Abstract
Polyhedra can encapsulate other proteins and have potential applications as protein stabilizers. The extremely stable polyhedra matrix may provide a platform for future engineered micro-crystal devices. However, the protein composition of the polyhedra matrix remains largely unknown. In this study, the occlusion-derived virus (ODV)-removed BmNPV polyhedra matrix fraction was subjected to SDS-PAGE and then an LC-ESI-MS/MS analysis using a Thermo Scientific Q Exactive mass spectrometer. In total, 28 host and 91 viral proteins were identified. The host components were grouped into one of six categories, i.e., chaperones, ubiquitin and related proteins, host helicases, cytoskeleton-related proteins, RNA-binding proteins and others, according to their predicted Pfam domain(s). Most viral proteins may not be essential for polyhedra assembly, as evidenced by studies in the literature showing that polyhedra formation occurs in the nucleus upon the disruption of individual genes. The structural role of these proteins in baculovirus replication will be of significant interest in future studies. The immobilization of enhanced green fluorescent protein (eGFP) into the polyhedra by fusing with the C-terminus of BM134 that is encoded by open reading frame (ORF) 134 suggested that the polyhedra had a powerful capacity to trap foreign proteins, and BM134 was a potential carrier for incorporating proteins of interest into the polyhedra.
Similar content being viewed by others
Introduction
Baculoviridae is a family of insect DNA viruses that have a large, circular, supercoiled and double-stranded DNA genome within a rod-shaped nucleocapsid1. During the life cycle of baculovirus, two typical progeny virions, i.e., budded virus (BV) and occlusion-derived virus (ODV), are produced. These virion phenotypes are genetically identical but differ structurally and functionally. BVs are responsible for cell-to-cell infections in cultured cells and tissues of susceptible hosts, while ODVs are required for horizontal insect-to-insect spreading1, 2. ODVs are embedded in a protein crystal called occlusion body (OB)3. Once ingested by a susceptible larva, the OBs are quickly dissolved due to the alkaline pH of the host midgut. ODVs are released to cross the peritrophic membrane and initiate the primary infection4. During the early stage of infection, nucleocapsids are transported through the nuclear membrane and migrate across the cytosol to the cell membrane for budding, thereby producing extracellular BVs. During the very late stage of infection, nucleocapsids in the nucleus are enveloped, resulting in ODVs that are packaged into the crystalline polyhedra matrix to form OBs.
The OBs, also named polyhedra, are stable and resistant to most normal environmental conditions due to the protective effect of multiple layers that are formed mainly by the polyhedrin envelope protein (PEP)5, thereby allowing the ODV virions to remain infectious in soil indefinitely1. Polyhedrin, a highly expressed protein with a molecular weight of ~29 kDa, is the major component protein in the polyhedra matrix. Polyhedrin is one of the most conserved proteins of alpha-baculovirus. Certain single point mutations in polyhedrin result in phenotypic changes, such as large, cuboid polyhedra that occlude no or few ODVs6,7,8,9. Polyhedra without ODVs have potential applications as stabilizers of proteins of interest, such as enzymes and growth factors, that are encapsulated in the polyhedra matrix10, 11. To explore the use of polyhedra as a framework for future engineered micro-crystal devices, an analysis of the component proteins associated with the polyhedra matrix is required.
The availability of genome sequences facilitates component analyses of baculoviruses on a proteomic scale. To date, the ODV components of eight baculoviruses, i.e., Autographa californica multiple nucleopolyhedrovirus (AcMNPV)12, Culex nigripalpus nucleopolyhedrovirus (CuniNPV)13, BmNPV14, Chrysodeixis chalcites nucleopolyhedrovirus (ChchNPV)15, Helicoverpa armigera nucleopolyhedrovirus (HaNPV)16, 17, Anticarsia gemmatalis multiple nucleopolyhedrovirus (AgMNPV)18, Mamestra brassicae nucleopolyhedrovirus (MabrNPV)19, Pieris rapae granulovirus (PrGV)20 and Clostera anachoreta granulovirus (ClanGV)21, and the BV components of four viruses, i.e., AcMNPV22, HaNPV17, AgMNPV18 and MabrNPV19, have been reported. These data shed light on the structure and assembly of baculovirus virions; however, in comparison, little is known regarding the structural composition of the polyhedra matrix.
In this study, a comprehensive analysis of the BmNPV polyhedra matrix-associated component proteins was performed using a Thermo Scientific Q Exactive mass spectrometer. In total, 28 host and 91 viral proteins were identified to be associated with the polyhedra matrix of BmNPV. Most of these viral proteins may not be essential for polyhedra assembly, as evidenced by studies in the literature that polyhedra formation in the nucleus was shown to occur upon the disruption of individual genes. The host components had been grouped into one of six categories, i.e., chaperones, ubiquitin and related proteins, host helicases, cytoskeleton-related proteins, RNA-binding proteins and others, according to their predicted Pfam domain(s). These host-derived proteins may be important for the assembly and maturation of polyhedra. These findings may provide novel insights into the baculovirus structure and shed light on the powerful ability of polyhedra to encapsulate foreign proteins.
Results and Discussion
SDS-PAGE analysis of ODV-removed polyhedra matrix
To identify proteins associated with the BmNPV polyhedra matrix, polyhedra from infected cells were purified, and the virions were released and pelleted by continuous sucrose centrifugation. The ODV pellet and sample over the top gradient were collected for the SDS-PAGE analysis and then made visible using Coomassie brilliant blue staining. The results showed that certain protein bands were present in both fractions (Supplementary Fig. S1), which is consistent with previous observations in an earlier proteomic investigation of CuniNPV ODV13. The total polyhedra matrix proteins on the gel were divided into 5 regions (M1 to M5) (Fig. 1) and subjected to an in-gel trypsin digestion and LC-ESI-MS/MS analysis using a Thermo Scientific Q Exactive mass spectrometer. A comprehensive compilation of the data was provided in supplementary Tables S1 to S9.
BmNPV polyhedra matrix proteins were separated on 12% (A) and 15% (B) SDS-PAGE gels. Purified polyhedra were treated with an alkaline solution. Undissolved polyhedra were removed by low-speed centrifugation at 500 × g. The resulting supernatant was collected and then centrifuged by continuous sucrose gradients to remove the ODVs. Protein sample over the upper gradient was separated by SDS-PAGE. Proteins on the gel were excised into five contiguous sections (M1 to M5) and subjected to in-gel digestion and LC-ESI-MS/MS analysis. Two M5 sections shown in (A) and (B) were combined for the determination. Lane M, protein marker.
Identification of host proteins associated with the BmNPV polyhedra matrix
Certain viruses trap the host proteins in their virions either specifically or randomly. An earlier study has shown that baculoviruses can trap heterologous chloramphenicol acetyltransferase in BV particles nonspecifically23. In this present study, more than 350 host proteins were identified to be associated with the BmNPV polyhedra matrix, which is indicative of the powerful ability of polyhedra to encapsulate foreign proteins. Twenty-eight of these identified proteins that had more than 1 peptide were listed and grouped, according to their predicted Pfam domain(s), into one of six categories, i.e., chaperones, ubiquitin and related proteins, host helicases, cytoskeleton-related proteins, RNA-binding proteins and others (Table 1). These proteins may play a structural role in the long-term stability of polyhedra in most environmental conditions or participate in the polyhedra assembly and maturation and then became trapped in the polyhedra.
During viral infections, many viral encoded proteins are synthesized in large amounts in a relatively short time and protein folding can become a limiting step during their active conformation and trafficking. Therefore, on the one hand, viruses require cellular chaperones for their protein folding processes; on the other hand, because chaperones are involved in the regulation of fundamental cellular processes, viruses must interact with chaperones24. The HSC/HSP70 family chaperones, which are central components of the cellular chaperone network, are involved in all stages of the viral life cycle of replication25, 26. In the case of baculovirus, the expression of HSP70 is often induced upon viral replication in vivo and in vitro 27. HSP70 plays a role in facilitating the genome synthesis of AcMNPV and the release of progeny BVs28, and is identified as a component of BV and ODV of BmNPV14, 29, BV of HaNPV17, and ODV of AgMNPV18. In this report, HSP70–4 was identified by an LC-ESI-MS/MS analysis to be associated with the BmNPV polyhedra matrix. The association between HSP70-4 and the polyhedra matrix was further confirmed by a Western blot analysis using a rabbit polyclonal anti-HSC/HSP70 antibody, which showed certain bands in five excised regions of the gel (Fig. 2A), and an immunofluorescence microscopy assay, which showed co-localization of HSP70-4 and polyhedrin in the polyhedra in the nucleus of infected cells (Fig. 2B). These results suggested that HSP70-4 was a component protein of the BmNPV polyhedra matrix. In addition to HSP70-4, this study identified HSP40 and two chaperone-like proteins, i.e., cyclophilin A and FKBP, to be associated with the BmNPV polyhedra matrix (Table 1). HSP40, which is a co-chaperone of HSP70, is involved in the enhancement or inhibition of pathogenesis in a wide range of viral infections30. Cyclophilin A has been shown to be recruited specifically into the virion for the infectivity of human immunodeficiency virus-131, 32. For baculovirus HaNPV, cyclophilin A is identified to be associated with virion ODV and BV17. FKBP is also a component of the HaNPV ODV17. However, to date, their roles in baculovirus infections are not investigated yet.
Host protein HSP70-4 was associated with the BmNPV polyhedra matrix. (A) Western blot analysis of HSP70-4 in the BmNPV polyhedra matrix. The fraction over the upper sucrose gradient was loaded for the Western blot analysis. Images were visualized using a chemiluminescence detection system. Lane M, the protein marker. (B) Co-localization of polyhedrin with HSP70-4 in the polyhedra in infected BmN cells. Cells were infected with the BmNPV T3 isolate at an MOI of 10 TCID50/cell. At 48 h p.i. the cells were fixed, permeabilized, and then incubated in a buffer containing mouse monoclonal anti-polyhedrin and rabbit polyclonal anti-HSC/HSP70 antibodies. Next, the cells were stained with the Texas Red-conjugated goat anti-mouse IgG, FITC-conjugated goat anti-rabbit IgG and Hoechst 33342, washed with 1× PBS, and then imaged under a Leica TCS SP5 confocal laser scanning microscope using the Texas Red filter set for the Texas Red-conjugated antibody and the FITC filter set for the FITC-conjugated antibody. In the merged images, yellow suggested an association between polyhedrin and HSP70-4 in the polyhedra matrix (Merge I), and magenta indicated the nuclear location of the polyhedra (Merge II).
Ubiquitin is a well-conserved protein that contains 76 amino acids and plays important roles in many diverse cellular activities33, and certain viral processes, such as virus entry and egress34. The ubiquitin-proteasome pathway plays a central role in BmNPV infections since its inhibition by the proteasome inhibitor MG-132 results in reduced BV and ODV formation35. The inhibitor bortezomib also leads to similar observations in terms of a reduced progeny BV production, restricted ODV proliferation, fewer copies of viral genomic DNA replication and a decreased number of ie1 and polyhedrin transcripts36. Ubiquitin homologs are found in most lepidopteran baculovirus genomes. ORF26 in the BmNPV genome encodes a ubiquitin homolog protein. During the very late stage of polyhedra formation, the BmNPV ubiquitin is largely and evenly localized in the nucleus, and in the cytoplasm, it exhibits a dotted distribution37. The identification of host and viral ubiquitins (Tables 1 and 2) prompted the question of whether certain proteins that are associated with the BmNPV polyhedra matrix were ubiquitylated. To address it, the polyhedra matrix sample was subjected to a Western blot analysis using antibodies specific to different forms of ubiquitin. Both the FK2 and FK1 antibodies recognize poly-ubiquitin conjugates, but the FK2 antibody also recognizes conjugated mono- and multiple mono-ubiquitin. Certain bands were observed by immunostaining with the FK2 antibody (Fig. 3A). When the sample was analyzed using the FK1 antibody, several bands were detected (Fig. 3B). These results demonstrated that certain mono- (and/or multiple mono-) and poly-ubiquitinated proteins were present in the BmNPV polyhedra matrix. Generally, proteins conjugated to poly-ubiquitin chains linked at the K48 residue of ubiquitin are targeted for degradation by the proteasome145, whereas proteins that are conjugated to K63-linked poly-ubiquitin chains play other roles, such as the regulation of protein function and cellular trafficking146. Staining with the K48- and K63-specific antibodies revealed a ubiquitination profile that was similar to that observed using the FK1 antibody (Fig. 3C, D). The LUB9 antibody recognizes the head-to-tail linear poly-ubiquitin chain in which the C-terminal Gly of one ubiquitin monomer was conjugated to the N-terminal Met of the next monomer. After staining with the LUB9 antibody, a visible band was detected (Fig. 3E). Thus, the differences observed in the staining using several antibodies were interpreted as mono- and/or multiple mono-ubiquitination.
Western blot analysis of ubiquitylated proteins associated with the polyhedra matrix of BmNPV. Polyhedra matrix proteins were separated on SDS-PAGE gel and then analyzed using mouse monoclonal anti-ubiquitin antibody clone FK2 (A) and FK1 (B), rabbit monoclonal anti-ubiquitin antibody Lys48-specific clone Apu2 (C) and Lys63-specific clone Apu3 (D), and mouse monoclonal anti-linear poly-ubiquitin antibody clone LUB9 (E). Images were visualized using a secondary antibody conjugated to HRP and a chemiluminescence detection system.
The LC-ESI-MS/MS analysis identified several ubiquitinated viral proteins, i.e., BV/ODV-C42, DNAPOL and PKIP, although this results should be further confirmed experimentally (Table 3). Ubiquitin exists in several linked forms, and each form plays distinct roles in the cell. Although BmNPV ubiquitin contains an additional and seven conserved lysines37, two linked forms, i.e., K48- and K63-linked poly-ubiquitin chains, were identified, whereas for the host ubiquitin, K6-, K11-, K29-, K48- and K63-linked forms were observed (Table 3). Notably, the identification and role exploration of these ubiquitinated proteins in baculovirus replication will be of interest in future investigations.
The presence of host cytoskeleton-related proteins, helicases, RNA binding and ribosomal proteins in the polyhedra matrix was not further verified due to a lack of corresponding antibodies. However, progress during the previous decade has revealed that certain viruses recruit numerous host proteins to facilitate each step of the viral replication process. Cytoskeleton-related proteins are one of the largest groups of cellular factors. The cellular actin cytoskeleton is involved in different stages of baculovirus infections17. This present study identified certain nucleocapsid proteins, such as P78/83, BV/ODV-C42 and VP80, to be components of the polyhedra matrix. The proteins P78/83 and BV/ODV-C42 are involved in actin polymerization and nucleocapsid trafficking147,148,149. VP80 interacts with filamentous actin103. Therefore, it is not surprising that actin became trapped in the polyhedra matrix. RNA-binding proteins bind to single or double stranded RNAs and play important roles in RNA splicing, export, stability, localization and translation150, thereby exerting significant control over numerous cellular functions. The identification of RNA binding proteins was reasonable due to the existence of certain small non-coding RNAs that could perform a structural function in polyhedra crystal formation in the BmNPV polyhedra matrix151,152,153. The present study is the first to report that certain DEAD/DEAH box helicases are components of the polyhedra matrix. The DEAD/DEAH box helicases are a family of proteins whose purpose is to unwind nucleic acids. The DEAD box helicases are involved in various aspects of RNA metabolism, including nuclear transcription, pre-mRNA splicing, ribosome biogenesis, nucleocytoplasmic transport, translation, RNA decay and organellar gene expression154, 155. The tomato bushy stunt virus recruits the host DEAD-box helicases Ded1p, Ded1 and RH20 to aid viral replication in infected cells, facilitate the maintenance of the full-length viral genome and suppress viral recombination, thus limiting the appearance of defective viral RNAs during replication156,157,158. Several ribosomal proteins were also identified to be incorporated into the polyhedra matrix. In addition to their roles in protein translation, some ribosomal proteins show extra-ribosomal functions, such as protein chaperone activity and the regulation of transcription159,160,161,162. An earlier proteomic analysis has identified the presence of the ribosomal proteins RPL18, RPL5, RPL3, and RPS6 in Ebola virions and demonstrated that reduced expression of each of these ribosomal proteins by RNA interference effectively inhibited Ebola infection in 293 T cells163. It would be interesting to investigate whether these identified host proteins are involved in the baculovirus life cycle.
Identification of viral components associated with the BmNPV polyhedra matrix
Baculovirus polyhedra contain ODVs that are embedded in the crystalline polyhedra matrix. The number of ODVs that are scattered with no apparent order generally ranges from 1 to 200164, implying that the polyhedra harbor flexible pools for encapsulating foreign proteins, including entities such as biological nanoparticles10. The genome of the BmNPV T3 strain contains 136 ORFs encoding predicted proteins of over 60 amino acids165. In total, 91 viral proteins with one or more peptides were identified to be associated with the polyhedra matrix of BmNPV (Table 2). Certain components were observed in more than one excised gel region. Among these proteins, most have been previously reported to be envelope (E)-, envelope-and-nucleocapsid (ENC)-, and/or nucleocapsid (NC)-associated components of BV and ODV in the literature. Apparently, most viral proteins localized in the nucleus were specifically or randomly trapped in the polyhedra matrix.
To further confirm that certain viral proteins that were associated with the ODV were encapsulated in the polyhedra matrix, Western blotting was performed. The results showed that the nucleocapsid-specific protein VP39 and certain ODV envelope-specific components, such as ODV-E66, P74, PIF1, PIF2, PIF3 and PIF5, were detected in the crystalline matrix fraction of the polyhedra (Fig. 4).
Certain proteins associated with ODV were detected in the polyhedra matrix. The polyhedra matrix (lane P) and ODV (lane V) fractions were loaded for the Western blot analyses. The antibodies used for the blotting were described previously17 and indicated to the right of each blot diagram. Lane M, the protein marker.
Notably, polyhedrin is a major component in the BmNPV polyhedra matrix. In this report, polyhedrin was identified in five excised gel sections by mass spectrometry. The Western blot analysis using a mouse monoclonal anti-polyhedrin antibody showed many bands with molecular weights larger and lower than the theoretical molecular size of polyhedrin of 28.8 kDa (Fig. 5). Polyhedrin is a well characterized protein. The presence of dodecameric or disulfide-linked octameric polyhedrin molecules in the polyhedra has been reported167, 168. Recently, polyhedrin was found to form aggregates and aggresomes in the cytoplasm of infected cells166, 169. Polyhedrin directly binds to Bombyx mori microtubule-associated protein 1-light chain 3 (BmLC3), an autophagosome marker, and is co-localized with BmLC3 to the isolation membrane of the autophagosome166. These findings suggest that polyhedrin may function as an autophagic adapter during the process of selective autophagy, which is regulated by post-translational modifications, such as phosphorylation and ubiquitination170, 171. The bands with molecular sizes larger than 28.8 kDa may represent aggregated and post-translationally modified polyhedrin, and the bands with molecular weights lower than 28.8 kDa may be partially degraded products of polyhedrin172. The involvement of polyhedrin in autophagy requires further clarification.
Western blot analysis of the protein polyhedrin in the polyhedra matrix. Total proteins of the polyhedra matrix fraction were subjected to SDS-PAGE and then Western blot using a mouse monoclonal anti-polyhedrin antibody166. Proteins were visualized using a goat anti-mouse secondary antibody conjugated to HRP and a chemiluminescence detection system. Lane M, the protein marker.
Among the 91 viral components, some proteins, such as FP25K and VLF-1, are involved in the transcription of very late genes, including polyhedrin 173, 174. Bm34 plays an important role in the transcription of vlf-1 and fp25K and indirectly controls the expression of polyhedrin 56. The deletion of these genes leads to a decreased production of polyhedra56, 57, 62, 66,67,68, 78. Although these proteins are important for polyhedra production, they may not be essential for the assembly of polyhedrin into polyhedra. This finding is similar to that observed in other proteins (Table 2) as shown in the literature by studies in which polyhedra formation occurred in the nucleus upon the disruption of individual genes.
The protein DNAPOL plays an important role in viral genome replication, and when deleted, the very late phase of infection will not proceed and undoubtedly prevent the emergence of polyhedra70. The other two proteins, i.e., PK1 and PKIP, should also be considered. PK1, which is a serine/threonine kinase, is a component of the viral very late gene transcription initiation complex175 and has been shown to regulate very late polyhedrin promoter expression176, 177. Deleting pk1 results in the failure of nucleocapsid assembly and polyhedra formation in the nucleus of infected cells41. PKIP is a protein that interacts with PK1 to stimulate the activity of PK1. A temperature-sensitive mutation in the pkip gene (amino acid A46 was mutated to T46) caused AcMNPV to lose its ability to form plaques and polyhedra at a non-permissive temperature of 33 °C upon the delayed synthesis of polyhedrin50. In the present study, PK1 and PKIP were identified by mass spectrometry (Table 2). Notably, further efforts are needed to better understand the roles of PK1 and PKIP in polyhedra formation.
BM134 acts as a fusion partner to incorporate eGFP into the polyhedra matrix
The usage of polyhedra to encapsulate foreign peptide fragments was first demonstrated by McLinden and coworkers178, rendering polyhedra as attractive immobilized platforms. The most common approach of constructing a recombinant baculovirus with improved insecticidal properties is to engineer a donor vector that over-expresses an insect-specific toxin. The over-expression of a toxin by a recombinant baculovirus rarely contributes sufficiently to the insecticidal activity179. However, the incorporation of a toxin protein into polyhedra is more effective because the toxin is thus directly delivered to its normal site of activity, i.e., the gut of the host, and is expressed in the cells of the host180. This strategy offers an alternative to producing baculovirus-based insecticides with improved properties.
This study paid much attention to three of these 91 viral proteins due to their potential applications as carriers of engineered polyhedra. These proteins were identified by mass spectrometry with quite a fewer peptides and higher coverages, and the deletion of their individual genes leads to no or a minor impact on BV production according to the literature. One protein was polyhedrin, which is a major component of the BmNPV polyhedra matrix. Over the past decade, polyhedrin and its partial sequences were widely used to be fused, and then the fusion protein was incorporated, along with the native polyhedrin, into polyhedra180,181,182,183,184. These immobilized proteins included insecticidal toxin Cry1Ac180, Cry1–5185,186,187 of Bacillus thuringiensis, and the insect-specific cyto-insectotoxin (Cit1a) from the venom of the Central Asian spider Lachesana tarabaevi 188. The Bioassays of these recombinant viruses showed that their speed of action and pathogenicity were enhanced. The N-terminal 179 amino acids of classical swine fever virus envelope glycoprotein E2 (E2ΔC) were successfully incorporated by polyhedrin into the polyhedra, and the immobilized E2ΔC was immunogenic182. Additionally, the eGFP was trapped in the polyhedra and, after one month of storage, remained fluorescent189. These studies demonstrate that polyhedrin is a strong carrier that can incorporate foreign proteins into polyhedra.
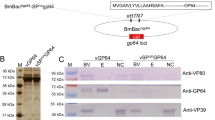
The other two proteins were ODV-E66 and BM134. ODV-E66 is an ODV envelope-specific protein58, and its N-terminal 23-amino-acid sequence is a hydrophobic domain that is sufficient to direct native and fusion proteins to be anchored on the viral envelope190. The deletion of odv-e66 had no effect on BV production, viral DNA replication and polyhedra formation in infected Sf9 cells but had an effect on oral infectivity59. BM134 is encoded by the ORF134 of BmNPV, and is a 109-amino-acid protein that is associated with the ODV144. BM134 is likely to be nonessential because when the ORF134 was deleted, the virus appeared normal191, 192. The lower molecular mass of 12.4 kDa may allow BM134 to be fused and then the fusion protein is incorporated into the BmNPV polyhedra. To confirm it, an ORF134-disrupted BmNPV bacmid was successfully generated via the Red/ET homologous recombination system in Escherichia coli as described previously193, 194 (Fig. 6A). The fusion protein BM134-eGFP-encoded nucleotide sequence and polyhedrin fragment were inserted into the polyhedrin locus by a site-specific transposon195, to produce the bacmids vBM134Re-eGFP and vBM134KO-Polh, respectively (Fig. 6B). These bacmids were transfected into BmN cells to obtain individual BV stocks. Equal MOI (10 TCID50/cell) of these viral stocks were used to co-infect BmN cells, and then, at 120 h post-infection (p.i.), the polyhedra were purified for the fluorescence visualization using a Leica TCS SP5 confocal laser scanning microscope. The results showed that the green fluorescence was observed on some polyhedra particles (Fig. 6C). The immobilization of eGFP into polyhedra was further confirmed by SDS-PAGE and a Western blot analysis using a mouse monoclonal anti-eGFP antibody. Theoretically, the molecular weight of the fusion protein BM134-eGFP is 39.6 kDa. A clear band with a molecular size larger than 39.6 kDa was observed (Fig. 6D), suggesting that the protein BM134 may undergo extensively post-translational modifications. The above results showed that BM134 could be used to immobilize foreign proteins into the polyhedra, and a protein of interest could be more effectively incorporated by fusion with the C-terminus of BM134.
BM134 as a carrier to incorporate eGFP into the polyhedra matrix. (A) PCR identification of the ORF134 knockout bacmid. Lane M, DNA marker. Lane 1, DNA template of wildtype BmNPV bacmid and lane 2, template of the ORF134-disrupted bacmid, vBM134KO. (B) Construction of the BM134-eGFP-encoded nucleotide sequence- and polyhedrin-inserted bacmids vBM134Re-eGFP and vBM134KO-Polh. (C) Fluorescence of the polyhedra purified from BmN cells co-infected with viral stocks of vBM134Re-eGFP and vBM134KO-Polh. (D) Western blot analysis of purified polyhedra immobilized with the fusion protein BM134-eGFP using a mouse monoclonal anti-eGFP antibody. Lane M, pre-stained protein marker.
Notably, the polyhedrin and BM134 could be used as carriers to immobilize double proteins into the polyhedra. If the ODV-E66 or its N-terminal sequence is demonstrated in the future to be an authentic carrier, polyhedra that trap triple proteins could be engineered.
In summary, the results presented here provided novel insight into the protein composition of the baculovirus polyhedra matrix and shed light on the cellular pathways that may play a structural role. The polyhedra displayed a powerful capacity to encapsulate foreign proteins as evidenced by the immobilization of the fusion protein BM134-eGFP in the polyhedra.
Methods
Cells, virus, and antibodies
The B. mori cell line BmN was cultured as previously described166. The BmNPV T3 isolate165 was propagated in BmN cells. The virus titer was determined by end-point dilution as previously described196 and was expressed as TCID50/ml.
Antibodies, such as a rabbit polyclonal antibody to HSC/HSP70 (sc-33575, Santa Cruz Biotechnology Inc., Santa Cruz, CA, USA), mouse monoclonal anti-ubiquitinylated proteins antibody clone FK1 (04–262) and FK2 (04–263), rabbit monoclonal anti-ubiquitin Lys48-specific clone Apu2 (05–1307) and Lys63-specific clone Apu3 (05–1308) from Millipore Corporation, Billerica, MA, USA, mouse monoclonal anti-linear poly-ubiquitin clone LUB9 (AB130, LifeSensors Inc., Malvern, PA, USA), mouse monoclonal antibody to BmNPV polyhedrin from Dr. Wen-Bin Wang of Jiangsu University, China, and rabbit polyclonal antibodies to AcMNPV VP39, ODV-E66, P74, PIF1, PIF2, PIF3 and PIF5 from Dr. Zhihong Hu of State Key Laboratory of Virology, Wuhan Institute of Virology, Chinese Academy of Sciences, Wuhan, People’s Republic of China, were used.
Polyhedra matrix protein preparation
Polyhedra were purified from infected BmN cells using a method proposed by Braunagel and Summers197 and then placed in a 70 °C water bath for 20 min. The protease inactivation was performed by suspending the purified polyhedra in a 10 mM HgCl2 solution198. The inhibitor HgCl2 was removed by washing the polyhedra with 10 mM Tris-HCl pH 7.6, 1 mM EDTA three times. ODVs were released by alkaline treatment and pelleted by continuous sucrose gradients199. The sample over the upper gradient was collected and added to equal volume of 2 × Laemmli buffer. The proteins were separated by 12% and 15% SDS-PAGE and stained using Coomassie brilliant blue R-250. The gel lane was excised into five contiguous sections spanning the complete gel lane based on a comparison with pre-stained molecular markers.
In-gel digestion, mass spectrometry and protein identification
The in-gel protein digestion, LC-ESI-MS/MS analysis using a Thermo Scientific Q Exactive mass spectrometer in data-dependent mode with an automatic switch between MS and MS/MS scans, data analysis were performed as previously described200, 201.
Western blot
The proteins in the polyhedra matrix and ODV fractions were separated by SDS-PAGE and transferred onto a PVDF membrane by wet electrophoresis transfer. Certain antibodies against HSC/HSP70, ubiquitin, polyhedrin, VP39, ODV-E66, P74, PIF1, PIF2, PIF3 and PIF5 were used as primary antibodies, and alkaline phosphatase or HRP-conjugated immunoglobulin G was applied as the secondary antibody. The signal was visualized using a BCIP/NBT development kit or a Tanon 5200 chemiluminescent imaging system (Tanon Science & Technology Co., Shanghai, China).
Generation of ORF134 knockout bacmid
The BmNPV bacmid was extracted from E. coli BmDH10Bac202 and then transformed into ElectroMaxTM DH10BTM cells (Invitrogen Life technologies, Carlsbad, CA, USA) by electroporation. The kanamycin-resistant, lacZ-positive colony was selected and transformed with the plasmid pRedET (Gene Bridges GmbH, Heidelberg, Germany). The colonies were isolated in the LB medium containing kanamycin and tetracycline, and designated E. coli BmDH10Bac-pRedET. A chloramphenicol resistance gene (cm) cassette with ORF134 flanking regions was amplified using the primers BM134Null-F (5′-CCAATAATATATTATGTATAGCACGTCAAAAATTAACAATGCGCGCTGAAAAGGGCGGCCGCGAAGTTCC-3′) and BM134Null-R (5′-TACGTCGCAAGCTATTTAGTTCGCGTTTTATTTGATCTCTATCATTCACTATAGGGCTCGAGGAAG-3′), and FRT-cm-FRT (Gene Bridges GmbH, Heidelberg, Germany) as the template. The underlined sequences are homologous to the upstream and downstream coding regions of ORF134. The disruption of ORF134 by cm was performed according to the protocol provided by the Quick & Easy Conditional Knockout Kit (Gene Bridges GmbH, Heidelberg, Germany). Colonies resistant to chloramphenicol, kanamycin and tetracycline were selected and verified by PCR using the primers BmORF134-F (5′-CGGATCCATGTATAGCACGTCAAAAAT-3′) (BamHI site underlined) and BmORF134-R (5′-ACTCGAGTACGGTGCATCTGCCATATT-3′) (XhoI site underlined). A positive colony containing the ORF134-disrupted bacmid vBM134KO was incubated at 42 °C to remove the plasmid pRedET and then chemically transformed with the helper plasmid pMON7124 to generate the E. coli BmDH10Bac-BM134KO/helper.
Construction of the polyhedrin-inserted and ORF134-rescued bacmids
The fragment at locus 128,008~1,446 nt of the BmNPV T3 isolate genome was amplified by PCR using the primers Polh-F (5′-AGAATTCCAATGTACCGCGCGGCG-3′) (EcoRI site underlined) and Polh-R (5′-ACTGCAGACCGCCTGCACCATCG-3′) (PstI site underlined) and ligated into EcoRI/PstI sites of plasmid pFast203. The resulting construct pFast-Polh was transformed into E. coli BmDH10Bac-BM134KO/helper to obtain the polyhedrin-inserted bacmid, which was named vBM134KO-Polh (Fig. 6B). ORF134 without the stop codon was amplified using the primers BmORF134-F and BmORF134-R and then inserted into the polyhedrin-removed pPph-Polyhedrin-eGFP166 to obtain pPph-ORF134-eGFP. This donor plasmid was transformed into E. coli BmDH10Bac-BM134KO/helper competent cells to produce the ORF134-repaired bacmid designated vBM134Re-eGFP (Fig. 6B). Positive transpositions were verified by PCR using the pUC/M13 forward and reverse primers.
Incorporation of eGFP into the polyhedra
Bacmids vBM134KO-Polh and vBM134Re-eGFP were transfected into BmN cells to obtain individual BV stocks. Equal MOI (10 TCID50/cell) of P2 viral stocks were used to co-infect BmN cells. At 120 h p.i., the polyhedra were purified for the fluorescence visualization under a Leica TCS SP5 confocal laser scanning microscope and a Western blot analysis using a mouse monoclonal anti-eGFP antibody.
References
Rohrmann, G. F. Ed. Baculovirus Molecular Biology. 9–22 (NCBI, 2013).
Blissard, G. W. Baculovirus-insect cell interactions. Cytotechnology 20, 73–93 (1996).
Slack, J. & Arif, B. M. The baculoviruses occlusion-derived virus: virion structure and function. Adv. Virus Res. 69, 99–165 (2007).
Friesen, P. D. Insect viruses. Fields virology. (ed. Knipe, D. M. & Howley, P. M.) (Lippincott-Williams & Wilkins, 2007).
Sajjan, D. B. & Hinchigeri, S. B. Structural organization of baculovirus occlusion bodies and protective role of multilayered polyhedron envelope protein. Food Environ. Virol. 8, 86–100 (2016).
Carstens, E. B., Krebs, A. & Gallerneault, C. E. Identification of an amino acid essential to the normal assembly of Autographa californica nuclear polyhedrosis virus polyhedra. J. Virol. 58, 684–688 (1986).
Katsuma, S. et al. Molecular characterization of baculovirus Bombyx mori nucleopolyhedrovirus polyhedron mutants. Arch. Virol. 144, 1275–1285 (1999).
Lin, G., Zhong, J. & Wang, X. Abnormal formation of polyhedra resulting from a single mutation in the polyhedrin gene of Autographa californica multicapsid nucleopolyhedrovirus. J. Inverteb. Pathol. 76, 13–19 (2000).
López, M. G., Alfonso, V., Carrillo, E. & Taboga, O. Description of a novel single mutation in the AcMNPV polyhedrin gene that results in abnormally large cubic polyhedra. Arch. Virol. 156, 695–699 (2011).
Ji, X. et al. How baculovirus polyhedra fit square pegs into round holes to robustly package viruses. EMBO J. 29, 505–514 (2010).
Chiu, E., Coulibaly, F. & Metcalf, P. Insect virus polyhedra, infectious protein crystals that contain virus particles. Curr. Opin. Struc. Biol. 22, 234–240 (2012).
Braunagel, S. C., Russell, W. K., Rosas-Acosta, G., Russell, D. H. & Summers, M. D. Determination of the protein composition of the occlusion-derived virus of Autographa californica nucleopolyhedrovirus. Proc. Natl. Acad. Sci. USA 100, 9797–9802 (2003).
Perera, O., Green, T. B., Stevens, S. M. Jr., White, S. & Becnel, J. J. Proteins associated with Culex nigripalpus nucleopolyhedrovirus occluded virions. J. Virol. 81, 4585–4590 (2007).
Liu, X., Chen, K., Cai, K. & Yao, Q. Determination of protein composition and host-derived proteins of Bombyx mori nucleopolyhedrovirus by 2-dimensional electrophoresis and mass spectrometry. Intervirology 51, 369–376 (2008).
Xu, F., Ince, İ. A., Boeren, S., Vlak, J. M. & van Oers, M. M. Protein composition of the occlusion derived virus of Chrysodeixis chalcites nucleopolyhedrovirus. Virus Res. 158, 1–7 (2011).
Deng, F. et al. Proteomics analysis of Helicoverpa armigera single nucleocapsid nucleopolyhedrovirus identified two new occlusion-derived virus-associated proteins, HA44 and HA100. J. Virol. 81, 9377–9385 (2007).
Hou, D. et al. Comparative proteomics reveal fundamental structural and functional differences between the two progeny phenotypes of a baculovirus. J. Virol. 87, 829–839 (2013).
Braconi, C. T. et al. Proteomic analyses of baculovirus Anticarsia gemmatalis multiple nucleopolyhedrovirus budded and occluded virus. J. Gen. Virol. 95, 980–989 (2014).
Hou, D., Chen, X. & Zhang, L. K. Proteomic analysis of Mamestra brassicae nucleopolyhedrovirus progeny virions from two different hosts. PLoS ONE 11, e0153365 (2016).
Wang, X. F. et al. ODV-associated proteins of the Pieris rapae granulovirus. J. Proteome Res. 10, 2817–2827 (2011).
Zhang, X., Liang, Z., Yin, X. & Shao, X. Proteomic analysis of the occlusion derived virus of Clostera anachoreta granulovirus. J. Gen. Virol. 96, 2394–2404 (2015).
Wang, R. et al. Proteomics of the Autographa californica nucleopolyhedrovirus budded virions. J. Virol. 84, 7233–7242 (2010).
Carbonell, L. F. & Miller, L. K. Baculovirus interaction with nontarget organisms: a virus-borne reporter gene is not expressed in two mammalian cell lines. Appl. Environ. Microbiol. 53, 1412–1417 (1987).
Lahaye, X., Vidy, A., Fouquet, B. & Blondel, D. Hsp70 protein positively regulates rabies virus infection. J. Virol. 86, 4743–4751 (2012).
Mayer, M. P. Recruitment of Hsp70 chaperone: a crucial part of viral survival strategies. Rev. Physiol. Biochem. Pharmacol. 153, 1–46 (2005).
Nagy, P. D., Wang, R. Y., Pogany, J., Hafren, A. & Makinen, K. Emerging picture of host chaperone and cyclophilin roles in RNA virus replication. Virology 411, 374–382 (2011).
Breitenbach, J. E. & Popham, H. J. Baculovirus replication induces the expression of heat shock proteins in vivo and in vitro. Arch. Virol. 158, 1517–1522 (2013).
Lyupina, Y. V. et al. An important role of the heat shock response in infected cells for replication of baculovirus. Virology 406, 336–341 (2010).
Iwanaga, M. et al. Involvement of HSC70-4 and other inducible HSPs in Bombyx mori nucleopolyhedrovirus infection. Virus Res. 179, 113–118 (2014).
Knox, C., Luke, G. A., Blatch, G. L. & Pesce, E. R. Heat shock protein 40 (Hsp40) plays a key role in the virus life cycle. Virus Res. 160, 15–24 (2011).
Franke, E. K., Yuan, H. E. & Luban, J. Specific incorporation of cyclophilin A into HIV-1 virions. Nature 372, 359–362 (1994).
Thali, M. et al. Functional association of cyclophilin A with HIV-1 virions. Nature 372, 363–365 (1994).
Hurley, J. H., Lee, S. & Prag, G. Ubiquitin-binding domains. Biochem. J. 399, 361–372 (2006).
Boname, J. M. & Lehner, P. J. Interactions between viruses and the ubiquitin-proteasome system. Protein degradation. Vol. 4: The ubiquitin-proteasome system and disease. (ed. Mayer, R. J., Ciechanover, A. J. & Rechsteiner, M.) 145–168 (WILEY-VCH Verlag GmbH & Co. KGaA, 2008).
Katsuma, S., Tsuchida, A., Matsuda-Imai, N., Kang, W. & Shimada, T. Role of the ubiquitin–proteasome system in Bombyx mori nucleopolyhedrovirus infection. J. Gen. Virol. 92, 699–705 (2011).
Xue, J. et al. Dynamic interactions between Bombyx mori nucleopolyhedrovirus and its host cells revealed by transcriptome analysis. J. Virol. 86, 7345–7359 (2012).
Guo, Z. J., Zhu, Y. M., Li, G. H., Chen, K. P. & Zhang, C. X. Ubiquitins of Bombyx mori nucleopolyhedrovirus and Helicoverpa armigera nucleopolyhedrovirus show distinct subcellular localization in infected cells. Acta Virol. 55, 101–106 (2011).
Russell, R. L. Q., Funk, C. J. & Rohrmann, G. F. Association of a baculovirus-encoded protein with the capsid basal region. Virology 227, 142–152 (1997).
Vialard, J. E. & Richardson, C. D. The 1,629-nucleotide open reading frame located downstream of the Autographa californica nuclear polyhedrosis virus polyhedrin gene encodes a nucleocapsid-associated phosphoprotein. J. Virol. 67, 5859–5866 (1993).
Li, A. et al. Posttranslational modifications of baculovirus protamine-like protein P6.9 and the significance of its hyperphosphorylation for viral very late gene hyperexpression. J. Virol. 89, 7646–7659 (2015).
Liang, C. et al. Autographa californica multiple nucleopolyhedrovirus PK-1 is essential for nucleocapsid assembly. Virology 443, 349–357 (2013).
Tao, X. Y. et al. Autographa californica multiple nucleopolyhedrovirus ORF11 is essential for budded-virus production and occlusion-derived-virus envelopment. J. Virol. 89, 373–383 (2015).
Kokusho, R., Koh, Y., Fujimoto, M., Shimada, T. & Katsuma, S. Bombyx mori nucleopolyhedrovirus BM5 protein regulates progeny virus production and viral gene expression. Virology 498, 240–249 (2016).
Katsuma, S. et al. Baculovirus-encoded protein BV/ODV-E26 determines tissue tropism and virulence in lepidopteran insects. J. Virol. 86, 2545–2555 (2012).
Beniya, H., Braunagel, S. C. & Summers, M. D. Autographa californica nuclear polyhedrosis virus: subcellular localization and protein trafficking of BV/ODV-E26 to intranuclear membranes and viral envelopes. Virology 240, 64–75 (1998).
Yang, Z. N. et al. Bombyx mori nucleopolyhedrovirus ORF9 is a gene involved in the budded virus production and infectivity. J. Gen. Virol. 90, 162–169 (2009).
Wang, Y. et al. ac18 is not essential for the propagation of Autographa californica multiple nucleopolyhedrovirus. Virology 367, 71–81 (2007).
Lung, O. Y., Cruz-Alvarez, M. & Blissard, G. W. Ac23, an envelope fusion protein homolog in the baculovirus Autographa californica multicapsid nucleopolyhedrovirus, is a viral pathogenicity factor. J. Virol. 77, 328–339 (2003).
Yu, L. L., Bray, D., Lin, Y. C. & Lung, O. Autographa californica multiple nucleopolyhedrovirus ORF 23 null mutant produces occlusion-derived virions with fewer nucleocapsids. J. Gen. Virol. 90, 1499–1504 (2009).
McLachlin, J. R., Yang, S. & Miller, L. K. A baculovirus mutant defective in PKIP, a protein which interacts with a virus-encoded protein kinase. Virology 246, 379–391 (1998).
Katsuma, S., Bando, H. & Shimada, T. Deletion analysis of a superoxide dismutase gene of Bombyx mori (Lepidoptera: Bombycidae) nucleopolyhedrovirus. Appl. Entomol. Zool. 50, 57–62 (2015).
Guarino, L. A., Smith, G. & Dong, W. Ubiquitin is attached to membranes of baculovirus particles by a novel type of phospholipid anchor. Cell 80, 301–309 (1995).
Gomi, S., Zhou, C. E., Yih, W., Majima, K. & Maeda, S. Deletion analysis of four of eighteen late gene expression factor gene homologues of the baculovirus, BmNPV. Virology 230, 35–47 (1997).
Wang, D., An, S. H., Guo, Z. J., Xu, H. J. & Zhang, C. X. Characterization of Helicoverpa armigera nucleopolyhedrovirus orf33 that encodes a novel budded virion derived protein, BV-e31. Arch. Virol. 150, 1505–1515 (2005).
Yu, M. & Carstens, E. B. Characterization of an Autographa californica multiple nucleopolyhedrovirus mutant lacking the ac39 (p43) gene. Virus Res. 155, 300–306 (2011).
Katsuma, S. & Shimada, T. Bombyx mori nucleopolyhedrovirus ORF34 is required for efficient transcription of late and very late genes. Virology 392, 230–237 (2009).
Tao, X. Y. et al. Functional characterization of Autographa californica multiple nucleopolyhedrovirus ORF43 and phenotypic changes of ORF43-knockout mutant. J. Microbiol. 51, 515–521 (2013).
Hong, T., Braunagel, S. C. & Summers, M. D. Transcription, translation, and cellular localization of PDV-E66: a structural protein of the PDV envelope of Autographa californica nuclear polyhedrosis virus. Virology 204, 210–222 (1994).
Xiang, X. et al. Autographa californica multiple nucleopolyhedrovirus odv-e66 is an essential gene required for oral infectivity. Virus Res. 158, 72–78 (2011).
Xu, H. J., Liu, Y. H., Yang, Z. N. & Zhang, C. X. Characterization of ORF39 from Helicoverpa armigera single-nucleocapsid nucleopolyhedrovirus, the gene containing RNA recognition motif. J. Biochem. Mol. Biol. 39, 263–269 (2006).
Marek, M., Romier, C., Galibert, L., Merten, O. W. & van Oers, M. M. Baculovirus VP1054 is an acquired cellular PURα, a nucleic acid-binding protein specific for GGN Repeats. J. Virol. 87, 8465–8480 (2013).
Lee, H. H. & Miller, L. K. Isolation, complementation, and initial characterization of temperature-sensitive mutants of the baculovirus Autographa californica nuclear polyhedrosis virus. J. Virol. 31, 240–252 (1979).
Olszewski, J. & Miller, L. K. Identification and characterization of a baculovirus structural protein, VP1054, required for nucleocapsid formation. J. Virol. 71, 5040–5050 (1997).
Li, L. et al. Characterization of Spodoptera exigua multicapsid nucleopolyhedrovirus ORF100 and ORF101, two homologues of E. coli ChaB. Virus Res. 121, 42–50 (2006).
Braunagel, S. C. et al. Mutations within the Autographa californica nucleopolyhedrovirus FP25K gene decrease the accumulation of ODV-E66 and alter its intranuclear transport. J. Virol. 73, 8559–8570 (1999).
Harrison, R. L. & Summers, M. D. Mutations in the Autographa californica multinucleocapsid nuclear polyhedrosis virus 25 kDa protein gene result in reduced virion occlusion, altered intranuclear envelopment and enhanced virus production. J. Gen. Virol. 76, 1451–1459 (1995).
Cheng, X. H. et al. Cell-dependent production of polyhedra and virion occlusion of Autographa californica multiple nucleopolyhedrovirus fp25k mutants in vitro and in vivo. J. Gen. Virol. 94, 177–186 (2013).
Nakanishi, T. et al. Comparative studies of Lepidopteran baculovirus-specific protein FP25K: development of a novel Bombyx mori nucleopolyhedrovirus-based vector with a modified fp25K gene. J. Virol. 84, 5191–5200 (2010).
Tian, C. H. et al. Bombyx mori nucleopolyhedrovirus ORF51 encodes a budded virus envelope associated protein. Virus Genes 38, 171–177 (2009).
Feng, G. et al. Selection and characterization of Autographa californica multiple nucleopolyhedrovirus DNA polymerase mutations. J. Virol. 86, 13576–13588 (2012).
Zhang, M. J. et al. Bombyx mori nucleopolyhedrovirus ORF54, a viral desmoplakin gene, is associated with the infectivity of budded virions. Arch. Virol. 157, 1241–1251 (2012).
Ke, J., Wang, J., Deng, R. & Wang, X. Autographa californica multiple nucleopolyhedrovirus ac66 is required for the efficient egress of nucleocapsids from the nucleus, general synthesis of preoccluded virions and occlusion body formation. Virology 374, 421–431 (2008).
Ke, J. et al. Characterization of AcMNPV with a deletion of ac69 gene. Microbiol. Res. 3, e4 (2011).
Hu, X. et al. Bm59 is an early gene, but is unessential for the propagation and assembly of Bombyx mori nucleopolyhedrovirus. Mol. Genet. Genomics 291, 145–154 (2016).
Wei, D. et al. Autographa californica nucleopolyhedrovirus Ac76: a dimeric type II integral membrane protein that contains an inner nuclear membrane-sorting motif. J. Virol. 88, 1090–1103 (2014).
Hu, Z. et al. Autographa californica multiple nucleopolyhedrovirus ac76 is involved in intranuclear microvesicle formation. J. Virol. 84, 7437–7447 (2010).
Yang, S. & Miller, L. K. Expression and mutational analysis of the baculovirus very late factor 1 (vlf-1) gene. Virology 245, 99–109 (1998).
Li, Y. et al. vlf-1 deletion brought AcMNPV to defect in nucleocapsid formation. Virus Genes 31, 275–284 (2005).
Wu, W. & Passarelli, A. L. The Autographa californica M nucleopolyhedrovirus ac79 gene encodes an early gene product with structural similarities to UvrC and intron-encoded endonucleases that is required for efficient budded virus production. J. Virol. 86, 5614–5625 (2012).
Dong, F., Wang, J., Deng, R. & Wang, X. Autographa californica multiple nucleopolyhedrovirus gene ac81 is required for nucleocapsid envelopment. Virus Res. 221, 47–57 (2016).
Iwanaga, M., Kurihara, M., Kobayashi, M. & Kang, W. Characterization of Bombyx mori nucleopolyhedrovirus orf68 gene that encodes a novel structural protein of budded virus. Virology 297, 39–47 (2002).
Russell, R. L. Q. & Rohrmann, G. F. Characterization of P91, a protein associated with virions of an Orgyia pseudotsugata baculovirus. Virology 233, 210–223 (1997).
Xiang, X. et al. Bombyx mori nucleopolyhedrovirus BmP95 plays an essential role in budded virus production and nucleocapsid assembly. J. Gen. Virol. 94, 1669–1679 (2013).
Zhu, S., Wang, W., Wang, Y., Yuan, M. & Yang, K. The baculovirus core gene ac83 is required for nucleocapsid assembly and per os infectivity of Autographa californica Nucleopolyhedrovirus. J. Virol. 87, 10573–10586 (2013).
Ishihara, G., Shimada, T. & Katsuma, S. Functional characterization of Bombyx mori nucleopolyhedrovirus CG30 protein. Virus Res. 174, 52–59 (2013).
Zhang, M. J. et al. Disruption of Bombyx mori nucleopolyhedrovirus ORF71 (Bm71) results in inefficient budded virus production and decreased virulence in host larvae. Virus Genes 45, 161–168 (2012).
Thiem, S. M. & Miller, L. K. Identification, sequence, and transcriptional mapping of the major capsid protein gene of the baculovirus Autographa californica nuclear polyhedrosis virus. J. Virol. 63, 2008–2018 (1989).
Wu, W. & Passarelli, A. L. Autographa californica multiple nucleopolyhedrovirus Ac92 (ORF92, P33) is required for budded virus production and multiply enveloped occlusion-derived virus formation. J. Virol. 84, 12351–12361 (2010).
Nie, Y., Fang, M. & Theilmann, D. A. Autographa californica multiple nucleopolyhedrovirus core gene ac92 (p33) is required for efficient budded virus production. Virology 409, 38–45 (2011).
Yuan, M. et al. Identification of Autographa californica nucleopolyhedrovirus ac93 as a core gene and its requirement for intranuclear microvesicle formation and nuclear egress of nucleocapsids. J. Virol. 85, 11664–11674 (2011).
Russell, R. L. Q. & Rohrmann, G. F. A 25-kDa protein is associated with the envelopes of occluded baculovirus virions. Virology 195, 532–540 (1993).
Chen, L. et al. Autographa californica multiple nucleopolyhedrovirus odv-e25 (Ac94) is required for budded virus infectivity and occlusion-derived virus formation. Arch. Virol. 157, 617–625 (2012).
Xu, H. J., Yang, Z. N., Wang, F. & Zhang, C. X. Bombyx mori nucleopolyhedrovirus ORF79 encodes a 28-kDa structural protein of the ODV envelope. Arch. Virol. 151, 681–695 (2006).
Dong, Z. Q. et al. Bombyx mori nucleopolyhedrovirus ORF79 is a per os infectivity factor associated with the PIF complex. Virus Res. 184, 62–70 (2014).
Fang, M., Nie, Y., Harris, S., Erlandson, M. A. & Theilmann, D. A. Autographa californica multiple nucleopolyhedrovirus core gene ac96 encodes a per os infectivity factor (pif-4). J. Virol. 83, 12569–12578 (2009).
Wu, W. et al. Autographa californica multiple nucleopolyhedrovirus 38K is a novel nucleocapsid protein that interacts with VP1054, VP39, VP80, and itself. J. Virol. 82, 12356–12364 (2008).
Wu, W. et al. Autographa californica multiple nucleopolyhedrovirus nucleocapsid assembly is interrupted upon deletion of the 38 K gene. J. Virol. 80, 11475–11485 (2006).
Wilson, M. E., Mainprize, T. H., Friesen, P. D. & Miller, L. K. Location, transcription, and sequence of a baculovirus gene encoding a small arginine-rich polypeptide. J. Virol. 61, 661–666 (1987).
Liu, X. et al. Distribution and phosphorylation of the basic protein P6.9 of Autographa californica nucleopolyhedrovirus. J. Virol. 86, 12217–12227 (2012).
Braunagel, S. C., Guidry, P. A., Rosas-Acosta, G., Engelking, L. & Summers, M. D. Identification of BV/ODV-C42, an Autographa californica nucleopolyhedrovirus orf101-encoded structural protein detected in infected-cell complexes with ODV-EC27 and p78/83. J. Virol. 75, 12331–12338 (2001).
Vanarsdall, A. L., Pearson, M. N. & Rohrmann, G. F. Characterization of baculovirus constructs lacking either the Ac 101, Ac 142, or the Ac 144 open reading frame. Virology 367, 187–195 (2007).
Yuan, M. et al. A highly conserved baculovirus gene p48 (ac103) is essential for BV production and ODV envelopment. Virology 379, 87–96 (2008).
Marek, M., Merten, O. W., Galibert, L., Vlak, J. M. & van Oers, M. M. Baculovirus VP80 protein and the F-actin cytoskeleton interact and connect the viral replication factory with the nuclear periphery. J. Virol. 85, 350–5362 (2011).
Müller, R., Pearson, M. N., Russell, R. L. & Rohrmann, G. F. A capsid-associated protein of the multicapsid nuclear polyhedrosis virus of Orgyia pseudotsugata: genetic location, sequence, transcriptional mapping, and immunocytochemical characterization. Virology 176, 133–144 (1990).
Chen, W. et al. Identification of Spodoptera litura multicapsid nucleopolyhedrovirus ORF97, a novel protein associated with envelope of occlusion-derived virus. Virus Genes 32, 79–84 (2006).
Tang, Q. et al. Bm91 is an envelope component of ODV but is dispensable for the propagation of Bombyx mori nucleopolyhedrovirus. J. Invertebr. Pathol. 113, 70–77 (2013).
Fang, M. et al. Open reading frame 94 of Helicoverpa armigera single nucleocapsid nucleopolyhedrovirus encodes a novel conserved occlusion-derived virion protein, ODV-EC43. J. Gen. Virol. 84, 3021–3027 (2003).
Fang, M., Nie, Y. & Theilmann, D. A. Deletion of the AcMNPV core gene ac109 results in budded virions that are non-infectious. Virology 389, 66–74 (2009).
Alfonso, V. et al. AcMNPV core gene ac109 is required for budded virion transport to the nucleus and for occlusion of viral progeny. PLoS ONE 7, e46146 (2012).
Lehiy, C. J., Wu, W., Berretta, M. F. & Passarelli, A. L. Autographa californica M nucleopolyhedrovirus open reading frame 109 affects infectious budded virus production and nucleocapsid envelopment in the nucleus of cells. Virology 435, 442–452 (2013).
Liang, G. et al. Bombyx mori nucleopolyhedrovirus ORF94, a novel late protein is identified to be a component of ODV structural protein. Curr. Microbiol. 61, 190–196 (2010).
Wei, W., Zhou, Y., Lei, C. & Sun, X. Autographa californica multiple nucleopolyhedrovirus orf114 is not essential for virus replication in vitro, but its knockout reduces per os infectivity in vivo. Virus Genes 45, 360–369 (2012).
Ohkawa, T., Washburn, J. O., Sitapara, R., Sid, E. & Volkman, L. E. Specific binding of Autographa californica M nucleopolyhedrovirus occlusion-derived virus to midgut cells of Heliothis virescens larvae is mediated by products of pif genes Ac119 and Ac022 but not by Ac115. J. Virol. 79, 15258–15264 (2005).
Li, X., Song, J., Jiang, T., Liang, C. & Chen, X. The N-terminal hydrophobic sequence of Autographa californica nucleopolyhedrovirus PIF-3 is essential for oral infection. Arch. Virol. 152, 1851–1858 (2007).
Song, J., Wang, R., Deng, F., Wang, H. & Hu, Z. Functional studies of per os infectivity factors of Helicoverpa armigera single nucleocapsid nucleopolyhedrovirus. J. Gen. Virol. 89, 2331–2338 (2008).
Lanier, L. M., Slack, J. M. & Volkman, L. E. Actin binding and proteolysis by the baculovirus AcMNPV: the role of virion-associated V-CATH. Virology 216, 380–388 (1996).
Wolgamot, G. M., Gross, C. H., Russell, R. L. Q. & Rohrmann, G. F. Immunocytochemical characterization of p24, a baculovirus capsid-associated protein. J. Gen. Virol. 74, 103–107 (1993).
Yang, M., Huang, C., Qian, D. D. & Li, L. L. Functional characterization of Autographa californica multiple nucleopolyhedrovirus gp16 (ac130). Virology 464-465, 341–352 (2014).
Gross, C. H., Russell, R. L. Q. & Rohrmann, G. F. Orgyia pseudotsugata baculovirus p10 and polyhedron envelope protein genes: analysis of their relative expression levels and role in polyhedron structure. J. Gen. Virol. 75, 1115–1123 (1994).
Yang, M., Wang, S., Yue, X. L. & Li, L. L. Autographa californica multiple nucleopolyhedrovirus orf132 encodes a nucleocapsid-associated protein required for budded-virus and multiply enveloped occlusion-derived virus production. J. Virol. 88, 12586–12598 (2014).
Han, Y. et al. The Bombyx mori nucleopolyhedrovirus Bm111 affects virulence but not virus replication. Curr. Microbiol. 69, 56–62 (2014).
Nakanishi, T., Shimada, T. & Katsuma, S. Characterization of a Bombyx mori nucleopolyhedrovirus mutant lacking both fp25K and p35. Virus Genes 41, 144–148 (2010).
Vlak, J. M. et al. Functional studies on the p10 gene of Autographa californica nuclear polyhedrosis virus using a recombinant expressing a p10-β-galactosidase fusion gene. J. Gen. Virol. 69, 765–776 (1988).
Williams, G. V., Rohel, D. Z., Kuzio, J. & Faulkner, P. A cytopathological investigation of Autographa californica nuclear polyhedrosis virus p10 gene function using insertion/deletion mutants. J. Gen. Virol. 70, 187–202 (1989).
Wang, L. et al. Characterization of a virion occlusion-defective Autographa californica multiple nucleopolyhedrovirus mutant lacking the p26, p10 and p74 genes. J. Gen. Virol. 90, 1641–1648 (2009).
Faulkner, P., Kuzio, J., Williams, G. V. & Wilson, J. A. Analysis of p74, a PDV envelope protein of Autographa californica nucleopolyhedrovirus required for occlusion body infectivity in vivo. J. Gen. Virol. 78, 3091–3100 (1997).
Liu, Y., de Jong, J., Nagy, É., Theilmann, D. A. & Krell, P. J. Nuclear translocation sequence and region in Autographa californica multiple nucleopolyhedrovirus ME53 that are important for optimal baculovirus production. J. Virol. 90, 3953–3965 (2016).
de Jong, J., Theilmann, D. A., Arif, B. M. & Krell, P. J. Immediate-early protein ME53 forms foci and colocalizes with GP64 and the major capsid protein VP39 at the cell membranes of Autographa californica multiple nucleopolyhedrovirus-infected cells. J. Virol. 85, 9696–9707 (2011).
de Jong, J., Arif, B. M., Theilmann, D. A. & Krell, P. J. Autographa californica multiple nucleopolyhedrovirus me53 (ac140) is a nonessential gene required for efficient budded-virus production. J. Virol. 83, 7440–7448 (2009).
Fang, M., Dai, X. & Theilmann, D. A. Autographa californica multiple nucleopolyhedrovirus EXON0 (ORF141) is required for efficient egress of nucleocapsids from the nucleus. J. Virol. 81, 9859–9869 (2007).
Dai, X., Stewart, T. M., Pathakamuri, J. A., Li, Q. & Theilmann, D. A. Autographa californica multiple nucleopolyhedrovirus exon0 (orf141), which encodes a RING finger protein, is required for efficient production of budded virus. J. Virol. 78, 9633–9644 (2004).
McCarthy, C. B., Dai, X., Donly, C. & Theilmann, D. A. Autographa californica multiple nucleopolyhedrovirus ac142, a core gene that is essential for BV production and ODV envelopment. Virology 372, 325–339 (2008).
Yang, Z. N., Xu, H. J., Park, E. Y. & Zhang, C. X. Characterization of Bombyx mori nucleopolyhedrovirus with a deletion of bm118. Virus Res. 135, 220–229 (2008).
Braunagel, S. C., He, H., Ramamurthy, P. & Summers, M. D. Transcription, translation, and cellular localization of three Autographa californica nuclear polyhedrosis virus structural proteins: ODV-E18, ODV-E35 and ODV-EC27. Virology 222, 100–114 (1996).
McCarthy, C. B. & Theilmann, D. A. AcMNPV ac143 (odv-e18) is essential for mediating budded virus production and is the 30th baculovirus core gene. Virology 375, 277–291 (2008).
Lapointe, R. et al. Characterization of two Autographa californica nucleopolyhedrovirus proteins, Ac145 and Ac150, which affect oral infectivity in a host-dependent manner. J. Virol. 78, 6439–6448 (2004).
Dickison, V. L., Willis, L. G., Sokal, N. R. & Theilmann, D. A. Deletion of AcMNPV ac146 eliminates the production of budded virus. Virology 431, 29–39 (2012).
Theilmann, D. A. et al. Characterization of a highly conserved baculovirus structural protein that is specific for occlusion-derived virions. Virology 218, 148–158 (1996).
Harrison, R. L., Sparks, W. O. & Bonning, B. C. The Autographa californica multiple nucleopolyhedrovirus ODV-E56 envelope protein is required for oral infectivity and can be functionally substituted by the Rachiplusia ou multiple nucleopolyhedrovirus ODV-E56. J. Gen. Virol. 91, 1173–1182 (2010).
Xiang, X. et al. The Bombyx mori nucleopolyhedrovirus (BmNPV) ODV-E56 envelope protein is also a per os infectivity factor. Virus Res. 155, 69–75 (2011).
Milks, M. L., Washburn, M. J., Willis, L. G., Volkman, L. E. & Theilmann, D. A. Deletion of pe38 attenuates AcMNPV genome replication, budded virus production, and virulence in Heliothis virescens. Virology 310, 224–234 (2003).
Li, Y. & Miller, L. K. Expression and localization of a baculovirus protein phosphatase. J. Gen. Virol. 76, 2941–2948 (1995).
Katsuma, S. et al. The baculovirus uses a captured host phosphatase to induce enhanced locomotory activity in host caterpillars. PLoS Pathog. 8, e1002644 (2012).
Lin, F. et al. Characterization of a late gene, ORF134, from Bombyx mori nucleopolyhedrovirus. Intervirology 54, 113–121 (2011).
Thrower, J. S., Hoffman, L., Rechsteiner, M. & Pickart, C. M. Recognition of the polyubiquitin proteolytic signal. EMBO J. 19, 94–102 (2000).
Newton, K. et al. Ubiquitin chain editing revealed by polyubiquitin linkage-specific antibodies. Cell 134, 668–678 (2008).
Ohkawa, T., Volkman, L. E. & Welch, M. D. Actin-based motility drives baculovirus transit to the nucleus and cell surface. J. Cell Biol. 190, 187–95 (2010).
Li, K. et al. The putative pocket protein binding site of Autographa californica nucleopolyhedrovirus BV/ODV-C42 is required for virus-induced nuclear actin polymerization. J. Virol. 84, 7857–7868 (2010).
Wang, Y. et al. Identification of a novel regulatory sequence of actin nucleation promoting factor encoded by Autographa californica multiple nucleopolyhedrovirus. J. Biol. Chem. 290, 9533–9541 (2015).
Keene, J. D. RNA regulons: coordination of post-transcriptional events. Nat. Rev. Genet. 8, 533–543 (2007).
Shirina, T. V., Bobrovskaia, M. T. & Kozlov, É. A. Small RNAs in Bombyx mori nuclear polyhedrosis virus polyhedra form complexes with polypeptide p14 and polyhedrin. Ukr. Biokhim. Zh. (1999) 82(6), 87–92 (2010).
Shirina, T. V., Bobrovskaia, M. T. & Kozlov, É. A. The search of miRNA genes in Bombyx mori nuclear polyhedrosis virus genomes regions complementary to the latest genes. Ukr. Biokhim. Zh. (1999) 83(5), 59–66 (2011).
Shirina, T. V., Gerashchenko, A. V., Bobrovskaia, M. T. & Kashuba, V. I. Identification of small RNA in polyhedra of Bombyx mori nuclear polyhedrosis virus. Ukr. Biochem. J. 86(2), 101–106 (2014).
Aubourg, S., Kreis, M. & Lecharny, A. The DEAD box RNA helicase family in Arabidopsis thaliana. Nucleic Acids Res. 27, 628–636 (1999).
de la Cruz, J., Kressler, D. & Linder, P. Unwinding RNA in Saccharomyces cerevisiae: DEAD-box proteins and related families. Trends Biochem. Sci. 24, 192–198 (1999).
Kovalev, N., Barajas, D. & Nagy, P. D. Similar roles for yeast Dbp2 and Arabidopsis RH20 DEAD-box RNA helicases to Ded1 helicase in tombusvirus plus-strand synthesis. Virology 432, 470–484 (2012).
Kovalev, N., Pogany, J. & Nagy, P. D. A co-opted DEAD-box RNA helicase enhances tombusvirus plus-strand synthesis. PLoS Pathog. 8, e1002537 (2012).
Chuang, C., Prasanth, K. R. & Nagy, P. D. Coordinated function of cellular DEAD-box helicases in suppression of viral RNA recombination and maintenance of viral genome integrity. PLoS Pathog. 11, e1004680 (2015).
Wool, I. G. Extraribosomal functions of ribosomal proteins. Trends Biochem. Sci. 21, 164–165 (1996).
Lindstrӧm, M. S. Emerging functions of ribosomal proteins in gene-specific transcription and translation. Biochem. Biophys. Res. Commun. 379, 167–170 (2009).
Warner, J. R. & McIntosh, K. B. How common are extraribosomal functions of ribosomal proteins? Mol. Cell 34, 3–11 (2009).
Kovacs, D. et al. Janus chaperones: assistance of both RNA-and protein-folding by ribosomal proteins. FEBS Lett. 583, 88–92 (2009).
Spurgers, K. B. et al. Identification of essential filovirion-associated host factors by serial proteomic analysis and RNAi screen. Mol. Cell. Proteomics 9, 2690–2703 (2010).
Ackermann, H. W. & Smirnoff, W. A. A morphological investigation of 23 baculoviruses. J. Invert. Pathol. 41, 269–280 (1983).
Gomi, S., Majima, K. & Maeda, S. Sequence analysis of the genome of Bombyx mori nucleopolyhedrovirus. J. Gen. Virol. 80, 1323–1337 (1999).
Guo, Z. J. et al. Characterization of aggregate/aggresome structures formed by polyhedrin of Bombyx mori nucleopolyhedrovirus. Sci. Rep. 5, 14601 (2015).
Rohrmann, G. F. Characterization of N-polyhedrin of two baculovirus strains pathogenic for Orgyia pseudotsugata. Biochemistry 16, 1631–1634 (1977).
Scharnhorst, D. W. & Weaver, R. F. Structural analysis of the matrix protein from the nuclear polyhedrosis virus of Heliothis zea. Virology 102, 468–472 (1980).
Sampieri, A., Luz-Madrigal, A., Zepeda, J. & Vaca, L. Identification of fragments from Autographa californica polyhedrin protein essential for self-aggregation and exogenous protein incorporation. BMC Biochem. 16, 5 (2015).
Wild, P. et al. Phosphorylation of the autophagy receptor Optineurin restricts Salmonella growth. Science 333, 228–233 (2011).
Liu, Z. et al. Ubiquitylation of autophagy receptor Optineurin by HACE1 activates selective autophagy for tumor suppression. Cancer Cell 26, 106–120 (2014).
Bae, S. M. et al. Hyper-enhanced production of foreign recombinant protein by fusion with the partial polyhedrin of nucleopolyhedrovirus. PLoS ONE 8, e60835 (2013).
Harrison, R. L., Jarvis, D. L. & Summers, M. D. The role of the AcMNPV 25K gene, “FP25,” in baculovirus polh and p10 expression. Virology 226, 34–46 (1996).
Yang, S. & Miller, L. K. Activation of baculovirus very late promoters by interaction with very late factor 1. J. Virol. 73, 3404–3409 (1999).
Mishra, G., Chadha, P. & Das, R. H. Serine/threonine kinase (pk-1) is a component of Autographa californica multiple nucleopolyhedrovirus (AcMNPV) very late gene transcription complex and it phosphorylates a 102 kDa polypeptide of the complex. Virus Res. 137, (147–149 (2008).
Fan, X., Thirunavukkarasu, K. & Weaver, R. F. Temperature-sensitive mutations in the protein kinase-1 (pk-1) gene of the Autographa californica nuclear polyhedrosis virus that block verylate gene expression. Virology 224, 1–9 (1996).
Mishra, G., Chadha, P., Chaudhury, I. & Das, R. H. Inhibition of Autographa californica nucleopolyhedrovirus (AcNPV) polyhedrin gene expression by DNAzyme knockout of its serine/threonine kinase (pk1) gene. Virus Res. 135, 197–201 (2008).
McLinden, J. H., Ploplis, V. A., Fraser, M. J. & Rosen, E. D. Expression of foreign epitopes on recombinant occlusion bodies of baculoviruses. Vaccine 10, 231–237 (1992).
Mäkelä, A. R. & Oker-Blom, C. The baculovirus display technology-an evolving instrument for molecular screening and drug delivery. Comb. Chem. High Throughput Screen. 11, 86–98 (2008).
Chang, J. H. et al. An improved baculovirus insecticide producing occlusion bodies that contain Bacillus thuringiensis insect toxin. J. Invertebr. Pathol. 84, 30–37 (2003).
Chen, L. et al. Immobilization of foreign protein in BmNPV polyhedra by fusion expression with partial polyhedrin fragments. J. Virol. Methods 194, 185–189 (2013).
Lee, K. S. et al. Production of classical swine fever virus envelope glycoprotein E2 as recombinant polyhedra in baculovirus-infected silkworm larvae. Mol. Biotechnol. 50, 211–220 (2012).
Je, Y. H. et al. Baculovirus expression vectors that incorporate the foreign protein into viral occlusion bodies. Biotechniques 34, 81–87 (2003).
Kim, B. Y. et al. Production of Aujeszky’s disease (pseudorabies) virus envelope glycoproteins gB and gC as recombinant polyhedra in baculovirus-infected silkworm larvae. J. Asia-Pac. Entomol. 15, 114–120 (2012).
Jung, M. P. et al. Insecticidal activity of recombinant baculovirus expressing both spider toxin isolated from Araneus ventricosus and Bacillus thuringiensis crystal protein fused to a viral polyhedrin. Entomol. Res. 42, 339–346 (2012).
Choi, J. Y. et al. Insecticidal activity of recombinant baculovirus co-expressing Bacillus thuringiensis crystal protein and Kunitz-type toxin isolated from the venom of bumblebee Bombus ignites. J. Asia-Pac. Entomol. 16, 75–80 (2013).
Shim, H. J. et al. NeuroBactrus, a novel, highly effective, and environmentally friendly recombinant baculovirus insecticide. Appl. Environ. Microbiol. 79, 141–149 (2013).
Ali, M. P., Kato, T. & Park, E. Y. Improved insecticidal activity of a recombinant baculovirus expressing spider venom cyto-insectotoxin. Appl. Microbiol. Biotechnol. 99, 10261–10269 (2015).
Cao, C. P. et al. An observation on immobilization of foreign protein in polyhedra of BmNPV. Sci. Sericulture 35, 790–795 (2009).
Hong, T., Summers, M. D. & Braunagel, S. C. N-terminal sequences from Autographa californica nuclear polyhedrosis virus envelope proteins ODV-E66 and ODV-E25 are sufficient to direct reporter proteins to the nuclear envelope, intranuclear microvesicles and the envelope of occlusion derived virus. Proc. Natl. Acad. Sci. USA 94, 4050–4055 (1997).
Ono, C. et al. Phenotypic grouping of 141 BmNPVs lacking viral gene sequences. Virus Res. 165, 197–206 (2012).
Kamita, S. G. et al. A baculovirus-encoded protein tyrosine phosphatase gene induces enhanced locomotory activity in a lepidopteran host. Proc. Natl. Acad. Sci. USA 102, 2584–2589 (2005).
Zhang, Y., Buchholz, F., Muyrers, J. P. P. & Stewart, A. F. A new logic for DNA engineering using recombination in Escherichia coli. Nat. Genet. 20, 123–128 (1998).
Muyrers, J. P. P., Zhang, Y., Testa, G. & Stewart, A. F. Rapid modification of bacterial artificial chromosomes by ET-recombination. Nucleic Acids Res. 27, 1555–1557 (1999).
Luckow, V. A., Lee, S. C., Barry, G. F. & Olins, P. O. Efficient generation of infectious recombinant baculoviruses by site-specific transposon-mediated insertion of foreign genes into a baculovirus genome propagated in Escherichia coli. J. Virol. 67, 4566–4579 (1993).
King, L.A. & Possee, R.D., Ed. The baculovirus expression system: a laboratory guide. (Chapman and Hall, 1992).
Braunagel, S. C. & Summers, M. D. Autographa californica nuclear polyhedrosis virus, PDV, and ECV viral envelopes and nucleocapsids: structural proteins, antigens, lipid and fatty acid profiles. Virology 202, 315–328 (1994).
Summers, M. D. & Smith, G. E. Trichoplusia ni granulosis virus granulin: a phenol-soluble, phosphorylated protein. J. Virol. 16, 1108–1116 (1975).
IJkel, W. F. J. et al. A novel baculovirus envelope fusion protein with a proprotein convertase cleavage site. Virology 275, 30–41 (2000).
Zhang, Y. et al. Comparative proteome analysis of multi-layer cocoon of the silkworm, Bombyx mori. PLoS ONE 10, e0123403 (2015).
Guo, X. et al. Proteins in the cocoon of silkworm inhibit the growth of Beauveria bassiana. PLoS ONE 11, e0151764 (2016).
Motohashi, T., Shimojima, T., Fukagawa, T., Maenaka, K. & Park, E. Y. Efficient large-scale protein production of larvae and pupae of silkworm by Bombyx mori nuclear polyhedrosis virus bacmid system. Biochem. Biophys. Res. Commun. 326, 564–569 (2005).
Guo, Z. J. et al. Open reading frame 60 of the Bombyx mori nucleopolyhedrovirus plays a role in budded virus production. Virus Res. 151, 185–191 (2010).
Acknowledgements
We would like to thank Dr. Wen-Bin Wang from the School of Medical Science and Laboratory Medicine, Jiangsu University for kindly providing the mouse monoclonal anti-polyhedrin antibody, Dr. Zhihong Hu from the State Key Laboratory of Virology, Wuhan Institute of Virology, Chinese Academy of Sciences, Wuhan, People’s Republic of China for offering the rabbit polyclonal antibodies against AcMNPV VP39, ODV-E66, P74, PIF1, PIF2, PIF3 and PIF5, and Prof. Enoch Y. Park from the Department of Applied Biological Chemistry, Faculty of Agriculture, Shizuoka University, Japan for supplying the BmNPV BAC-TO-BAC system. This research study was supported by grants from the National Nature Science Foundation of China (No. 31370184, No. 31101767), the Nature Science Foundation of Jiangsu Province, China (No. BK20161365) and a project funded by the Priority Academic Program Development (PAPD) of Jiangsu Higher Education Institutions.
Author information
Authors and Affiliations
Contributions
Z-J.G., X.-D.T. and W.-L.W. conceived and designed these experiments and wrote the paper. M.-H.Y., X.-Y.D., T.T. and X.-Y.Y. contributed material and reagents, and performed the experiments. Z.-J.G., M.-H.Y. and X.-Y.D. analyzed the data. All authors reviewed the manuscript.
Corresponding authors
Ethics declarations
Competing Interests
The authors declare that they have no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Guo, ZJ., Yu, MH., Dong, XY. et al. Protein composition analysis of polyhedra matrix of Bombyx mori nucleopolyhedrovirus (BmNPV) showed powerful capacity of polyhedra to encapsulate foreign proteins. Sci Rep 7, 8768 (2017). https://doi.org/10.1038/s41598-017-08987-8
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-017-08987-8
- Springer Nature Limited
This article is cited by
-
Biotechnological applications of occlusion bodies of Baculoviruses
Applied Microbiology and Biotechnology (2018)