of the biology (particularly ultrastructure) of haemosporidians belonging to different families [3, 5,6,7] have been supported by molecular phylogenies, which indicate that these parasites are relatives and likely have a common origin [8,9,10,11,12]. In other word, accumulation of new knowledge about different groups of haemosporidians, which are remarkably diverse and flourishing in wildlife [5, 6, 13,14,15], would be beneficial for better understanding the biology of the entire group of Haemosporida, including agents of diseases of domestic animals and humans, as well as malaria [10].
Molecular phylogenetic data show that Haemoproteus parasites (Haemoproteidae) are a sister group to malaria agents of the genus Plasmodium (Plasmodiidae) [8, 9, 11]. Haemosporidians of both these genera do not digest haemoglobin completely, resulting in the accumulation of residual pigment (haemozoin) in their blood stages [1,2,3]. This feature unites species of Haemoproteus and Plasmodium and distinguishes them from species of the Leucocytozoidae and Garniidae, which do not produce residual pigment when developing in red blood cells. The life cycles of Haemoproteus and Plasmodium parasites differ mainly due to the inability of the haemoproteids (i) to multiply in blood cells (erythrocytic merogony is absent) and (ii) to complete sporogony in mosquitoes [7]. Haemoproteus parasites are transmitted mainly by Culicoides biting midges (Ceratopogonidae), and a few species are vectored by louse flies (Hippoboscidae) [2, 7, 16]. In spite of these differences, the knowledge on Haemoproteus spp. is important for better understanding evolutionary biology of haemosporidians [17,18,19], including human malaria parasites of genus Plasmodium [10]. For example, the application of Haemoproteus tartakovskyi genomic information in phylogenetic studies contributes to understanding the evolutionary relationships of Laverania parasites [8].
Haemosporidians of the genus Haemoproteus parasitize only birds and reptiles [1, 3, 6]. These protists remain a neglected group of blood pathogens mainly because they have been traditionally considered to be relatively benign to their hosts [20]. This seems to be true in regard to the blood pathology during haemoproteosis due to the absence of multiplication in blood cells and predominantly light or moderate parasitaemia, which rarely reaches 5%, but usually is less than 1% in wild-caught animals [3]. However, recent studies show that haemoproteids are pathogenic to blood-sucking insects, including mosquitoes, who often die within 12 h after taking heavily infected blood meals due to damage caused by the migration of ookinetes [21,22,23]. Furthermore, the application of molecular diagnostic tools has proven that large-sized megalomeronts (up to 300 µm and even bigger) develop in many Haemoproteus infections, resulting in the damage of various organs [2, 24,25,26,27,28,29,30]. These findings call for research aimed at better understanding the biology of haemoproteids in regard of animal health.
Haemoproteus species are cosmopolitan and often prevalent in birds [3, 5, 14, 31,32,33,34]. Molecular studies have revealed over 4600 unique cytochrome b gene lineages of these parasites, and many more likely exist [13] (see MalAvi database http://130.235.244.92/Malavi, accessed in April 2022). Over 170 species of Haemoproteus have been described, including 49 new species described during the past 20 years. Difficulties in species identification using morphological data preclude comprehensive parasite diversity research. This is unfortunate because morphological and molecular data complement each other and are essential in obtaining a true understanding of pathogen diversity in certain bird populations [35, 36], particularly during co-infections of parasites belonging to same genus [37,38,39]. Species-specific molecular markers are absent for the majority of avian Haemoproteus, and currently they are difficult to design and use due to the vast genetic diversity of these pathogens, most of which remain insufficiently investigated or even non-described in wildlife. Morphological identification using microscopic examination of blood films supplements the information providing by polymerase chain reaction (PCR)-based diagnostic tools and remains important in the research of wildlife haemosporidians [36, 38, 40]. Thus, the available keys for the identification of avian Haemoproteus species [3] should be reworked and supplemented.
This review aimed to simplify the identification of avian Haemoproteus species by developing easy-to-use keys based on morphological characters of gametocytes, the parasite development stage easily accessed due to presence in the peripheral circulation. This article provides comprehensive keys, which include new Haemoproteus species that are not already found in formerly published keys [3]. This should assist academic and veterinary medicine researchers in the identification of Haemoproteus pathogens. The available information about molecular markers (molecular barcodes), which can be used for the detection of described Haemoproteus species and the comparative research, was summarized as well. This study generalizes the over 40-years of experience of the authors in taxonomy of Haemoproteus species, and is designed for researchers who are interested in wildlife pathogens.
Methods
Full-length papers with descriptions of new Haemoproteus species as well as articles with re-descriptions and molecular characterizations of these parasites published in peer-reviewed journals were considered. In all, 201 articles and books were reviewed, and 191 publications containing the representative morphological and/or PCR-based information related to identification of these parasites were cited and incorporated in the References.
Type and voucher preparation as well as gametocyte images of avian Haemoproteus parasites were obtained from the collections of Nature Research Centre (Vilnius, Lithuania), International Reference Centre for Avian Haematozoa (Queensland Museum, Queensland, Australia), the US National Parasite Collection (National Museum of Natural History, Washington DC, USA), Natural History Museum (London, UK), Muséum National d’Histoire Naturelle (Paris, France), Grupo de Estudio Relación Parásito Hospedero, Universidad Nacional de Colombia (Bogotá, Colombia) and individual researchers. All accessed preparations were examined. An Olympus BX61 light microscope (Olympus, Tokyo, Japan) equipped with an Olympus DP70 digital camera and imaging software AnalySIS FIVE (Olympus Soft Imaging Solution GmbH, Münster, Germany) was used to examine preparations and prepare illustrations. It is important to note that the staining quality of some old type specimens of Haemoproteus species, which were obtained from museums, was of insufficient quality mainly due to fading, resulting in some poorly visible diagnostic characters. However, images of the parasites from such preparations were included in this review and were shown in the corresponding species figures if they provided valuable information about the general shape of gametocytes, the size of pigment granules, the mode of influence on host cells and others readily visible taxonomic characters. Use of this information helps to distinguish some species, and also is important for future taxonomic studies being the source of illustrations from valuable type specimens. The quality of these slides could hardly be improved in the future, so worth documentation at this stage. Black and white drawings were also provided to illustrate morphological details of the parasites, of whose high-quality photographs were not available.
Classical dichotomous keys were developed for the identification of Haemoproteus species [3]. Each key consists of steps divided into two alternatives, which identifies the species of a specimen following a series of simple choices that lead the user to the correct name of a given species. The most difficult choices, which might lead to ambiguity, were accompanied with references to the corresponding pictures that further illustrate meaning of the text information. This simplifies the comparison of diagnostic features used in the keys and minimizes possible misunderstanding. All species names in the keys were accompanied with references to the original parasite descriptions, re-descriptions and (or) other publications, which contain description and (or) illustrations of corresponding parasites. Published articles containing valuable morphological descriptions were collected, analysed and cited. These references help to access parasite descriptions and confirm a parasite identification. Information about the barcoding DNA sequences, which can be used for molecular detection and identification of corresponding parasites are also provided.
The experimental observations showed that Haemoproteus species vary in vertebrate host specificity, but the same parasite usually cannot complete life cycle and produce invasive stages (gametocytes) in birds belonging to different orders (see review in [3, p. 69]. Molecular sequence information is in accordance with these empirical data and indicates only rare cases when the same Haemoproteus lineages could be found in birds belonging to different orders [13]. Importantly, the rare reports of the same Haemoproteus lineages in birds of different orders have never been supported by the observation of the corresponding species gametocytes, an invasive parasite stage for vectors, indicating abortive (incomplete) development, which is a dead-end of infection [7]. Thus, morphologically similar parasites in birds belonging to different orders are different species in most cases. This conclusion was confirmed by molecular data [7, 9, 11, 13]. Due to vertebrate host specificity and the resulting restriction of parasite distributions by bird orders, the natural host range of haemosporidians remains helpful in species identification. This provided an opportunity to design separate keys for identification of parasites inhabiting birds of different orders. This approach simplifies parasite identification by minimizing the number of species, which are needed for comparison before making the final conclusion about a species identity. This approach was used in the keys.
Approximately 50% of all described Haemoproteus species parasitize birds of the order Passeriformes. Due to the marked species diversity of haemoproteids in passerines, the keys for their identification were developed for closely related passeriform bird families, which were grouped in suborders or superfamilies as suggested by the current bird phylogenies [41, 42]. Such keys provide the opportunity to easily identify the majority of described parasites of passeriform birds. However, it should be noted that some Haemoproteus parasites of passerines might infect and produce gametocytes in birds belonging to different families within the same order [43], supporting information that host taxonomic characteristic cannot be considered as the main parasite taxonomic character [3, 44]. This might lead to circumstances when a sample under identification could be not found in a certain key. This also certainly will happen if a researcher is dealing with a new (non-described) parasite species. Further comparison of such samples with parasites of most closely related avian groups is suggested before making a final conclusion about the identity of a sample. This is a relatively weak point of the keys for the identification of Haemoproteus species parasitizing passeriform birds. However, experience shows [3] that creating one big key for all parasites of Passeriformes birds would be even more difficult to use due to the need of too many minor morphological characters, which often are difficult to estimate in practical work, particularly during low parasitaemia. This usually make the identification even more complicated in comparison to the approach, which was used here.
Genus Haemoproteus includes two subgenera—Haemoproteus and Parahaemoproteus. Species of these subgenera differ in patterns of sporogony, which occur in louse flies (family Hippoboscidae) and Culicoides biting midges (family Ceratopogonidae), respectively [3, 7]. Haemoproteus and Parahaemoproteus parasites are indistinguishable at the gametocyte stages, but might be present in birds of the same orders. They also sometimes occur in co-infection [3]. The current examples are the parasites of Columbiformes and Suliformes birds [45, 46]. To facilitate identification of species of both subgenera using morphological characters of gametocytes, these parasites were given in the same keys. Most of avian haemoproteid species belong to Parahaemoproteus. All species of subgenus Haemoproteus were indicated in notes to the corresponding keys.
Results
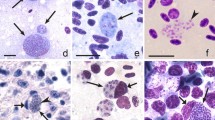
Birds are hosts of various intracellular blood parasites belonging to Apicomplexa. For example, they are infected by haemosporidians of the genera Plasmodium, Leucocytozoon, Garnia, Fallisia and related species of Babesia, Isospora, Lankesterella and Hepatozoon [2,3,4, 47, 48]. These organisms (Fig. 1a–o) often occur in co-infections. Haemoproteus species can be readily distinguished from all other avian intracellular haematozoa, except Plasmodium parasites, because their blood stages (gametocytes) develop exclusively in red blood cells and always contain refractive pigment granules (haemozoin granules) (Fig. 1a–c), which are absent in all other blood parasites (Fig. 1h–o). Even the smallest Haemoproteus gametocytes contain pigment granules and can be distinguished from other intracellular blood parasites (Fig. 1a), except for Plasmodium species, which sometimes look similar at the gametocyte or early trophozoite stages (Fig. 1d, e). Furthermore, it is important to note that gametocytes of some species of Haemoproteus are similar to the elongate gametocytes of malaria parasites (Fig. 1b–d). Species of Plasmodium can be readily distinguished because they multiply in blood cells and produce meronts (= schizonts), in which invasive merozoites develop (Fig. 1f, g). This is not the case in species of Haemoproteus. Extensive examination of blood films at magnification 500× usually provides opportunity to visualize meronts of Plasmodium even during low parasitaemia and thus to identify malaria infections. Examination of blood films using an oil immersion objective 50× is particularly convenient for this diagnostic procedure. However, an obstacle still might be a low Plasmodium sp. parasitaemia (< 0.001% of infected red blood cells), which however is the obstacle for identification of any haematozoan infection using microscopic examination of blood films and sometimes even sensitive PCR-based methods [38].
Main morphological features of blood stages, which are helpful to distinguish Haemoproteus parasites (a–c) from other avian intracellular protists (d–o). Young (a) and fully grown (b, c) gametocytes of Haemoproteus species. Fully grown gametocytes (d, e) and mature erythrocytic meronts (f, g) of Plasmodium species. Growing meronts of Garnia (h) and Fallisia (i) species. Gametocytes (j, k) in roundish host cell (j) and fusiform host cell (k) of Leucocytozoon species. Growing meront (l) of Babesia species. Merozoite (m) of Isospora species. Sporozoite (n) of Lankesterella and gamont (o) of Hepatozoon species. Note presence of malarial pigment (haemozoin) in species of Haemoproteus (a–c) and Plasmodium (d–g) and its absence in species of other avian blood parasites (h–o). Elongate gametocytes of malaria parasites belonging to the subgenera Giovannolaia and Huffia (d) are similar to gametocytes of Haemoproteus species in forms, but the gametocytes of malaria parasites usually are more irregular in shape (d) and the outline of their macrogametocyte nuclei often is not so well indistinct (compare b and d). Presence of merogony in blood cells (f, g) clearly shows malaria infection. Long simple arrows—host cell nuclei. Short simple arrows—parasite nuclei. Simple arrowheads—pigment granules. Simple small arrowheads—merozoites. Simple wide short arrows—vacuoles. Triangle arrows—remnants of host cell cytoplasm. Other explanations are given in the text
It should be noted that a shape of fully grown gametocytes is an important character for many Haemoproteus species identifications [2, 3, 49]. Several major gametocyte forms are readily distinguishable and were used in the keys (Fig. 2a–r). These forms are microhalteridial (fully grown gametocytes are small and do not reach the poles of infected erythrocytes, Fig. 2b–f), halteridial (fully grown gametocytes reach and occupy the poles of infected erythrocytes, Fig. 2g–j), circumnuclear (fully grown gametocytes encircle the nuclei of infected erythrocytes completely, Fig. 2o, p) and rhabdosomal (fully grown gametocytes push the nuclei of erythrocytes to polar position and finally enucleate the host cells, Fig. 2q, r). A non-infected erythrocyte was shown in Fig. 2a for comparison with the infected erythrocytes. Sometimes subdivisions of these main forms were also helpful during species identification, and they were mentioned in the keys. These forms are broadly-halteridial (fully grown gametocytes occupy the poles of infected erythrocytes and markedly displace the nuclei laterally, Fig. 2k, l) and close to circumnuclear (fully grown gametocytes encircle the nuclei of infected erythrocytes nearly completely, Fig. 2m, n). It is important to remind in this regard that the form of only fully grown gametocytes was considered in the keys.
Main morphological forms of fully grown gametocytes, which are used in keys for identification of Haemoproteus species: roundish (b), microhalteridial (c–f), halteridial (g–j) and broadly-halteridial (k, l), close to circumnuclear (m, n), circumnuclear (o, p) and rhabdosomal (q, r). Uninfected erythrocyte (a) was shown for comparison purpose. All pictures show only fully grown gametocytes. Note that form of growing gametocytes often is different from the form of the fully grown gametocytes. That is why the form of young growing gametocytes usually was not mentioned in the keys, except for some rare cases, which were specified in the tables for species identification. The gametocyte forms depend on mode of parasite growth in red blood cells, which often is species-specific and was used in taxonomy. Various variations of halteridial (c–l) and circumnuclear (m–p) gametocyte forms predominate in avian haemoproteids. Roundish form (a) is particularly rare. Rhabdosomal forms (q, r) develop in several Haemoproteus species whose fully grown gametocytes enucleate infected erythrocytes. Dumbbell-like form of gametocytes (e) was often mentioned in the keys. Long simple arrows—host cell nuclei. Short simple arrows—parasite nuclei. Simple arrowheads—pigment granules. Other explanations are given in the text
Blood films should be carefully examined before parasite identification. When the most typical growing and fully-grown gametocytes of Haemoproteus are recognized in samples, the next step is the species identification using the keys. The latter were designed in the form of separate tables, each created for a separate group of avian hosts. To facilitate the use of the keys, a guide to all keys has been provided in Table 1. It lists all the tables for Haemoproteus species identification in relation to bird orders or relative families, as grouped in the keys.
Discussion
The recent discoveries of severe damage caused by tissue stages (meronts and megalomeronts) of Haemoproteus parasites in various organs, including the brain, heart, lungs, kidneys and skeletal muscles of naturally infected birds call for research aimed at better understanding the diversity and patterns of development of these pathogens [2, 24, 25, 27, 28, 170, 186]. This work requires parasite species identification and molecular data analysis. The combination of microscopic and PCR-based tools complements each other and increases the significance of research on haemosporidian parasite diversity, so is preferable, particularly in wildlife studies [36, 38, 40]. However, the progress in developing morphological identifications of Haemoproteus species is slow and noticeably falls behind the accumulation of DNA sequence information. This is unfortunate because the identification of the parasite species helps in accessing the basic data on the life cycles and patterns of pathogens’ development in various hosts and would be helpful for better understanding of haemoproteosis. The easy-to-use keys should stimulate taxonomic research and contribute to the discovery of new pathogen species by providing direct indications how to distinguish the parasites, which are morphologically different from described ones. In other words, the samples under identification, which are absent in the keys, likely represent new pathogens and would be worth targeted taxonomic investigation.
The following possible obstacles should be considered during the identification of Haemoproteus parasites using samples collected in wildlife. First, the intensity of parasitaemia is often low in naturally infected birds, and all necessary blood stages (young, growing or fully grown gametocytes) might be absent in a single sample. Experienced taxonomists can often perform the species identification by visualization of several Haemoproteus gametocytes, which are typical for the species, however this is hardly achievable for novices. As a result, low parasitaemia might limit the use of the keys. The negative effect of this factor can be minimized by sampling a sufficient number of host individuals belonging to the same species at the same study site. Extensive sampling of the same host species is common in population studies. The sample size needed for parasite taxonomical work depends on the prevalence of infection in a certain bird population [187]. The sampling of 10–30 host individuals often is sufficient and usually provides an opportunity to access various intensities of parasitaemia of the same parasite and then to visualize the full range of gametocytes, which are necessary for parasite species identification. This is preferable for the development of comprehensive descriptions of new pathogens.
Second, gametocytes of Haemoproteus inhabit erythrocytes, which are fragile cells and might be deformed during the preparation of blood films, resulting in presence of unusual parasite forms, which are artefacts from the taxonomic point of view and should be ignored during species identification and parasite descriptions. Morphological characters of typical (the most common) non-deformed gametocytes and their host cells should be selected for identification. Only such cells were used and shown in all corresponding illustrations in the keys (Figs. 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, 30, 31, 32, 33, 34, 35, 36).
Morphological features of gametocytes, which are used for identification of Haemoproteus species parasitizing Accipitriformes birds. Microgametocytes (a–c) and macrogametocytes (d–f) of Haemoproteus janovyi (a–c), H. elani (d), H. buteonis (e) and H. nisi (f). Note the markedly variable form of H. janovyi gametocytes (a–c), the broadly halteridial form of H. elani gametocyte (d), the presence of unfilled spaces on the poles of infected erythrocytes during H. buteonis infection (e), and the circumnuclear form of H. nisi gametocyte, which is overfilled with volutin granules (f). Images a–c are from the type material, which is fading, resulting in pale staining and poorly recognizable pigment granules and nuclei, however the overall form of the gametocytes is readily visible. Long simple arrows—host cell nuclei. Short simple arrows—parasite nuclei. Simple arrowheads—pigment granules. Simple wide long arrows—unfilled space on poles of infected erythrocytes. Triangle wide arrowheads—volutin granules. Other explanations are given in the text
Morphological features of gametocytes, which are used for identification of Haemoproteus species parasitizing Anseriformes birds. Macrogametocytes (a, b, e, d) and microgametocytes (c, f) of Haemoproteus macrovacuolatus (a, b), H. greineri (c, d) and H. nettionis (e, f). Note the presence of large vacuoles in the cytoplasm of H. macrovacuolatus macrogametocytes (a, b). Fully grown gametocytes of H. greineri are predominantly close to circumnuclear (d) and circumnuclear (c) in form, but fully grown gametocytes of H. nettionis are predominantly broadly halteridial (e, f). Prominent volutin granules (c, d) are present in gametocytes of H. greineri and H. nettionis (volutin was not shown in white-and black pictures e, f); these species are indistinguishable based on this character. Long simple arrows—host cell nuclei. Short simple arrows—parasite nuclei. Simple arrowheads—pigment granules. Simple wide short arrows—vacuoles. Triangle wide arrowheads—volutin granules. Other explanations are given in the text
Morphological features of gametocytes, which are used for identification of Haemoproteus species parasitizing Apodiformes birds. Macrogametocytes (a–e, g, h) and microgametocyte (f) Haemoproteus archilochus (a, b), H. witti (c, d), H. apodus (e, f) and H. trochili (g, h). Note the circumnuclear (a, b) and broadly halteridial (c, d) forms of fully grown gametocytes in H. archilochus (a, b) and H. witti (c, d), respectively. Gametocytes of H. apodus (e, f) and H. trochili (g, h) are microhalteridial in form. Haemoproteus apodus is readily distinguishable due to presence of numerous dumbbell-like growing gametocytes (e, f). Long simple arrows—host cell nuclei. Short simple arrows—parasite nuclei. Simple arrowheads—pigment granules. Simple wide long arrows—unfilled space between growing gametocytes and envelope of infected erythrocyte. Other explanations are given in the text
Morphological features of gametocytes, which are used for identification of Haemoproteus species parasitizing Bucerotiformes birds. Macrogametocytes (a, b) of Haemoproteus upupae (a) and H. bucerotis (b). Note the different position of nuclei in macrogametocytes—close to central (a) and subcentral (b)—of these species, and the different morphology and mode of distribution of pigment granules in the cytoplasm (compare a with b). Long simple arrows—host cell nuclei. Short simple arrows—parasite nuclei. Simple arrowheads—pigment granules. Other explanations are given in the text
Morphological features of gametocytes, which are used for identification of Haemoproteus species parasitizing Caprimulgiformes birds. Macrogametocytes (a, b) and microgametocyte (c) of Haemoproteus caprimulgi (a–c). Note the markedly pleomorphic form of fully grown gametocytes (b, c). Advanced growing gametocytes often do not adhere to the nuclei of infected erythrocytes (a). Long simple arrows—host cell nuclei. Short simple arrows—parasite nuclei. Simple arrowheads—pigment granules. Simple wide long arrows—unfilled space between growing gametocyte and the nucleus of infected erythrocyte. Other explanations are given in the text
Morphological features of gametocytes, which are used for identification of Haemoproteus species parasitizing Cariamiformes birds. Macrogametocyte (a) and microgametocyte (b) of H. pulcher (a, b). Note that advanced growing gametocytes often do not adhere to the nuclei of infected erythrocytes (b), and the infected erythrocytes nuclei assume roundish form (a, b). Long simple arrows—host cell nuclei. Short simple arrows—parasite nuclei. Simple arrowhead—pigment granules. Other explanations are given in the text
Morphological features of gametocytes, which are used for identification of Haemoproteus species parasitizing Cathartiformes birds. Macrogametocyte (a) and microgametocyte (b) of Haemoproteus catharti (a, b). Note that advanced growing gametocytes often do not adhere to erythrocyte nuclei (a). Pigment granules are of medium size and numerous (a, b). Long simple arrows—host cell nuclei. Short simple arrows—parasite nuclei. Simple arrowheads—pigment granules. Simple wide long arrows—unfilled space between growing gametocyte and nucleus of infected erythrocyte. Other explanations are given in the text
Morphological features of gametocytes, which are used for identification of Haemoproteus species parasitizing Charadriiformes birds. Macrogametocytes (a–f, h, j–m, o, p) and microgametocytes (g, i, n) of Haemoproteus rotator (a–c), H. scolopaci (d, e), H. jenniae (f, g), H. larae (h, i), H. abdusalomovi (j), H. burhini (k), H. skuae (l–n), H. contortus (o, p). Note that fully grown gametocytes of H. rotator markedly rotate the nuclei of infected erythrocytes (a–c). Nucleolus is readily visible in nuclei of H. scolopaci macrogametocytes (d, e). Fully grown gametocytes of H. jenniae contain predominantly roundish or slightly oval and of approximately uniform size and form pigment granules (f, g), which are different from the predominantly elongate rod-like pigment granules in gametocytes of H. larae (h, i). More or less evident unfilled spaces are present between circumnuclear macrogametocytes and nuclei of infected erythrocytes in H. skuae (m) and H. contortus (p). Images a, c, l–n, o are from the type material, which is fading, resulting in pale staining and poorly recognizable pigment granules and nuclei, however the overall form of the gametocytes is readily visible. Long simple arrows—host cell nuclei. Short simple arrows—parasite nuclei. Simple arrowheads—pigment granules. Simple wide short arrows—vacuoles. Simple wide long arrows—unfilled spaces between gametocytes and nuclei of infected erythrocytes. Other explanations are given in the text
Morphological features of gametocytes, which are used for identification of Haemoproteus species parasitizing Ciconiiformes birds. Macrogametocytes (a, b) of Haemoproteus crumenium (a) and H. ciconiae (b). Note that nucleus of H. crumenium macrogametocyte locate close the erythrocyte nucleus (a), but this is usually not a case in H. ciconiae (b). Long simple arrows—host cell nuclei. Short simple arrows—parasite nuclei. Simple arrowheads—pigment granules. Simple wide short arrows—vacuoles. Other explanations are given in the text
Morphological features of gametocytes, which are used for identification of Haemoproteus species parasitizing Coliiformes birds. Macrogametocytes of Haemoproteus undulatus (a, b). Note that growing gametocytes are usually appressed to the envelope of infected erythrocytes but do not touch the nuclei of erythrocytes along their entire margin (a). Form of advanced gametocytes is close to circumnuclear (b) or circumnuclear. Long simple arrows—host cell nuclei. Short simple arrows—parasite nuclei. Simple arrowheads—pigment granules. Simple wide long arrows—unfilled spaces between gametocytes and nuclei of infected erythrocytes. Other explanations are given in the text
Morphological features of fully grown (a–g, i, k, l) and young (h, j) gametocytes, which are used for identification of Haemoproteus species parasitizing Columbiformes birds. Macrogametocytes (a, d–k) and microgametocytes (b, c, l) of Haemoproteus columbae (a, b), H. turtur (c, d), H. palumbis (e), H. multivolutinus (f), H. multipigmentatus (g, h), H. paramultipigmentatus (i, j) and H. sacharovi (k, l). Note that big roundish volutin granules present in gametocytes of H. columbae (a, b), and the small pigment granules locate inside these volutin granules but usually are not present free in the cytoplasm (b). Numerous discrete roundish volutin granules present in gametocytes of H. turtur (c, d) and H. palumbis (e), and pigment granules are readily visible in the cytoplasm of both these species (d, e). Volutin overfills the cytoplasm in gametocytes in H. multivolutinus (f). Young gametocytes of H. multipigmentatus (h) and H. paramultipigmentatus (j) are markedly different in form. Gametocytes of H. sacharovi (k, l) are outwardly similar to gametocytes of Leucocytozoon parasites. Long simple arrows—host cell nuclei. Short simple arrows—parasite nuclei. Simple arrowheads—pigment granules. Triangle wide arrowheads—volutin. Other explanations are given in the text
Morphological features of gametocytes, which are used for identification of Haemoproteus species parasitizing Coraciiformes birds. Macrogametocytes (a, c, e, g–j, l, n, o, q, s) and microgametocytes (b, d, f, k, m, p, r, t) of Haemoproteus lairdi (a, b), H. enucleator (c, d), H. gavrilovi (e, f), H. fuscae (g–i), H. coraciae (j, k), H. eurystomae (l–n), H. manwelli (o, p), H. meropis (q, r) and H. halcyonis (s, t). Note the presence of rhabdosomal gametocytes in H. lairdi (b) and H. enucleator (c, d). One big circular vacuole is often present in the cytoplasm of H. gavrilovi macrogametocyte (e). Nucleus of H. halcyonis microgametocyte is condensed (t) and is similar in size to macrogametocyte nucleus (s), which is a rare feature in avian haemoproteids. Images c, g, h, l, m, s, t are from the type material, which is fading, resulting in pale staining and poorly recognizable pigment granules and nuclei, however the overall form of gametocytes is readily visible. Long simple arrows—host cell nuclei. Short simple arrows—parasite nuclei. Simple arrowheads—pigment granules. Simple wide short arrows—vacuoles. Triangle wide arrowheads—volutin granules. Other explanations are given in the text
Morphological features of gametocytes, which are used for identification of Haemoproteus species parasitizing Cuculiformes birds. Macrogametocytes (a–f) of Haemoproteus cuculis (a), H. centropi (b–e) and H. clamatori (f). Note that pigment granules in H. centropi gametocytes tend to aggregate into compact large masses (c) or loosely aggregated clumps (e). Long simple arrows—host cell nuclei. Short simple arrows—parasite nuclei. Simple arrowheads—pigment granules. Other explanations are given in the text
Morphological features of gametocytes, which are used for identification of Haemoproteus species parasitizing Falconiformes birds. Macrogametocytes (a–f) of Haemoproteus tinnunculi (a–c) and H. brachiatus (d–f). Note that growing gametocytes of H. brachiatus are highly irregular or amoeboid in outline (d, e). Advanced growing gametocytes of H. brachiatus often do not adhere to envelope of erythrocytes (d), which is not a case in H. tinnunculi (a). Long simple arrows—host cell nuclei. Short simple arrows—parasite nuclei. Simple arrowheads—pigment granules. Simple wide long arrows—a space between gametocyte and envelope of infected erythrocyte. Triangle wide arrows—ameboid outgrowths. Other explanations are given in the text
Morphological features of gametocytes, which are used for identification of Haemoproteus species parasitizing Galliformes birds. Macrogametocytes (a–h, j–l) and microgametocytes (i) of Haemoproteus lophortyx (a, b), H. stableri (c), H. mansoni (d), H. pratasi (e), H. ammoperdix (f), H. rileyi (g), H. cracidarum (h, i), H. paraortalidum (j) and H. ortalidum (k, l). Note that macrogametocytes of H. paraortalidum and H. ortalidum contain one large (bigger than 1 µm in diameter) circular vacuole (j–l). An unfilled space (a ‘cleft’) is present between the fully grown gametocytes and the nuclei of infected erythrocytes during development of H. lophortyx (a, b) and H. stableri (c). Vacuole-like unstained spaces (j) are present on both ends of H. paraortalidum macrogametocyte. Images a, k, l are from the type material, which is fading, resulting in pale staining and the poorly recognizable pigment granules and nuclei, however the overall form of the gametocytes is readily visible. Long simple arrows—host cell nuclei. Short simple arrows—parasite nuclei. Simple arrowheads—pigment granules. Simple wide long arrows—unfilled spaces between gametocytes and nuclei of infected erythrocytes. Simple wide short arrows—vacuoles. Simple wide arrowheads—unstained spaces on the ends of macrogametocyte. Other explanations are given in the text
Morphological features of gametocytes, which are used for identification of Haemoproteus species parasitizing Gruiformes birds. Macrogametocytes (a, c–h) and microgametocyte (b) of Haemoproteus balearicae (a, b), H. gallinulae (c, d), H. antigonis (e), H. porzanae (f–h). Note the slender form of H. balearicae gametocytes (a, b) and the presence of dumbbell-shaped growing gametocytes in H. porzanae (f, g). Long simple arrows—host cell nuclei. Short simple arrows—parasite nuclei. Simple arrowheads—pigment granules. Simple wide long arrows—spaces between gametocytes and envelope of infected erythrocytes. Other explanations are given in the text
Morphological features of gametocytes, which are used for identification of Haemoproteus species parasitizing Musophagiformes birds. Macrogametocytes (a-c) of Haemoproteus montezi (a, b) and H. minchini (c). Note that both ends of advanced H. montezi gametocytes usually are more or less narrowed in comparison to the widths of the gametocytes (b), but this is not a case in H. minchini gametocytes, which both ends usually are approximately similarly rounded (c). Images a, b are from the type material, which is fading, resulting in pale staining and the poorly recognizable pigment granules and nuclei, however the overall form of the gametocytes is readily visible. Long simple arrows—host cell nuclei. Short simple arrows—parasite nucleus. Simple arrowheads—pigment granules. Triangle wide arrowheads—volutin granules. Other explanations are given in the text
Morphological features of gametocytes, which are used for identification of Haemoproteus species parasitizing Otidiformes birds. Macrogametocytes of Haemoproteus telfordi (a, b). Note that the growing gametocytes are closely appressed to the erythrocyte envelope but usually do not touch the erythrocyte nuclei (a). Long simple arrow—host cell nucleus. Short simple arrows—parasite nuclei. Simple arrowheads—pigment granules. Simple wide long arrows—a space between gametocyte and nucleus of infected erythrocyte. Other explanations are given in the text
Morphological features of gametocytes, which are used for identification of Haemoproteus species parasitizing Pelecaniformes birds. Macrogametocytes (a, b, d–i) and microgametocytes (c) Haemoproteus plataleae (a–c), H. pelouroi (d–f) and H. herodiadis (g–i). Note the pleomorphic form of fully grown gametocytes of H. plataleae (a–c). Gametocyte outline is predominantly markedly irregular in H. pelouroi (d–f), but is smooth in H. herodiadis (g–i). Images d–f are from the type material, which is fading, resulting in pale staining and the poorly recognizable pigment granules and nuclei, however the overall form of the gametocytes is readily visible. Long simple arrows—host cell nuclei. Short simple arrows—parasite nuclei. Simple arrowheads—pigment granules. Other explanations are given in the text
Morphological features of gametocytes, which are used for identification of Haemoproteus species parasitizing Piciformes birds. Macrogametocytes (a–e, i–l, n, p) and microgametocytes (f–h, m, o) of Haemoproteus xantholaemae (a), H. cornuata (b), H. velans (c), H. homovelans (d), H. bennetti (e–g), H. bucconis (h), H. thereicerycis (i–k), H. bilobata (l, m), H. indicator (n, o) and H. borgesi (p). Note that H. velans gametocytes contain numerous prominent volutin granules and/or distinct volutin clumps (c), but this is not the case in H. homovelans (d). The advanced gametocytes of H. bennetti, H. bucconis and H. thereicerycis markedly displace nuclei of infected erythrocytes to the erythrocyte poles (f, i, k) and can enucleate the infected erythrocytes (e, g, h). Fully grown gametocytes of H. bilobata assume a unique dumbbell-like or bilobed form (l, m) and cause a readily visible flattening of nuclei of infected erythrocytes at the nuclei side, which is opposite to the gametocytes (l, m). Images f–h, j–m are from the type material, which is fading, resulting in pale staining and the poorly recognizable pigment granules and nuclei, however the overall form of the gametocytes is readily visible. Long simple arrows—host cell nuclei. Short simple arrows—parasite nuclei. Simple arrowheads—pigment granules. Simple wide long arrows—unfilled spaces between gametocytes and the envelope of infected erythrocytes or/and between gametocytes and nuclei of the erythrocytes. Triangle wide arrowhead—volutin granule. Other explanations are given in the text
Morphological features of fully grown gametocytes, which are used for identification of Haemoproteus species parasitizing Psittaciformes birds. Microgametocyte (a) and macrogametocytes (b–e) of Haemoproteus handai (a, b), H. homohandai (c) and H. psittaci (d, e). Note the markedly different form, size and number of pigment granules in gametocytes of H. handai (a), H. homohandai (c) and H. psittaci (e). Image b is from the type material, which is fading, resulting in pale staining and the poorly recognizable pigment granules and nuclei, however the overall form of the gametocyte is readily visible. Long simple arrows—host cell nuclei. Short simple arrows—parasite nuclei. Simple arrowheads—pigment granules. Other explanations are given in the text
Morphological features of gametocytes, which are used for identification of Haemoproteus species parasitizing Pterocliformes birds. Macrogametocytes of Haemoproteus krylovi (a) and H. pteroclis (b). Long simple arrows—host cell nuclei. Short simple arrow—parasite nucleus. Simple arrowheads—pigment granules. Other explanations are given in the text
Morphological features of gametocytes, which are used for identification of Haemoproteus species parasitizing Strigiformes birds. Macrogametocytes (a, d, e) and microgametocytes (b, c) of Haemoproteus syrnii (a–c) and H. noctuae (d, e). Note that volutin in H. syrnii gametocytes is arranged in compact roundish or circular granules (a, b), but this is not a case in H. noctuae (d, e). Long simple arrows—host cell nuclei. Short simple arrows—parasite nuclei. Simple arrowheads—pigment granules. Triangle wide arrowheads—volutin. Other explanations are given in the text
Morphological features of fully grown gametocytes, which are used for identification of Haemoproteus species parasitizing Suliformes birds. Macrogametocytes (a, c) and microgametocytes (b, d) of Haemoproteus iwa (a, b) and H. valkiunasi (c, d). Note that the number of pigment granules in fully grown H. iwa macrogametocytes (a) is at least twice that in fully grown microgametocytes (b), but this is not the case in H. valkiunasi (c, d). Long simple arrows—host cell nuclei. Short simple arrows—parasite nuclei. Simple arrowheads—pigment granules. Other explanations are given in the text
Morphological features of gametocytes, which are used for identification of Haemoproteus species parasitizing passeriform birds (suborder Tyranni) of the families Eurylaimidae, Furnariidae, Pittidae, Thamnophilidae, Tyrannidae. Macrogametocytes (a–c, e–i) and microgametocytes (d) of H. circumnuclearis (a, b), H. pittae (c, d), H. tyranni (e), H. furnarius (f, g), H. formicarius (h), H. eurylaimus (i). Note that attenuated (snake-like) advanced gametocytes (f) develop in H. furnarius. Image a, c are from the type material, which is fading, resulting in pale staining and the poorly recognizable pigment granules and nuclei, however the overall form of gametocytes is readily visible. Long simple arrows—host cell nuclei. Short simple arrows—parasite nuclei. Simple arrowheads—pigment granules. Simple wide long arrows—space between developing gametocyte and envelope of infected erythrocyte. Other explanations are given in the text
Morphological features of gametocytes, which are used for identification of Haemoproteus species parasitizing passeriform birds (suborder Passeri) of the families Meliphagidae, Oriolidae, Pachycephalidae, Vireonidae birds. Macrogametocytes of H. vireonis (a, b), H. ptilotis (c, d), H. pachycephalus (e, f) and H. orioli (g, h). Note that advanced growing gametocytes of H. pachycephalus (size greater than length of erythrocyte nuclei) do not touch the envelope of infected erythrocytes along their entire margin (e, f). Pigment granules are small in gametocytes of H. orioli, and nuclei assume terminal position in macrogametocytes of this parasite (g, h). Images a, c, e are from the type material, which is fading, resulting in pale staining (e) and the poorly recognizable nuclei (a, c, e) and pigment granules (e), however the overall form of the gametocytes is readily visible. Long simple arrows—host cell nuclei. Short simple arrows—parasite nuclei. Simple arrowheads—pigment granules. Simple wide long arrows—space between developing gametocyte and envelope of infected erythrocyte. Other explanations are given in the text
Morphological features of gametocytes, which are used for identification of Haemoproteus species parasitizing passeriform birds (suborder Passeri) of the families Aegithinidae, Artamidae, Malaconotidae, Vangidae birds. Macrogametocytes (a, b, d–f) and microgametocytes (c) of H. aegithinae (a–c), H. bukaka (d, e) and H. cublae (f). Note that advanced growing gametocytes of H. aegithinae are closely appressed to nuclei of infected erythrocytes but do not touch envelope of the erythrocytes along their entire margin (b, c). On the opposite, the advanced growing gametocytes of H. cublae do not touch the nuclei of infected erythrocytes along their entire margin but are closely appressed to erythrocyte envelope (f). Macrogametocytes of H. bukaka contain large vacuoles (d, e). Images b, c are from the type material, which is fading, resulting in pale staining and the poorly recognizable pigment granules and nuclei, however the overall form of gametocytes is readily visible. Long simple arrows—host cell nuclei. Short simple arrows—parasite nuclei. Simple arrowheads—pigment granules. Simple wide long arrows—space between developing gametocytes and envelope or nucleus of infected erythrocytes. Simple wide short arrows—vacuoles. Other explanations are given in the text
Morphological features of gametocytes, which are used for identification of Haemoproteus species parasitizing passeriform birds (suborder Passeri) of the families Corvidae, Dicruridae, Laniidae, Monarchidae birds. Macrogametocytes (a, b, e, g–j) and microgametocytes (c, d, f) of H. homopicae (a–c), H. picae (d), H. danilewskii (e, f), H. lanii (g), H. dicruri (h) and H. monarchus (i, j). Note that the cytoplasm is markedly vacuolated (c) in growing and fully grown microgametocytes of H. homopicae, but this is not a case (d) in microgametocytes of H. picae. Macrogametocyte nucleus is closely appressed to the nucleus of infected erythrocyte (g) in H. lanii. Growing gametocyte of H. monarchus is dumbbell-shaped and does not touch the envelope of erythrocyte along its entire margin (i). Image i is from the type material, which is fading, resulting in pale staining and the poorly recognizable pigment granules and nucleus, however the overall form of the gametocyte is readily visible. Long simple arrows—host cell nuclei. Short simple arrows—parasite nuclei. Simple arrowheads—pigment granules. Simple wide long arrows—spaces between gametocytes and envelope of infected erythrocytes. Simple wide short arrows—vacuoles. Other explanations are given in the text
Morphological features of gametocytes, which are used for identification of Haemoproteus species parasitizing passeriform birds (suborder Passeri) of the families Alaudidae, Cisticolidae, Melanocharitidae, Paridae birds. Macrogametocytes of H. wenyoni (a), H. majoris (b, c), H. alaudae (d), H. parus (e, f), H. calandrellae (g), H. nucleophilus (h, i). Note that the growing gametocyte of H. majoris assume dumbbell-like shape (c). Fully grown gametocytes of H. parus are roundish in form (e, f). Macrogametocyte nucleus assume median positions and adheres to the infected erythrocyte nucleus (g) in H. calandrellae. Fully grown gametocytes are closely appressed to the nuclei of erythrocytes but do not touch the envelope of the erythrocytes along their entire margin (h, i) in H. nucleophilus. Image h is from the type material, which is fading, resulting in pale staining and the poorly recognizable pigment granules and nucleus, however the overall form of the gametocyte is readily visible. Long simple arrows—host cell nuclei. Short simple arrows—parasite nuclei. Simple arrowheads—pigment granules. Simple wide long arrows—spaces between gametocytes and envelope of infected erythrocytes. Other explanations are given in the text
Morphological features of gametocytes, which are used for identification of Haemoproteus species parasitizing passeriform birds (suborder Passeri) of the families Acrocephalidae and Hirundinidae birds. Macrogametocytes (a, d, f–i, k, l) microgametocytes (b, c, e, j) of H. payevskyi (a, b), H. nucleocondensus (c, d), H. belopolskyi (e–g), H. parahirundinis (h), H. hirundinis (i, j) and H. stellaris (k, l). Note the markedly condensed nuclear material in fully grown microgametocytes of H. payevskyi (b) and H. nucleocondensus (c), resulting in similar size of the nuclei in microgametocytes (b, c) and macrogametocytes (a, d), a rare character in Haemoproteus parasites. Macrogametocytes nucleus is median in position (h) in H. parahirundinis, but this is not a case in H. hirundinis (i). Gigantic pigment granules (k, l) are present in fully grown gametocytes of H. stellaris; the granules might be arranged like stars (k, l). Image k is from the type material, which is fading, resulting in pale staining and the poorly recognizable pigment granules and nucleus, however the overall form of the gametocyte and pigment granules are readily visible. Long simple arrows—host cell nuclei. Short simple arrows—parasite nuclei. Simple arrowheads—pigment granules. Simple wide long arrows—spaces between gametocytes and envelope of infected erythrocytes. Other explanations are given in the text
Morphological features of gametocytes, which are used for identification of Haemoproteus species parasitizing passeriform birds (suborder Passeri) of the families Leiothrichidae, Phylloscopidae, Pycnonotidae, Sylviidae and Zosteropidae birds. Macrogametocytes (a–c, e–h, j–q, s, t, v), microgametocytes (d, r, u) and young gametocyte (i) of H. killangoi (a, b), H. parabelopolskyi (c–e), H. leiothrichus and H. homoleiothrichus (f, g), H. timalus (h, i), H. philippinensis (j), H. vacuolatus (k), H. palloris (l), H. homogeneae (m), H. otocompsae (n, o), H. sanguinis (p), H. pallidulus (q, r), H. homopalloris (s) and H. zosteropis (t-v). Note that dumbbell-shaped gametocyte does not touch envelope of infected erythrocyte along its entire margin (j) in H. philippinensis. The majority of advanced macrogametocytes contain a clear roundish discrete vacuole (k, l) in H. vacuolatus and H. palloris. The cytoplasm is homogenous in appearance (m) in macrogametocytes of H. homogeneae. Advanced growing macrogametocytes of H. otocompsae and H. sanguinis are closely appressed to the nuclei of infected erythrocytes but do not touch the envelope of the erythrocytes along their entire margin (n). The macrogametocyte cytoplasm is relatively pale-stained (q) and is similar to microgametocyte (r) based this character. Fully grown gametocytes of H. killangoi and H. zosteropis contain large size (greater than 1 µm) pigment granules (a, b, t–v). Images j, v are from the type material, which is fading, resulting in pale staining and the poorly recognizable pigment granules and nuclei, however the overall form of the gametocytes is readily visible. Long simple arrows—host cell nuclei. Short simple arrows—parasite nuclei. Simple arrowheads—pigment granules. Simple wide long arrows—spaces between gametocytes and envelope of infected erythrocytes. Simple wide short arrows—vacuoles. Other explanations are given in the text
Morphological features of gametocytes, which are used for identification of Haemoproteus species parasitizing passeriform birds (suborder Passeri) of the families Mimidae, Muscicapidae, Sittidae, Sturnidae and Turdidae birds. Macrogametocytes (a, d–f, h, j–m, o–r) and microgametocytes (b, c, g, i, n, s, t) of H. attenuatus (a, b), H. balmorali (c, d), H. pastoris (e), H. homominutus (f, g), H. kairullaevi (h, i), H. sittae (j), H. asymmetricus (k), H. fallisi (l), H. minutus (m, n), H. beckeri (o), H. neseri (p, q), H. nipponensis (r) and H. pallidus (s, t). Note that advanced growing microgametocytes of H. attenuatus are markedly narrow (attenuated) in width (b). Gametocytes of H. attenuatus and H. balmorali are overfilled with volutin granules (a–d). The asymmetrical position of advanced growing gametocytes in regard of erythrocyte nuclei (k) is a characteristic feature of H. asymmetricus. Advanced growing gametocytes of H. neseri (p, q) have well-pronounces dumbbell-like form (p, q). Fully grown gametocytes of H. pallidus are closely appressed to the nuclei of infected erythrocytes, but do not touch the erythrocyte envelope along their entire margin (s, t). Images l, p are from the type material, which is fading, resulting in pale staining and the poorly recognizable pigment granules and nucleus, however the overall form of the gametocytes is readily visible. Long simple arrows—host cell nuclei. Short simple arrows—parasite nuclei. Simple arrowheads—pigment granules. Simple wide long arrows—spaces between gametocytes and envelope of infected erythrocytes. Triangle wide arrowheads—volutin granules. Simple wide short arrow—vacuole. Other explanations are given in the text
Morphological features of gametocytes, which are used for identification of Haemoproteus species parasitizing passeriform birds (suborder Passeri) of the families Dicaeidae, Estrildidae, Fringillidae, Motacillidae, Nectariniidae, Passeridae and Ploceidae birds. Macrogametocytes (b, e–h, k–y, z–gg), microgametocytes (a, c, d), and young gametocytes (i, j) of H. nucleofascialis (a, b), H. micronuclearis (c), H. africanus (d, e), H. homobelopolskyi (f, g), H. passeris (h, i), H. fringillae (j, k), H. dolniki (l), H. queleae (m), H. dicaeus (n), H. tartakovskyi (o, p), H. anthi (q), H. orizivorae (r), H. globulosus (s, t), H. concavocentralis (u, v), H. uraeginthus (w, x), H. paranucleophilus (y), H. cyanomitrae (z), H. magnus (aa, bb), H. macropigmentatus (cc), H. motacillae (dd–ff) and H. bubalornis (gg). Note that fully grown microgametocytes of H. nucleofascialis and H. micronuclearis contain markedly compressed (not diffuse) nuclei (a, c). Nucleus of fully grown H. africanus macrogametocyte locates centrally (e). The rod-like (thin) pigment granules (f, g) are common in advanced and fully grown gametocytes of H. homobelopolskyi. During growth, the young gametocytes of H. passeris first adhere to envelope of erythrocytes and then grow towards the erythrocyte nuclei (i), but the opposite pattern of growth (j) is characteristic in H. fringillae. Growing advanced macrogametocyte of H. fringillae is dumbbell-shaped and assume a distinct linear form; the parasite deforms infected erythrocytes by causing the envelop protrusion, which is located in the non-invaded cytoplasmic region of the erythrocyte (k). A readily distinguishable space is present between the growing advanced gametocyte of H. concavocentralis and the nucleus of infected erythrocyte, resulting in the gametocyte concave form (u, v). Fully grown gametocytes of H. uraeginthus often assume rhabdosomal form (w, x) and enucleate infected erythrocytes (x). Fully grown gametocytes of H. paranucleophilus are closely appressed to the nuclei of infected erythrocytes but do not touch the envelope of the erythrocytes along their entire margin (y). Advanced growing gametocytes of H. cyanomitrae are closely appressed to the nuclei of infected erythrocytes but often do not touch the envelope of the erythrocytes along their entire margin (z); the similar feature is characteristic of H. sequeirae (not shown). Fully grown gametocytes of H. magnus, H. macropigmentatus, H. motacillae and H. bubalornis contain the large-size pigment granules (aa–cc, ff, gg). Images w, x are from the type material, which is fading, resulting in pale staining and the poorly recognizable pigment granules and nucleus, however the overall form of the gametocytes is readily visible. Long simple arrows—host cell nuclei. Short simple arrows—parasite nuclei. Simple arrowheads—pigment granules. Simple wide long arrows—spaces between gametocytes and structures of infected erythrocytes. Simple wide short arrows—vacuoles. Triangle long arrow—protrusion of the erythrocyte envelope. Other explanations are given in the text
Morphological features of gametocytes, which are used for identification of Haemoproteus species parasitizing passeriform birds (suborder Passeri) of the families Emberizidae, Icteridae, Parulidae, Passerellidae and Thraupidae birds. Macrogametocytes of H. erythrogravidus (a), H. coatneyi (b), H. nucleocentralis (c) and H. quiscalus (d). Note that the fully grown gametocyte of H. erythrogravidus markedly deform the host cell by causing a balloon-like protrusion (a), which is located in the non-invaded cytoplasmic region of the erythrocytes. The nuclei assume predominantly central position (c) in fully grown macrogametocytes of H. nucleocentralis. Advanced dumbbell-shaped macrogametocytes, which do not touch envelope of erythrocytes along their entire margin (d) is a distinctive character of H. quiscalus. Image d is from the type material, which is fading, resulting in pale staining and the poorly recognizable pigment granules and nucleus, however the overall form of the gametocyte is readily visible. Long simple arrows—host cell nuclei. Short simple arrows—parasite nuclei. Simple arrowheads—pigment granules. Simple wide long arrows—spaces between gametocytes and the envelop of erythrocytes. Triangle long arrow—protrusion of the erythrocyte envelope. Other explanations are given in the text
Third, the morphometric characters of Haemoproteus gametocytes and their host cells can be similar and might overlap in some parasite species. Thus, they should be carefully considered during parasite identification and description. It is important to note that morphometric characters are functions of gametocyte size (age). In other words, the length and shape of gametocytes as well as number of pigment granules of the same parasite species can change markedly as the gametocytes grow and mature. As a result, the morphometric data are considered taxonomically valuable only if the gametocytes measurements are standardized and accessed only in fully grown gametocytes, which selection requires some experience and good quality preparations. Minor morphometric differences might be a result of incorrect measurements and are usually of low taxonomic value [3]. Furthermore, it is important to note that measurement of parasites is time consuming and requires subsequent statistical evaluation. This is why the use of morphometric characters was minimized in the keys. However, some readily distinguishable morphometric data (the nuclear displacement ratio, number of pigment granules and their size, etc.) were essential for some species identifications, but they were provided only for fully grown gametocytes in the keys. Measurements of the growing gametocytes will distort taxonomic values of the morphometric characters and might abate their applicability during species identifications and thus should be discouraged.
Fourth, the co-infections of Haemoproteus and Plasmodium species often occur in naturally infected birds and require some experience to distinguish between co-existing parasites [35, 38, 39, 182]. The co-infections of several morphologically similar species belonging to the same genus might be particularly difficult to distinguish; such co-infections also often are hardly distinguishable by commonly used PCR-based methods, which apply general primers for the parasite detection [38, 40]. However, the co-infections of many Haemoproteus and other haemosporidian species belonging to same genus are readily distinguishable using morphological characters of blood stages (for example see Fig. 37a, b).
Examples of co-infections of two different species of Haemoproteus in same blood films. Co-infection of Haemoproteus fringillae (a, on the left) and Haemoproteus magnus (a, on the right) from the blood of common chaffinch Fringilla coelebs. Co-infection of Haemoproteus pallidus (b, on the left) and Haemoproteus balmorali (b, on the right) from the blood of European pied flycatcher Ficedula hypoleuca. Macrogametocytes of these species were shown. Note that H. fringillae and H. magnus can be readily distinguished due to different form and size of pigment granules (a). The cytoplasm of H. balmorali macrogametocyte is densely stained and contains numerous volutin granules, which are not a case in H. pallidus (b). Long simple arrows—host cell nuclei. Short simple arrows—parasite nuclei. Simple arrowheads—pigment granules. Other explanations are given in the text
Fifth, the good quality of blood films is essential for the visualization of gametocyte morphological characters, which are needed for identification (Fig. 38a–d), but this is hardly possible in thick blood films (Fig. 38h) or in preparations affected by incorrect procedures of their drying, fixation, staining or storage [188]. Insufficient quality of preparations (Fig. 37e–h) is a significant obstacle in haemosporidian parasite species identification and description. It is essential to master methods of blood film preparation and storage before sample collection. These procedures are simple, relatively cheap and can be accomplished in any laboratory by careful application of well-described routine protocols [1, 3, 188, 189].
Gametocytes of Haemoproteus parasites as they are seen in good-quality (a-d) and bad-quality (e–h) blood films. The gametocyte structures (nuclei, pigment granules, volutin granules and position of the parasites in erythrocytes) are readily visible in good-quality blood films (a–d), but are hardly visible in preparations affected by incorrect procedures of staining (e), fixation (f, g) and blood film preparation (h). Note thick blood film (h), bad staining (e–h) and fixation (f, g), resulting in hardly visible parasite structures (e, g, h), destroyed host cell nuclei (g) and envelops (f). Bad-quality preparations are useless for haemosporidian species identification. Long simple arrows—host cell nuclei. Short simple arrows—parasite nuclei. Simple arrowheads—pigment granules. Simple wide short arrow—vacuole. Triangle wide arrowheads—volutin granules. Simple wide long arrows—Haemoproteus sp. parasites as they seen in bad-quality blood films. Triangle wide long arrows—artefacts of fixation, resulting in destroyed erythrocyte nuclei. Other explanations are given in the text
Sixth, the mature gametocytes of Haemoproteus parasites change morphology (round up and tend to escape from host cells) within one minute after the exposure to air [3, 148, 172]. This process naturally occurs in vectors before the exflagellation and gametogenesis [1,2,3, 7]. The form of gametocytes and their position in erythrocytes changes dramatically during this process, which readily occurs in vitro, including slowly drying blood films. As a result, such material can present distorted parasites and host cells and be unsuitable for species identification, and thus should be eliminated from the taxonomic work. The use of fans is recommended to quicken blood film drying, particularly during fieldwork in humid environments.
Molecular markers (barcodes) were developed and can be used for the detection and identification of approximately 42% of the named species (Table 36). Partial sequences of the mitochondrial cytochrome b gene (cytb) are easy to obtain using blood samples; they are known for many Haemoproteus species, and consequently are convenient and often used for haemoproteid and other haemosporidian parasite species delimitation (barcoding) [38]. However, most detected Haemoproteus parasite lineages remain non-characterized at the parasite species levels. Molecular characterization is best developed for the parasites of passeriform birds breeding and migrating in Europe, and it is weakest for the organisms inhabiting tropical bird species, which are more difficult to access for blood sampling [7, 13]. Parasites of most non-passeriform birds remained non-characterized molecularly for the same reason. This is a prominent obstacle in developing generalizations in biodiversity research of Haemoproteus and other haemosporidian parasites. It is important to note that some parasite molecular characterizations were based on incorrect species identifications and thus should be treated with caution. Several such cases were reported in Table 36. The development of molecular markers for the diagnosis of agents of haemoproteosis and other haemosporidioses is an important undertaking of current parasitology studies.
Approximately 177 described species of avian Haemoproteus currently can be distinguished using morphological characters of their blood stages (Tables 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, 30, 31, 32, 33, 34, 35). The molecular data supported the validity of most comprehensive morphological species descriptions, which were based on a package of distinct morphological characters (Table 36).
The application of molecular diagnostic tools provides new opportunities to distinguish haemosporidian parasites using their unique DNA sequences [38]. This stimulates biodiversity studies of wildlife haemosporidians, particularly because the molecular markers make the research readily repeatable due to DNA sequence information. Additionally, parasite species detection becomes possible at all stages of the life cycle, including the relatively insubstantially studied sporogonic and tissue stages, which are still difficult to distinguish to species and even the subgeneric or generic levels using morphological characters. Due to the easy accessibility of peripheral blood of birds, the gametocytes of Haemoproteus parasites likely will remain the main stage used for parasite morphological identification in the nearest future. However, it is worth noting that haemosporidian species are morphologically remarkably diverse, and often are readily distinguishable at other stages of their life cycle, including gametes (Fig. 39a, b), zygotes (Fig. 39c, d), ookinetes (Fig. 39e–h), oocysts (Fig. 39i, j), sporozoites (Fig. 39k, l) and exoerythrocytic meronts (Fig. 39m–p). Morphological characters of these life cycle stages can be applied in species taxonomy, and this might reduce the number of so-called morphologically cryptic haemosporidian parasites, which are similar or non-distinguishable at the gametocyte stage [3, 7, 165]. It is interesting to note in this regard that many species of Haemoproteus can be distinguished due to the markedly different size of their mature ookinetes (Fig. 39g, h) and the rate of their development, which also can be considered taxonomic characters [3, 149]. For example, the fully grown ookinetes of Haemoproteus minutus, H. pallidus and H. asymmetricus usually do not exceed 12 µm on average in length, and they complete development rapidly (within 1 h at 18–20 °C) in vitro, but the ookinetes of H. tartakovskyi, H. fringillae and H. balmorali are at least 1/3 bigger and develop 3-times more slowly at the same conditions [3, 63].
Examples of marked variation in morphological characters of different Haemoproteus species during sporogony (a–l) and exo-erythrocytic development (m–p). Microgametes (a, b), zygotes (c, d), developing ookinetes (e, f), fully grown ookinetes (g, h), oocysts (i, j), sporozoites (k, l), exo-erythrocytic meront (o) and megalomeronts (m, n, p) of H. minutus (a, g), H. tartakovskyi (b, c, h, k), H. fringillae (d, i), H. palumbis (j, l), H. majoris (m), H. mansoni (n), H. attenuatus (o) and H. passeris (p). Note that these Haemoproteus species are readily different not only on gametocyte stages (see Figs. 13, 17, 31, 34, 35), but also due to the following characters of their sporogonic and exo-erythrocytic stages: length of microgametes (a, b), mode of the cytoplasm vacuolization in zygotes (c, d), mode of elongation during initial stage of ookinete development (e, f), size and form of fully grown ookinetes (g, h), oocyst structure (i, j), size and form of sporozoites (k, l), form of exo-erythrocytic stages (m–p). Microgametes (a, b), ookinetes (g, h), sporozoites (k, l), oocysts (i, j) and exo-erythrocytic meronts (m–p) are clearly different in size. A large conspicuous vacuole (d) is present in zygote of H. fringillae, which is not a case in zygote of H. tartakovskyi (c). A long outgrowth appears in apical end of developing H. tartakovskyi ookinete (f), which is not a case in H. minutus ookinete (e). Developing oocyst of H. palumbis (j) contains numerous germinative centres (j), which is not a case in small oocyst of H. fringillae (i). One end of H. palumbis sporozoite is rounded (l), which is not the case in H. tartakovskyi sporozoite (k). Morphology of exo-erythrocytic meronts is markedly variable in different species of Haemoproteus (m–p). These examples show that different Haemoproteus and other haemosporidian species can be distinguished due to many characters of sporogonic and exo-erythrocytic stages, which remain unexplored in taxonomy. Long simple arrows—capsular-like wall of megalomeronts. Short simple arrows—parasite nuclei. Simple arrowheads—pigment granules. Simple wide arrowheads—apical end of ookinetes. Simple wide short arrows—vacuoles. Simple small arrowhead—merozoites. Simple wide long arrows—megalomeront cytomeres. Triangle wide short arrows—oocysts. Scale bars 10 µm (a–i, k, l), 50 µm (j, m–p). Other explanations are given in the text
Numerous morphologically indistinguishable Haemoproteus species have been described and are currently considered as synonyms of valid parasite names [3, 31, 32]. The lists of synonymous Haemoproteus species names as well as the names of unknown taxonomic position (incertae sedis), the names of species of doubtful identity (species inquirenda), were justified and are available in [3]; these data did not change significantly and were not repeated in this review. Most of these parasite descriptions were incomplete and came from old studies published in the twentieth century. Some new names of doubtful identity were reported in keys. Because of the high genetic diversity of avian Haemoproteus parasites and their relatively high vertebrate host specificity, some of the available synonymous names might be considered valid in the future. In other words, such names might be a reserve for future nomenclature studies [198]. Further morphological research combined with molecular parasite characterization are needed to prove the validity of available synonyms [3]. For taxonomic clearance, invalid species names (nomen nudum) of haemosporidian parasites should be excluded from taxonomic use [198, 199]. The Haemoproteus parasites belonging to categories of incertae sedis and species inquirenda [3, 31, 32] will require further investigation before their final taxonomic status is determined.
It should be mentioned that GenBank contains DNA sequences of haemosporidian parasites, where the molecular characterization was developed incorrectly or might be questionable [200, 201]. Parasite species identification and molecular characterization is the responsibility of researchers who deposit the sequence information in GenBank. With regard to Haemoproteus species, some such insufficiently dependable molecular characterizations are mentioned in the footnotes of Table 36. Selection of the sequences for phylogenetic analysis and related molecular research requires targeting efforts and the MalAvi database is a helpful resource of information for this purpose [13].
Conclusion
Haemoproteus is a sister genus to malaria parasites of genus Plasmodium. The parasites of both genera are common in birds globally, however, Haemoproteus species remain neglected. This is unfortunate because knowledge about close relatives of malaria pathogens is essential for understanding the evolutionary history of malaria and also the entire group of haemosporidian infections, which are flourishing in wildlife. Recent histopathological findings show that Haemoproteus parasites can damage bird organs during exo-erythrocytic development and thus can contribute to the overall bird health. However, these avian pathogens remain insufficiently studied partly due to difficulties in their species identification, which is an obstacle for comprehensive biodiversity research. This study developed illustrated dichotomous keys for the identification of described avian Haemoproteus species allowing their recognition using morphological features of blood stages (gametocytes). The most taxonomically valuable morphological characters of gametocytes and their host cells were systemized and used in the keys. In all, 177 species of Haemoproteus species parasitizing birds can be identified using the morphological characters of blood stages. The DNA sequences (taxonomic barcodes) are available for approximately 42% of them, and can simplify the identification of these parasites. The easy-to-use keys should be helpful not only for identification of described parasites, but also for distinguishing new pathogens, which still need discovery and description. It worth noting that species of Haemoproteus are remarkably morphologically diverse not only on gametocyte stage, but also on the exo-erythrocytic and vector stages, which remain insufficiently investigated and remain as unexplored reserves for future taxonomic and biodiversity studies. The majority of described Haemoproteus parasites also remain non-characterized using molecular markers; their development is an important task for current haemosporidian parasite researchers. This is particularly true because such markers are invaluable for haemosporidian parasite species identification in the sporogonic and tissue stages, which remain unassessed in most of these wildlife pathogens.
Availability of data and materials
All data generated during this study are included in this published article.
References
Garnham PCC. Malaria parasites and other Haemosporidia. Oxford: Blackwell; 1966.
Atkinson CT, Thomas NJ, Hunter DB. Parasitic diseases of wild birds. Oxford: Wiley-Blackwell; 2008.
Valkiūnas G. Avian malaria parasites and other Haemosporidia. Boca Raton: CRC; 2005.
Lainson R. Atlas of protozoan parasites of the Amazonian fauna of Brazil. Haemosporida of reptiles, vol. 1. Ananindeua: Instituto Evandro Chagas; 2012.
Santiago-Alarcon D, Marzal A. Avian malaria and related parasites in the tropics: ecology, evolution and systematics. Cham: Springer; 2020.
Telford SR. The hemoparasites of the reptilian. Boca Raton: CRC; 2009.
Valkiūnas G, Atkinson CT. Introduction to life cycles, taxonomy, distribution and basic research techniques. In: Santiago-Alarcon D, Marzal A, editors. Avian malaria and related parasites in the tropics: ecology, evolution and systematics. Berlin: Springer; 2020. p. 45–80.
Bensch S, Canbäck B, DeBarry JD, Johansson T, Hellgren O, Kissinger JC, et al. The genome of Haemoproteus tartakovskyi and its relationship to human malaria parasites. Genome Biol Evol. 2016;8:1361–73.
Pacheco MA, Matta NE, Valkiūnas G, Parker PG, Mello B, Stanley CE Jr, et al. Mode and rate of evolution of Haemosporidian mitochondrial genomes: timing the radiation of avian parasites. Mol Biol Evol. 2018;35:383–403.
Venugopal K, Hentzschel F, Valkiūnas G, Marti M. Plasmodium asexual growth and sexual development in the haematopoietic niche of the host. Nat Rev Microbiol. 2020;18:177–89.
Toscani Field J, Weinberg J, Bensch S, Matta NE, Valkiūnas G, Sehgal RNM. Delineation of the genera Haemoproteus and Plasmodium using RNA-Seq and multi-gene phylogenetics. J Mol Evol. 2018;86:646–54.
Galen SC, Borner J, Martinsen ES, Schaer J, Austin CC, West CJ, et al. The polyphyly of Plasmodium: comprehensive phylogenetic analyses of the malaria parasites (order Haemosporida) reveal widespread taxonomic conflict. R Soc Open Sci. 2018;5: 171780.
Bensch S, Hellgren O, Pérez-Tris J. MalAvi: a public database of malaria parasites and related haemosporidians in avian hosts based on mitochondrial cytochrome b lineages. Mol Ecol Resour. 2009;9:1353–8.
Clark NJ, Clegg SM, Lima MR. A review of global diversity in avian haemosporidians (Plasmodium and Haemoproteus: Haemosporida): new insights from molecular data. Int J Parasitol. 2014;44:329–38.
Schaer J, Reeder DM, Vodzak ME, Olival KJ, Weber N, Mayer F, et al. Nycteria parasites of Afrotropical insectivorous bats. Int J Parasitol. 2015;45:375–84.
Bukauskaitė D, Iezhova TA, Ilgūnas M, Valkiūnas G. High susceptibility of the laboratory-reared biting midges Culicoides nubeculosus to Haemoproteus infections, with review on Culicoides species that transmit avian haemoproteids. Parasitology. 2018;4:1–9.
Outlaw DC, Ricklefs RE. Rerooting the evolutionary tree of malaria parasites. Proc Natl Acad Sci USA. 2011;108:13183–7.
Ricklefs RE, Outlaw DC. A molecular clock for malaria parasites. Science. 2010;329:226–9.
Bensch S, Hellgren O, Križanauskienė A, Palinauskas V, Valkiūnas G, Outlaw D, et al. How can we determine the molecular clock of malaria parasites? Trends Parasitol. 2013;29:363–9.
Bennett GF, Peirce MA, Ashford RW. Avian haematozoa: mortality and pathogenicity. J Nat Hist. 1993;27:993–1001.
Valkiūnas G, Iezhova TA. Detrimental effects of Haemoproteus infections on the survival of biting midge Culicoides impunctatus (Diptera: Ceratopogonidae). J Parasitol. 2004;90:194–6.
Valkiūnas G, Kazlauskienė R, Bernotienė R, Bukauskaite D, Palinauskas V, Iezhova TA. Haemoproteus infections (Haemosporida, Haemoproteidae) kill bird-biting mosquitoes. Parasitol Res. 2014;113:1011–8.
Bukauskaite D, Bernotienė R, Iezhova TA, Valkiūnas G. Mechanisms of mortality in Culicoides biting midges due to Haemoproteus infection. Parasitology. 2016;143:1748–54.
Valkiūnas G, Iezhova TA. Exo-erythrocytic development of avian malaria and related haemosporidian parasites. Malar J. 2017;16:101.
Duc M, Ilgūnas M, Kubiliūnaitė M, Valkiūnas G. First report of Haemoproteus (Haemosporida, Haemoproteidae) megalomeronts in the brain of an avian host, with description of megalomerogony of Haemoproteus pastoris, the blood parasite of the Common starling. Animals. 2021;10:2824.
Atkinson CT, Forrester DJ, Greiner EC. Pathogenicity of Haemoproteus meleagridis (Haemosporina: Haemoproteidae) in experimentally infected domestic turkeys. J Parasitol. 1988;74:228.
Himmel T, Harl J, Kübber-Heiss A, Konicek C, Fernández N, Juan-Sallés C, et al. Molecular probes for the identification of avian Haemoproteus and Leucocytozoon parasites in tissue sections by chromogenic in situ hybridization. Parasit Vectors. 2019;12:282.
Ilgūnas M, Chagas CRF, Bukauskaitė D, Bernotienė R, Iezhova T, Valkiūnas G. The life-cycle of the avian haemosporidian parasite Haemoproteus majoris, with emphasis on the exoerythrocytic and sporogonic development. Parasit Vectors. 2019;12:516.
Yoshimoto M, Ozawa K, Kondo H, Echigoya Y, Shibuya H, Sato Y, et al. A fatal case of a captive snowy owl (Bubo scandiacus) with Haemoproteus infection in Japan. Parasitol Res. 2021;120:277–88.
Kelly EJ, Baldwin TJ, Frame DD, Childress AL, Wellehan JFX. Haemoproteus (Parahaemoproteus) spp. in captive-bred bobwhite quail (Colinus virginianus) in southern Utah, USA. J Wildl Dis. 2018;54:726–33.
Bennett GF, Whiteway M, Woodworth-Lynas C. A host-parasite catalogue of the avian haematozoa. Mem Univ Nfld Occ Pap Biol. 1982;5:1–243.
Bishop MA, Bennett GF. Host-parasite catalogue of the avian haematozoa: supplement 1, and Bibliography of the avian blood-inhabiting haematozoa. Mem Univ Nfld Occ Pap Biol. 1992;15:1–244.
Greiner EC, Bennett GF, White EM, Coombs RF. Distribution of the avian hematozoa of north America. Can J Zool. 1975;53:1762–87.
McClure HE. Migration and survival of the birds of Asia. Bangkok; 1974.
Valkiūnas G, Bensch S, Iezhova TA, Križanauskienė A, Hellgren O, Bolshakov CV. Nested cytochrome b polymerase chain reaction diagnostics underestimate mixed infections of avian blood haemosporidian parasites: microscopy is still essential. J Parasitol. 2006;92:418–22.
Lotta IA, Valkiūnas G, Pacheco MA, Escalante AA, Hernández SR, Matta NE. Disentangling Leucocytozoon parasite diversity in the neotropics: descriptions of two new species and shortcomings of molecular diagnostics for leucocytozoids. Int J Parasitol Parasites Wildl. 2019;9:159–73.
Braga EM, Silveira P, Belo NO, Valkiūnas G. Recent advances in the study of avian malariae: an overview with an emphasis on the distribution of Plasmodium spp. in Brazil. Mem Inst Oswaldo Cruz. 2011;1:3–11.
Bensch S, Hellgren O. The use of molecular methods in studies of avian haemosporidians. In: Santiago-Alarcon D, Marzal A, editors. Avian malaria and related parasites in the tropics: ecology, evolution and systematics. Berlin: Springer; 2020. p. 113–36.
Ciloglu A, Ellis VA, Bernotienė R, Valkiūnas G, Bensch S. A new one-step multiplex PCR assay for simultaneous detection and identification of avian haemosporidian parasites. Parasitol Res. 2019;118:191–201.
Valkiūnas G, Duc M, Iezhova TA. Increase of avian Plasmodium circumflexum prevalence, but not of other malaria parasites and related haemosporidians in northern Europe during the past 40 years. Malar J. 2022;21:105.
Oliveros CH, Field DJ, Ksepka DT, Barker FK, Aleixo A, Andersen MJ, et al. Earth history and the passerine superradiation. Proc Natl Acad Sci USA. 2019;116:7916–25.
Kuhl H, Frankl-Vilches C, Bakker A, Mayr G, Nikolaus G, Boerno ST, et al. An unbiased molecular approach using 3’UTRs resolves the avian family-level tree of life. Mol Biol Evol. 2021;38:108–27.
Križanauskienė A, Hellgren O, Kosarev V, Sokolov L, Bensch S, Valkiūnas G. Variation in host specificity between species of avian hemosporidian parasites: evidence from parasite morphology and cytochrome b gene sequences. J Parasitol. 2006;92:1319–24.
Valkiūnas G, Ashford RW. Natural host range is not a valid taxonomic character. Trends Parasitol. 2002;18:528–9.
Križanauskienė A, Iezhova TA, Sehgal RNM, Carlson JS, Palinauskas V, Bensch S, et al. Molecular characterization of Haemoproteus sacharovi (Haemosporida, Haemoproteidae), a common parasite of columbiform birds, with remarks on classification of haemoproteids of doves and pigeons. Zootaxa. 2013;3613:085–94.
Merino S, Hennicke J, Martínez J, Ludynia K, Torres R, Work TM, et al. Infection by Haemoproteus parasites in four species of frigatebirds and the description of a new species of Haemoproteus (Haemosporida: Haemoproteidae). J Parasitol. 2012;98:388–97.
Chagas CRF, Harl J, Preikša V, Bukauskaitė D, Ilgūnas M, Weissenböck H, et al. Lankesterella (Apicomplexa, Lankesterellidae) blood parasites of passeriform birds: prevalence, molecular and morphological characterization, with notes on sporozoite persistence in vivo and development in vitro. Animals. 2021;11:1451.
Valkiūnas G, Mobley K, Iezhova TA. Hepatozoon ellisgreineri n. sp. (Hepatozoidae): description of the first avian apicomplexan blood parasite inhabiting granulocytes. Parasitol Res. 2016;115:609–13.
Bennett GF, Peirce MA. Morphological form in the avian Haemoproteidae and an annotated checklist of the genus Haemoproteus Kruse, 1890. J Nat Hist. 1988;22:1683–96.
Greiner EC, Mundy PJ. Hematozoa from Southern African vultures, with a description of Haemoproteus janovyi sp. N. J Parasitol. 1979;651:147–53.
de Mello IF. New haemoproteids of some Indian birds. Proc Indian Acad Sci. 1935;2:469–75.
Peirce MA, Bennett GF, Bishop M. The haemoproteids of the avian order Falconiformes. J Nat Hist. 1990;24:1091–100.
Wingstrand KG. On some haematozoa of Swedish birds with remarks on the schizogony of Leucocytozoon sakharoffi. Kungl Sven Vetenskapsakad Handl. 1947;24:1–31.
Peirce MA, Marquiss M. Haematozoa of British birds. VII. Haematozoa of raptors in Scotland with a description of Haemoproteus nisi sp. nov. from the sparrowhawk (Accipiter nisus). J Nat Hist. 1983;1:813–21.
Matta NE, Pacheco MA, Escalante AA, Valkiūnas G, Ayerbe-Quiñones F, Acevedo-Cendales LD. Description and molecular characterization of Haemoproteus macrovacuolatus n. sp. (Haemosporida, Haemoproteidae), a morphologically unique blood parasite of black-bellied whistling duck (Dendrocygna autumnalis) from South America. Parasitol Res. 2014;113:2991–3000.
Bennett GF, Turner B, Whiteway M. Avian Haemoproteidae. 18. Haemoproteus greineri, a new species of haemoproteids from the waterfowl family Anatidae. Can J Zool. 1984;62:2290–2.
Williams NA, Bennett GF. Avian Haemoproteidae. 13. The haemoproteids of the ducks and geese (Anatidae). Can J Zool. 1980;58:88–93.
Johnston HT, Cleland JB. Notes on some parasitic Protozoa. Proc Linn Soc NSW. 1909;34:501–13.
Coatney GR, West E. Some blood parasites from Nebraska birds—II. Am Midl Nat. 1938;19:601–12.
White EM, Bennett GF, Williams NA. Avian Haemoproteidae. 11. The haemoproteids of the hummingbird family Trochilidae. Can J Zool. 1979;57:908–13.
Moens MJ, Valkiūnas G, Paca A, Bonaccorso E, Aguirre N, Pérez-Tris J. Parasite specialization in a unique habitat: hummingbirds as reservoirs of generalist blood parasites of Andean birds. J Anim Ecol. 2016;85:1234–45.
González AD, Lotta IA, García LF, Moncada LI, Matta NM. Avian haemosporidians from Neotropical highlands: evidence from morphological and molecular data. Parasitol Int. 2015;64:48–59.
Valkiūnas G, Ilgūnas M, Bukauskaitė D, Duc M, Iezhova TA. Description of Haemoproteus asymmetricus n. sp. (Haemoproteidae), with remarks on predictability of the DNA haplotype networks in haemosporidian parasite taxonomy research. Acta Trop. 2021;218:105905.
Bennett GF, Caines JR, Whiteway MA. Avian Haemoproteidae. 20. The haemoproteids of the avian families Apodidae (swifts), Bucconidae (puffbirds), and Indicatoridae (honeyguides). Can J Zool. 1986;64:766–70.
Bennett GF, Earlé RA, Squires-Parsons D. Additional new species of Haemoproteus, Hepatozoon and Leucocytozoon from South African birds. S Afr J Wildl Res. 1995;25:1–7.
Williams NA, Bennett GF, Mahrt JL. Avian Haemoproteidae. 6. Description of Haemoproteus caprimulgi sp. nov., and a review of the haemoproteids of the family Caprimulgidae. Can J Zool. 1975;53:916–9.
Vanstreels RET, Anjos CC, Leandro HJ, Carvalho AM, Santos AP, Egert L, et al. A new haemosporidian parasite from the Red-legged Seriema Cariama cristata (Cariamiformes, Cariamidae). Int J Parasitol Parasites Wildl. 2022;28(18):12–9.
Greiner EC, Fedynich AM, Webb SL, DeVault TL, Rhodes OE. Hematozoa and a new haemoproteid species from Cathartidae (New World vulture) in South Carolina. J Parasitol. 2011;97:1137–9.
Yabsley MJ, Vanstreels RET, Martinsen ES, Wickson AG, Holland AE, Hernandez SM, et al. Parasitaemia data and molecular characterization of Haemoproteus catharti from New World vultures (Cathartidae) reveals a novel clade of Haemosporida. Malar J. 2018;8(17):2.
Bennett GF. Avian Haemoproteidae. 10. The haemoproteids of the avian family Scolopacidae. Can J Zool. 1979;57:901–7.
Galli-Valerio B. Notes parasitologie. Zentralbl Bakteriol Parasitenkd Infektionskr Hyg I Abt Orig. 1929;112:54–9.
Levin II, Valkiūnas G, Iezhova TA, O’brien SL, Parker PG. Novel Haemoproteus species (Haemosporida: Haemoproteidae) from the swallow-tailed gull (Lariidae), with remarks on the host range of hippoboscid-transmitted avian hemoproteids. J Parasitol. 2012;98:847–54.
Yakunin MP. Blood parasites of the wild birds of South-East Kazakhstan (in Russian). Trudi Inst Zool AN KazSSR. 1972;33:69–79.
Inumaru M, Aratani S, Shimizu M, Yamamoto M, Sato Y, Murata K, Valkiūnas G. Penguins are competent hosts of Haemoproteus parasites: the first detection of gametocytes, with molecular characterization of Haemoproteus larae. Parasit Vectors. 2020;13:307.
Subkhonov M. New species of blood parasites (Haemosporidia, Sporozoa) from birds of Tadzhikistan (in Russian). Parazitologiya. 1980;14:45–7.
Nola J, Parsons NL, Peirce MA, Strauss V. New species of haematozoa in Phalacrocoracidae and Stercorariidae in South Africa. Ostrich. 2010;81:103–8.
Peirce MA, Cooper JE. Haematozoa of East African birds. IV. Redescription of Haemoproteus crumenium, a parasite of the marabou stork. E Afr Wildl J. 1977;15:169–72.
Valkiūnas G, Ilgūnas M, Bukauskaitė D, Iezhova TA. Description of Haemoproteus ciconiae sp. nov. (Haemoproteidae, Haemosporida) from the white stork Ciconia ciconia, with remarks on insensitivity of established polymerase chain reaction assays to detect this infection. Parasitol Res. 2016;115:2609–16.
Bennett GF, Earlé RA. New species of Haemoproteus, Hepatozoon and Leucocytozoon from South African birds. S-Afr Tydskr Natuurnav. 1992;22:114–8.
Bennett GF, Peirce MA. The haemoproteid parasites of the pigeons and doves (family Columbidae). J Nat Hist. 1990;24:311–25.
Coral AA, Valkiūnas G, González AD, Matta NE. In vitro development of Haemoproteus columbae (Haemosporida: Haemoproteidae), with perspectives for genomic studies of avian haemosporidian parasites. Exp Parasitol. 2015;157:163–9.
Valkiūnas G, Iezhova TA, Evans E, Carlson JS, Martínez-Gómez JE, Sehgal RNM. Two new Haemoproteus species (Haemosporida: Haemoproteidae) from columbiform birds. J Parasitol. 2013;99:513–21.
Valkiūnas G, Santiago-Alarcon D, Levin II, Iezhova TA, Parker PG. A new Haemoproteus species (Haemosporida: Haemoproteidae) from the indemic Galapagos dove Zenaida galapagoensis, with remarks on the parasite distribution, vectors, and molecular diagnostics. J Parasitol. 2010;96:783–92.
Covaleda Ortega J, Gállego BJ. Haemoproteus aviares. Rev Ibér Parasitol. 1950;10:141–85.
Martinsen ES, Paperna I, Schall JJ. Morphological versus molecular identification of avian Haemosporidia: an exploration of three species concepts. Parasitology. 2006;133:279–88.
Baker JR. Haemoproteus palumbis sp. nov. (Sporozoa, Haemosporina) of the English Wood-Pigeon Columba p. palumbus. J Protozool. 1966;13:515–9.
Coatney GR, Roudabus RL. Some blood parasites from Nebraska birds. Am Midl Nat. 1937;18:1005–30.
Bennett GF. Avian Haemoproteidae. 8. The haemoproteids of the bee-eater family (Meropidae). Can J Zool. 1978;56:1721–5.
Bennett GF, Okia NO, Ashford RW, Campbell AG. Avian Haemoproteidae. II. Haemoproteus enucleator sp. n. from the Kingfisher, Ispidina picta (Boddaert). J Parasitol. 1972;58:1143–7.
Valkiūnas G, Iezhova T. Parasitic Protozoa of the blood of birds in the USSR 6 Haemoproteidae of Columbiformes and Coraciiformes (in Russian). Ekologija. 1990;2:86–103.
Dimitrov D, Iezhova TA, Zehtindjiev P, Bobeva A, Ilieva M, Kirilova M, et al. Molecular characterisation of three avian haemoproteids (Haemosporida, Haemoproteidae), with the description of Haemoproteus (Parahaemoproteus) palloris n. sp. Syst Parasitol. 2016;93:431–49.
Bennett GF, Campbell AG. Avian Haemoproteidae. III. Description of Haemoproteus fusca sp. n. and redescription of Haemoproteus halcyonis de Mello, 1935, parasites of the kingfisher family Alcedinidae. J Parasitol. 1973;59:337–43.
Ivanova K, Zehtindjieva P, Mariaux J, Dimitrov D, Georgiev BB. Avian haemosporidians from rain forests in Madagascar: molecular and morphological data of the genera Plasmodium, Haemoproteus and Leucocytozoon. Infect Genet Evol. 2018;58:115–24.
Václav R, Betáková T, Švančarová P, Pérez-Serrano J, Criado-Fornelio Á, Škorvanová L, et al. Nest ecology of blood parasites in the European roller and its ectoparasitic carnid fly. Exp Parasitol. 2016;165:71–80.
Bishop A, Bennett GF. Avian Haemoproteidae. 23. The haemoproteids of the avian family Coraciidae (rollers). Can J Zool. 1986;64:1860–3.
Savage AF, Greiner EC. Hematozoa of the avian family Brachypteraciidae (the ground-rollers). J Parasitol. 2004;90:1468–72.
Peirce MA, Adlard RD. The haemoproteids of the Cuculidae. J Nat Hist. 2005;39:2281–7.
Peirce MA. Haematozoa of East African birds. II. Redescription of Haemoproteus centropi, a parasite of the Cuculidae. E Afr Wildl J. 1977;15:57–60.
Wasielewski T, Wülker G. Die Hämoproteus-Infektion des Turmfalken. Arch Schiffs Tropenhyg. 1918;22:1–100.
Valkiūnas G, Ilgūnas M, Bukauskaitė D, Chagas CRF, Bernotienė R, Himmel T, et al. Molecular characterization of six widespread avian haemoproteids, with description of three new Haemoproteus species. Acta Trop. 2019;197: 105051.
Puech M-P, Chavatteb J-M, Landau I, Karadjian G. Haemoparasites of falcons in France: a 2-year survey in the Cevennes, with description of two new haemoproteid species from poly-parasitised birds. J Nat Hist. 2019;53:2023–37.
O’Roke EC. The morphology of Haemoproteus lophortyx sp. nov. Science. 1929;70:432.
White EM, Bennett GF. Avian Haemoproteidae. 12. The haemoproteids of the grouse family Tetraonidae. Can J Zool. 1979;57:1465–72.
Atkinson CT. Host specificity and morphometric variation of Haemoproteus meleagridis Levine, 1961 (Protozoa: Haemosporina) in gallinaceous birds. Can J Zool. 1986;64:2634–8.
Bennett GF, Peirce MA. The haemoproteids of the avian family Phasianidae. Can J Zool. 1989;67:1557–65.
Nandi NC, Mandal AK. Haemoproteus megapodius sp. nov. in Megapodius freycinet Abbotti Oberholser (Megapodiidae) from the South Nicobar. Rec Zool Surv India. 1980;77:51–4.
Bennett GF, Gabaldon A, Ulloa G. Avian Haemoproteidae. 17. The haemoproteids of the avian family Cracidae (Galliformes); the guans, curassows, and chachalacas. Can J Zool. 1982;60:3105–12.
Ferreira-Junior FC, de Angeli DD, Martins NRS, Valkiūnas G, Braga ÉM. Haemoproteus paraortalidum n. sp. in captive Black-fronted Piping-guans Aburria jacutinga (Galliformes, Cracidae): high prevalence in a population reintroduced into the wild. Acta Trop. 2018;188:93–100.
Gabaldon A, Ulloa G. Subespecie de Haemoproteus rotundus Oliger, 1956 (Haemosporina: Haemoproteidae) presente en Venezuela. Bol Dir Malariol San Amb. 1978;18:165–74.
Peirce MA. Haemoproteus balearicae sp. nov., from crowned cranes, Balearica pavonina pavonina and B. pavonina gibbericeps. Bull Epizoot Dis Africa. 1973;2:467–75.
Sacchi L, Prigioni C. Haematozoa of Italian birds. II: First European record of Haemoproteus gallinulae de Mello, 1935 from Gallinula chloropus and redescription (Apicomplexa, Haemosporina). Atti Soc ital Sci nat Museo civ Stor nat Milano. 1986;127:27–32.
Bennett GF. Avian Haemoproteidae. 14. The haemoproteids of the avian family Rallidae. Can J Zool. 1980;58:321–5.
Bennett GF, Forrester DJ, Greiner EC, Campbell AG. Avian Haemoproteidae. 4. Description of Haemoproteus telfordi sp. nov., and a review of the haemoproteids of the families Gruidae and Otidae. Can J Zool. 1975;53:72–81.
Travassos Santos Dias JA. Resultados de um reconhecimento zoologico no alto dos de um reconhecimento zoologico no alto limpopo efectuado pelos Drs. F. Zumpt e J. A. T. Santos Dias. V. Hematozoários das aves: generos Haemoproteus Kruse 1890 e Carpanoplasma n. gen. Moçambique. 1953;73:69–99.
Chavatte J-M, Okumura C, Landau I. Haematozoa of the Great Blue Turacos, Corythaeola cristata (Vieillot, 1816) (Aves: Musophagiformes: Musophagidae) imported to Singapore Jurong Bird Park with description and molecular characterisation of Haemoproteus (Parahaemoproteus) minchini new species (Apicomplexa: Haemosporidia: Haemoproteidae). Raffles Bull Zool. 2017;65:325–40.
Bennett GF, Greiner EC, Campbell AG. Avian Haemoproteidae. 5. The haemoproteids of the family Threskiornithidae. Can J Zool. 1975;53:634–8.
Tendeiro J. Acerca dos hematozoários de algumas aves da Guiné Portuguesa. Rev Med Ve. 1947;42:287–350.
Mohammed AHH. Systematic and experimental studies on protozoan blood parasites of Egyptian birds. Cairo: Cairo Univ Press; 1958.
Bennett GF, Nandi NC. Avian Haemoproteidae. 16. The haemoproteids of the avian family Capitonidae (the barbets). Can J Zool. 1981;59:2064–71.
Groff TC, Lorenz TJ, Crespo R, Iezhova TA, Valkiūnas G, Sehgal RNM. Haemoproteosis lethality in a woodpecker, with molecular and morphological characterization of Haemoproteus velans (Haemosporida, Haemoproteidae). Int J Parasitol Parasites Wildl. 2019;10:93–100.
Dimitrov D, Zehtindjiev P, Bensch S, Ilieva M, Iezhova T, Valkiūnas G. Two new species of Haemoproteus Kruse, 1890 (Haemosporida, Haemoproteidae) from European birds, with emphasis on DNA barcoding for detection of haemosporidians in wildlife. Syst Parasitol. 2014;87:135–51.
Greiner EC, Mandal AK, Nandi NC. Haemoproteus bennetti sp. n. and a review of the haemoproteids from the Picidae (Woodpeckers). J Parasitol. 1977;63:651–6.
Maqsood M. Haemoproteus handai (sp. nov.) occurring in Indian parakeet (Psittacula cyanocephala). Indian Vet J. 1943;20:109–11.
Bennett GF, Peirce MA. Avian Haemoproteidae. 21. The haemoproteids of the parrot family Psittacidae. Can J Zool. 1986;64:771–3.
Valkiūnas G, Pendl H, Olias P. New Haemoproteus parasite of parrots, with remarks on the virulence of haemoproteids in naive avian hosts. Acta Trop. 2017;176:256–62.
Bennett GF, Peirce MA. Haemoproteus psittaci n. sp. (Haemoproteidae) from the African grey parrot Psittacus erithacus L. Syst Parasitol. 1992;23:21–4.
Shamsuddin M, Mohammad MK. Haematozoa of some Iraqi birds with description of two new species, Haemoproteus pteroclis and Leucocytozoon nycticoraxi (Protozoa: Haemosporina). Bull Nat Hist Res Centre. 1980;7:111–55.
Valkiūnas G, Palinauskas V, Ilgūnas M, Bukauskaite D, Dimitrov D, Bernotienė R, et al. Molecular characterization of five widespread avian haemosporidian parasites (Haemosporida), with perspectives on the PCR-based detection of haemosporidians in wildlife. Parasitol Res. 2014;113:2251–63.
Karadjian G, Puech M-P, Duval L, Chavatte J-M, Snounou G, Landau I. Haemoproteus syrnii in Strix aluco from France: morphology, stages of sporogony in a hippoboscid fly, molecular characterization and discussion on the identification of Haemoproteus species. Parasite. 2013;20:32.
Bukauskaitė D, Žiegytė R, Palinauskas V, Iezhova TA, Dimitrov D, Ilgūnas M, et al. Biting midges (Culicoides, Diptera) transmit Haemoproteus parasites of owls: evidence from sporogony and molecular phylogeny. Parasit Vectors. 2015;8:303.
Levin II, Valkiūnas G, Santiago-Alarcon D, Cruz LL, Iezhova TA, O’Brien SL, et al. Hippoboscid-transmitted Haemoproteus parasites (Haemosporida) infect Galapagos pelecaniform birds: evidence from molecular and morphological studies, with a description of Haemoproteus iwa. Int J Parasitol. 2011;15:1019–27.
Work TM, Rameyer RA. Haemoproteus iwa n. sp. in great frigatebirds (Fregata minor [Gmelin]) from Hawaii: parasite morphology and prevalence. J Parasitol. 1996;82:489–91.
Bennett GF, Caine JR, Whiteway M. Avian Haemoproteidae. 22. The haemoproteids of the New World flycatchers, the Tyrannidae. Can J Zool. 1986;64:774–7.
Bennett GF, Bishop MA, Peirce MA. The haemoproteids of the avian families Eurylaimidae (broadbills) and Pittidae (pittas). Syst Parasitol. 1991;18:111–6.
Bennett GF, Caines JR, Woodworth-Lynas CB. Avian Haemoproteidae. 24. The haemoproteids of the New World passeriform families Formicariidae, Furnariidae, Mimidae, and Vireonidae. Can J Zool. 1987;65:317–21.
Perkins SL. Molecular systematics of the three mitochondrial protein-coding genes of malaria parasites: corroborative and new evidence for the origins of human malaria. Mitochondrial DNA. 2008;19:471–8.
Laird M, Laird E. Culicidae and haematozoa from Bellona and Rennell. Nat Hist Rennell Island, British Solomon Islands, Copenhagen. 1959;2:213–34.
Clark NJ, Adlard RD, Clegg SM. Molecular and morphological characterization of Haemoproteus (Parahaemoproteus) ptilotis, a parasite infecting Australian honeyeaters (Meliphagidae), with remarks on prevalence and potential cryptic speciation. Parasitol Res. 2015;114:1921–8.
Bennett GF, Bishop MA, Peirce MA. The species and distribution of the haemoproteids of the avian family Muscicapidae sensu latu (Passeriformes). J Nat Hist. 1991;25:23–43.
Peirce MA. Haematozoa of Zambian birds. VII. Redescription of Haemoproteus orioli from Oriolus oriolus (Oriolidae). J Nat Hist. 1984;18:785–7.
Bennett GF, Peirce MA. The haemoproteids of the Old World avian families Alaudidae (larks), Irenidae (leaf birds), and Motacillidae (wagtails and pipits). J Nat Hist. 1990;24:939–47.
Savage AF, Ariey F, Greiner EC. Hematozoa of the avian family Vangidae (the Vangas). J Parasitol. 2004;90:1475–9.
Goulding W, Adlard RD, Clegg SM, Clark NJ. Molecular and morphological description of Haemoproteus (Parahaemoproteus) bukaka (species nova), a haemosporidian associated with the strictly Australo-Papuan host subfamily Cracticina. Parasitol Res. 2016;115:3387–400.
Peirce MA. Haematozoa of Zambian birds. III. Description of Haemoproteus cublae sp. nov. from Dryoscopus cubla (Malaconotidae). J Nat Hist. 1984;18:219–21.
Valkiūnas G, Iezhova T. Parasitic Protozoa of the blood of birds in the USSR. 8. Haemoproteidae of Passeriformes (in Russian). Ekologija. 1992;1:59–83.
Bishop MA, Bennett GF. The haemoproteids of the avian families Corvidae (crows and jays) and Sturnidae (starlings and mynas) (Passeriformes). Can J Zool. 1990;68:2251–6.
Hellgren O, Križanauskienė A, Valkiūnas G, Bensch S. Diversity and phylogeny of mitochondrial cytochrome b lineages from six morphospecies of avian Haemoproteus (Haemosporida: Haemoproteidae). J Parasitol. 2007;93:889–96.
Valkiūnas G, Palinauskas V, Križanauskienė A, Bernotienė R, Kazlauskienė R, Iezhova TA. Further observations on in vitro hybridization of hemosporidian parasites: patterns of ookinete development in Haemoproteus spp. J Parasitol. 2013;99:124–36.
Chagas CRF, Bukauskaitė D, Ilgūnas M, Bernotienė R, Iezhova T, Valkiūnas G. Sporogony of four Haemoproteus species (Haemosporida: Haemoproteidae), with report of in vitro ookinetes of Haemoproteus hirundinis: phylogenetic inference indicates patterns of haemosporidian parasite ookinete development. Parasit Vectors. 2019;12:422.
Valkiūnas G, Iezhova TA, Palinauskas V, Ilgūnas M, Bernotienė R. The evidence for rapid gametocyte viability changes in the course of parasitemia in Haemoproteus parasites. Parasitol Res. 2015;144:2903–9.
Peirce MA. Haematozoa of Zambian birds. VIII. Redescription of Haemoproteus dicruri from Dicrurus adsimilis (Dicruridae). J Nat Hist. 1984;18:789–91.
Peirce MA. Haematozoa of British birds. V. Redescription of Haemoproteus majoris (Laveran) from the great tit Parus major. J Nat Hist. 1981;15:151–4.
Žiegytė R, Markovets MY, Bernotienė R, Mukhin A, Iezhova TA, Valkiūnas G, et al. The widespread biting midge Culicoides impunctatus (Ceratopogonidae) is susceptible to infection with numerous Haemoproteus (Haemosporida) species. Parasit Vectors. 2017;10:397.
Bennett GF. New species of haemoproteids from the avian families Paridae and Sittidae. Can J Zool. 1989;67:2685–8.
Valkiūnas G, Iezhova T. Parasitic Protozoa of the blood of birds in the USSR 9 Haemoproteidae of Passeriformes, Charadriiformes and Anseriformes (in Russian). Ekologija. 1992;2:52–73.
Bennett GF, Bishop MA. The haemoproteids (Apicomplexa: Haemoproteidae) of the flowerpeckers of the avian family Dicaeidae (Passeriformes). Syst Parasitol. 1990;18:159–64.
Valkiūnas G, Iezhova T, Chernetsov NS. Haemoproteus payevskyi sp. n. (Haemosporidia: Haemoproteidae) from reed warbler Acrocephalus scirpaceus (Passeriformes)] in Russian). Parazitologiya. 1994;28:469–74.
Križanauskienė A, Iezhova TA, Palinauskas V, Chernetsov N, Valkiūnas G. Haemoproteus nucleocondensus n. sp. (Haemosporida, Haemoproteidae) from a Eurasian songbird, the Great Reed Warbler Acrocephalus arundinaceus. Zootaxa. 2012;3441:36–46.
Valkiūnas G, Križanauskienė A, Iezhova TA, Hellgren O, Bensch S. Molecular phylogenetic analysis of circumnuclear hemoproteids (Haemosporida: Haemoproteidae) of sylviid birds, with a description of Haemoproteus parabelopolskyi sp. nov. J Parasitol. 2007;93:680–7.
Žiegytė R, Palinauskas V, Bernotienė R, Iezhova TA, Valkiūnas G. Haemoproteus minutus and Haemoproteus belopolskyi (Haemoproteidae): complete sporogony in the biting midge Culicoides impunctatus (Ceratopogonidae), with implications on epidemiology of haemoproteosis. Exp Parasitol. 2014;145:74–9.
Valkiūnas G, Ilgūnas M, Chagas CRF, Bernotienė R, Iezhova TA. Molecular characterization of swallow haemoproteids, with description of one new Haemoproteus species. Acta Trop. 2020;207: 105486.
White EM, Bennett GF. Avian Haemoproteidae. 9. Description of Haemoproteus stellaris n. sp. and a review of the haemoproteids of the swallow family Hirundinidae. Can J Zool. 1978;56:2110–6.
Bennett GF, Peirce M. Avian Haemoproteidae. 15. The haemoproteids of the avian family Zosteropidae. Can J Zool. 1981;59:1155–60.
Clark NJ, Wells K, Dimitrov D, Clegg SM. Co-infections and environmental conditions drive the distributions of blood parasites in wild birds. J Animal Ecol. 2016;85:1461–70.
Ishtiaq F, Rao M, Palinauskas V. Molecular characterization and morphological description of cryptic haemoproteids in the laughingthrushes (Leiothrichidae) in the western and eastern Himalaya. India Wellcome Open Res. 2018;3:94.
Rahal EM, Bishop MA, Bennett GF. Avian Haemoproteidae. 25. The haemoproteids of the avian family Pycnonotidae (bulbuls). Can J Zool. 1987;65:322–8.
Valkiūnas G, Iezhova TA, Loiseau C, Chasar A, Smith TB, Sehgal RNM. New species of haemosporidian parasites (Haemosporida) from African rainforest birds, with remarks on their classification. Parasitol Res. 2008;103:1213–28.
Chagas CRF, Bukauskaitė D, Ilgūnas M, Iezhova T, Valkiūnas G. A new blood parasite of leaf warblers: molecular characterization, phylogenetic relationships, description and identification of vectors. Parasit Vectors. 2018;11:538.
Križanauskienė A, Pérez-Tris J, Palinauskas V, Hellgren O, Bensch S, Valkiūnas G. Molecular phylogenetic and morphological analysis of haemosporidian parasites (Haemosporida) in a naturally infected European songbird, the blackcap Sylvia atricapilla, with description of Haemoproteus pallidulus sp. nov. Parasitology. 2010;137:217–27.
Hernández-Lara C, Duc M, Ilgūnas M, Valkiūnas G. Massive infection of lungs with exo-erythrocytic meronts in European Robin Erithacus rubecula during natural Haemoproteus attenuatus haemoproteosis. Animals. 2021;11:3273.
Peirce MA. Haematozoa of Zambian birds. II. Description of Haemoproteus balmorali sp. nov. from Turdidae and Muscicapidae. J Nat Hist. 1984;18:123–5.
Valkiūnas G, Iezhova TA, Križanauskienė A, Palinauskas V, Bensch S. In vitro hybridization of Haemoproteus spp.: an experimental approach for direct investigation of reproductive isolation of parasites. J Parasitol. 2008;94:1385–94.
Bukauskaitė D, Chagas CRF, Bernotienė R, Žiegytė R, Ilgūnas M, Iezhova T, Valkiūnas G. A new methodology for sporogony research of avian haemoproteids in laboratory reared Culicoides spp., with description of the complete sporogonic development of Haemoproteus pastoris. Parasit Vectors. 2019;12:582.
Valkiūnas G, Iezhova T. Haemoproteus kairullaevi sp. n. (Haemosporidia: Haemoproteidae) from common mynah Acridotheres tristis (Passeriformes) (in Russian). Zool zhurnal. 1993;72:141–6.
Bennett GF, Campbell AG. Avian Haemoproteidae. I. Description of Haemoproteus fallisi n. sp. and a review of the haemoproteids of the family Turdidae. Can J Zool. 1972;50:1269–75.
Roudabush RL, Coatney GR. Haemoproteus beckeri n. sp. and Trypanosoma laverani var toxostomae n. var. from the brown thrasher (Toxostoma rufum). Iowa State Coll J Sci. 1935;10:1–6.
Iezhova TA, Dodge M, Sehgal RN, Smith TB, Valkiūnas G. New avian Haemoproteus species (Haemosporida: Haemoproteidae) from African birds, with a critique of the use of host taxonomic information in hemoproteid classification. J Parasitol. 2011;97:682–94.
Bennett GF, Peirce MA. The haemoproteids of the weaver finches (Passeriformes). J Ant Hist. 1991;25:7–22.
Zehtindjiev P, Križanauskienė A, Scebba S, Dimitrov D, Valkiūnas G, Heggemann A, et al. Haemosporidian infections in skylarks (Alauda arvensis): a comparative PCR-based and microscopy study on the parasite diversity and prevalence in southern Italy and the Netherlands. Eur J Wildl Res. 2012;58:335–44.
Iezhova TA, Valkiūnas G, Loiseau C, Smith TB, Sehgal RNM. Haemoproteus cyanomitrae sp. nov. (Haemosporida, Haemoproteidae) from a widespread African songbird, the olive sunbird Cyanomitrae olivacea. J Parasitol. 2010;96:137–43.
Dimitrov D, Valkiūnas G, Zehtindjiev P, Ilieva M, Bensch S. Molecular characterization of haemosporidian parasites (Haemosporida) in yellow wagtail (Motacilla flava), with description of in vitro ookinetes of Haemoproteus motacillae. Zootaxa. 2013;3666:369–81.
Mantilla JS, Gonzalez AD, Lotta IA, Moens M, Pacheco MA, Escalante AA, et al. Haemoproteus erythrogravidus n. sp. (Haemosporida, Haemoproteidae): description and molecular characterization of a widespread blood parasite of birds in South America. Acta Trop. 2016;159:83–94.
Anjos CC, Chagas CRF, Fecchio A, Schunck F, Costa-Nascimento MJ, Monteiro EF, et al. Avian malaria and related parasites from resident and migratory birds in the Brazilian Atlantic forest, with description of a new Haemoproteus species. Pathogens. 2021;10:103.
Burry-Caines JR, Bennett GF. The Haemoproteidae (Apicomplexa: Haemosporina) of the avian families Fringillidae and Emberizidae. Can J Zool. 1992;70:1149–60.
Ricklefs RE, Fallon SM. Diversification and host switching in avian malaria parasites. Proc R Soc Lond, B, Biol Sci. 2002;269:885–92.
Ortiz-Catedral L, Brunton D, Stidworthy MF, Elsheikha HM, Pennycott T, Schulze C. Haemoproteus minutus is highly virulent for Australasian and South American parrots. Parasit Vectors. 2019;12:40.
Bush AO, Lafferty KD, Lotz JM, Shostak AW. Parasitology meets ecology on its own terms: Margolis et al. revisited. J Parasitol. 1997;83:575–83.
Valkiūnas G, Iezhova TA, Križanauskienė A, Palinauskas V, Sehgal RNM, Bensch S. A comparative analysis of microscopy and PCR-based detection methods for blood parasites. J Parasitol. 2008;94:1395–401.
Campbell TW. Avian hematology and cytology. Ames: Iowa State University Press; 1995.
Bertram MR, Hamer SA, Hartup BK, Snowden KF, Medeiros MC, Outlaw DC, et al. A novel Haemosporida clade at the rank of genus in North American cranes (Aves: Gruiformes). Mol Phylogenet Evol. 2017;109:73–9.
Chagas CRF, Guimarães LO, Monteiro EF, Valkiūnas G, Michele Viana Katayama MV, Santos SV, et al. Hemosporidian parasites of free-living birds in the São Paulo Zoo, Brazil. Parasitol Res. 2016;115:1443–52.
Waldenström J, Bensch S, Kiboi S, Hasselquist D, Ottosson U. Cross-species infection of blood parasites between resident and migratory songbirds in Africa. Mol Ecol. 2002;11:1545–54.
Yumoto F, Tokiwa T, Chou S, Ueda M. Haemoproteus columbae infection in a straggler racing pigeon sheltered in Japan. J Vet Med Sci. 2021;83:1760–3.
Beadell JS, Ishtiaq F, Covas R, Melo M, Warren BH, Atkinson CT, et al. Global phylogeographic limits of Hawaii’s avian malaria. Proc Biol Sci. 2006;273:2935–44.
Chagas CR, Valkiūnas G, de Oliveira GL, Monteiro EF, Guida FJ, Simões RF, et al. Diversity and distribution of avian malaria and related haemosporidian parasites in captive birds from a Brazilian megalopolis. Malar J. 2017;16:83.
Marzal A, Ricklefs RE, Valkiūnas G, Albayrak T, Arriero E, Bonneaud C, et al. Diversity, loss and gain of malariae parasites in a globally invasive bird. PLoS ONE. 2011;6: e21905.
Martinsen ES, Perkins SL, Schall JJ. A three-genome phylogeny of malaria parasites (Plasmodium and closely related genera): evolution of life-history traits and host switches. Mol Phylogenet Evol. 2008;47:261–73.
International Commission on Zoological Nomenclature. International code of zoological nomenclature. 4th ed. London: The International Trust for Zoological Nomenclature; 1999.
Valkiūnas G, Iezhova TA. Keys to the avian malaria parasites. Malar J. 2018;17:212.
Valkiūnas G, Atkinson CT, Bensch S, Sehgal RN, Ricklefs RE. Parasite misidentifications in GenBank: how to minimize their number? Trends Parasitol. 2008;24:247–8.
Valkiūnas G, Ashford RW, Bensch S, Killick-Kendrick R, Perkins S. A cautionary note concerning Plasmodium in apes. Trends Parasitol. 2011;27:231–2.
Acknowledgements
We thank Dr. R. Adlard, Dr. C. T. Atkinson, Dr. S. Bensch, Dr. E. M. Braga, Dr. M. Bryant, Dr. D. Dimitrov, Dr. F. C Ferreira-Junior, Dr. E. Hoberg, Dr. N.E. Matta, Dr. M. A. J. Moens, Dr. Dr. P. G. Parker, Dr. R. N. M. Sehgal, Dr. A. Warren, Dr. H. Weissenböck, and Dr. P. Zehtindjiev for assistance in accessing parasite material. Dr. R. N. M. Sehgal kindly commented on an early draft of the article.
Funding
This study was supported by the Open Access to research infrastructure of the NRC under Lithuanian open access network initiative.
Author information
Authors and Affiliations
Contributions
GV collected published articles and collection material, analysed the literature data and wrote the manuscript; TAI and GV analysed preparations of the blood stages; TAI and GV prepared plates of images. Both authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Ethics approval and consent to participate
Not applicable.
Consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/. The Creative Commons Public Domain Dedication waiver (http://creativecommons.org/publicdomain/zero/1.0/) applies to the data made available in this article, unless otherwise stated in a credit line to the data.
About this article
Cite this article
Valkiūnas, G., Iezhova, T.A. Keys to the avian Haemoproteus parasites (Haemosporida, Haemoproteidae). Malar J 21, 269 (2022). https://doi.org/10.1186/s12936-022-04235-1
Received:
Accepted:
Published:
DOI: https://doi.org/10.1186/s12936-022-04235-1