Abstract
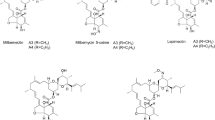
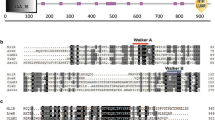
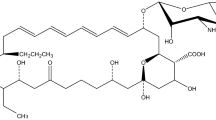
Milbemycin, a group of 16-membered macrolide antibiotics produced by Streptomyces milbemycinicus, has been widely used as an insecticide and an anthelmintic. To enhance the production of milbemycin, ribosomal engineering with heterologous expression of a regulatory gene was used to screen for a high milbemycin-producing mutant. The mutant S. milbemycinicus R2-6-5 isolated after the treatment of S. milbemycinicus A2079 with 6 μg/mL streptomycin produced 172.5 and 163.1% of milbemycins A3 and A4, respectively, compared with original strain. Analysis of the gene rsmG revealed a frameshift mutation, one cytidine unit being inserted into the 21 position (21C → 21CC). The heterologous regulatory gene aveR, which belongs to the LAL-family was integrated into the genome of S. milbemycinicus R2-6-5 denoted S. milbemycinicus J37 to enhance production of milbemycin. The production of milbemycins A3 and A4 in S. milbemycinicus J37 reached 758.9 and 279.0 μg/g respectively, representing 142 and 61% higher yields over S. milbemycinicus R2-6-5. The combination of ribosomal engineering and heterologous regulatory gene expression in S. milbemycinicus J37 resulted in an increase by 12.4-fold for milbemycin A3 and 11.7-fold for milbemycin A4, respectively, when compared to the original strain. Overall, these results demonstrate that combining available technologies for strain modification such as ribosome engineering technology and heterologous regulatory gene expression is an effective approach for development of high milbemycin-producing strains.





Similar content being viewed by others
REFERENCES
Satoi, S., Muto, N., Hayashi, M., Fujii, T., and Otani, M., J. Antibiot., 1980, vol. 33, no. 4, pp. 364–376.
Shoop, W. L., Vet. Parasitol., 1995, vol. 59, no. 2, pp. 139–156.
Cerkvenik-Flajs, V., Keegan, J., Kolar, L., Radeck, W., and Danaher, M., Curr. Pharm. Biotechnol., 2012, vol. 13, no. 6, pp. 936–951.
Jacobs, C.T. and Scholtz, C.H., Onderstepoort J. Vet. Res., 2015, vol. 82, no. 1, pp. 1–8.
Pluschkell, U., Horowitz, A. R., and Ishaaya, I., Phytoparasitica, 1999, vol. 27, no. 3, p. 183.
Wang, X. J., Wang, X. C., and Xiang, W. S., World J. Microb. Biot., 2009, vol. 25, no. 6, pp. 1051–1056.
Wang, H.Y., Zhang, J., Zhang Y.J., Zhang, B., Liu, C.X., Wang, X.J., and Xiang, W.S., Appl. Microbiol. Biot., 2014, vol. 98, no. 23, pp. 9703–9712.
Hosaka, T., Mol. Microbiol., 2010, vol. 61, no. 4, pp. 883–897.
Ochi, K., Susumu Okamoto, S., Tozawa, Y., Inaoka, T., and Kurosawa, K., Adv. Appl. Microbiol., 2003, vol. 56, no. 1, pp. 155–184.
Nishimura, K., Hosaka, T., Tokuyama, S., Okamoto, S., and Ochi, K., J. Bacteriol., 2007, vol. 189, no. 10, pp. 3876–3883.
Xu, J., Tozawa, Y., Lai, C., Hayashi, H., and Ochi, K., Mol. Genet. Genomics, 2002, vol. 268, no. 2, pp. 179–189.
Wang, G., Hosaka, T., and Ochi, K., Appl. Environ. Microb., 2008, vol. 74, no. 9, pp. 2834–2840.
Lu, F., Hou, Y., Zhang, H., Chu, Y., Xia, H., and Tian, Y., 3 Biotech., 2017, vol. 7, no. 4, p. 250.
Liu, G., Chater, K. F., Chandra, G., Niu, G., and Tan, H., Microbiol. Mol. Biol. Rev., 2013, vol. 77, no. 1, pp. 112–143.
Guo, J., Zhao, J., Li, L., Chen, Z., Wen, Y., and Li, J., Mol. Genet. Genomics, 2010, vol. 283, no. 2, pp. 123–133.
Wang, X.J., Yan, Y.J., Zhang, B., An, J., and Xiang, W.S., J. Bacteriol., 2010, vol. 192, no. 17, pp. 4526–4527.
Zhang, Y., He, H., Liu, H., Wang, H., Wang, X. et al., Microb. Cell Fact., 2016, vol. 15, no. 1, p. 152.
Kitani, S., Ikeda, H., Sakamoto, T., Noguchi, S., and Nihira, T., Appl. Microbiol. Biot., 2009, vol. 82, no. 6, pp. 1089–1096.
Wilson, D., Xue, Y., and Reynolds, K., J. Bacteriol., 2001, vol. 183, no. 11, pp. 3468–3475.
Huang, M., Li, M., Feng, Z., Liu, Y., Chu, Y., and Tian, Y., World J. Microb. Biot., 2011, vol. 27, no. 9, pp. 2103–2109.
Chen, C., Zhao, X., Jin, Y., Zhao, Z., and Suh, J.W., Plasmid, 2014, vol. 76, pp. 79–86.
Gust, B., Challis, G., Fowler, K., Kieser, T., and Chater, K., Proc. Natl. Acad. Sci. U. S. A., 2003, vol. 100, no. 4, pp. 1541–1546.
Takiguchi, Y., Ono, M., Muramatsu, S., Ide, J., Mishima, H., and Terao, M., J. Antibiot., 1983, vol. 36, no. 5, p. 502.
Ikeda, H., Ishikawa, J., Hanamoto, A., Shinose, M., Kikuchi, H., Shiba, T., et al., Nat. Biotechnol., 2003, vol. 21, no. 5, pp. 526–531.
Jung, W. S., Yoo, Y. J., and Park, J. W., Appl. Microbiol. Biotechnol., 2011, vol. 91, no. 5, pp. 1389–1397.
Okamoto-Hosoya, Y., Okamoto, S., and Ochi, K., Appl. Environ. Microb., 2003, vol. 69, no. 7, pp. 4256–4259.
Kagan, R. M. and Clarke, S., Arch. Biochem. Biophys., 1994, vol. 310, no. 2, pp. 417–427.
Okamoto, S., Lezhava, A., Hosaka, T., Okamoto-Hosoya, Y., and Ochi, K., J. Bacteriol., 2003, vol. 185, no. 2, pp. 601–609.
Leskiw, B. K., Bibb, M. J., and Chater, K. F., Mol. Microbiol., 1991, vol. 5, no. 12, pp. 2861–2867.
Leskiw, B.K., Mah, R., Lawlor, E J., and Chater, K.F., J. Bacteriol., 1993, vol. 175, no. 7, pp. 1995–2005.
Shima, J., Hesketh, A., Okamoto, S., Kawamoto, and Ochi, K., J. Bacteriol., 1996, vol. 178, no. 24, pp. 7276–7284.
Funding
This work was financially supported by National Key Research and Development Program of China (2018YFC1802201) and the Opening Project of Key Laboratory of Leather Chemistry and Engineering (China, Sichuan University), Ministry of Education (20826041C4159).
Author information
Authors and Affiliations
Contributions
All the authors contributed to this work. Yongqiang tian conceived, designed and support the experiments; Shiting Wang and Fenjuan Lu performed the experiments and analyzed the data; Shiting Wang wrote the paper; Zhigang Yang and Zhenjiang Li participated in the data analysis of untranslated region. All authors reviewed and approved the manuscript.
Corresponding authors
Ethics declarations
The authors declare that there are no conflicts of interest. The authors declare that there are no ethical issues in this study.
Rights and permissions
About this article
Cite this article
Wang, S., Lu, F., Yang, Z. et al. Combining Ribosomal Engineering with Heterologous Expression of a Regulatory Gene to Improve Milbemycin Production in Streptomyces milbemycinicus A2079. Appl Biochem Microbiol 57, 303–310 (2021). https://doi.org/10.1134/S0003683821030133
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1134/S0003683821030133