Abstract
Localized in the plasma membrane, death domain-containing TNF-related apoptosis-inducing ligand (TRAIL) receptors, TRAIL-R1 and TRAIL-R2, induce apoptosis and non-apoptotic signaling when crosslinked by the ligand TRAIL or by agonistic receptor-specific antibodies. Recently, an increasing body of evidence has accumulated that TRAIL receptors are additionally found in noncanonical intracellular locations in a wide range of cell types, preferentially cancer cells. Thus, besides their canonical locations in the plasma membrane and in intracellular membranes of the secretory pathway as well as endosomes and lysosomes, TRAIL receptors may also exist in autophagosomes, in nonmembraneous cytosolic compartment as well as in the nucleus. Such intracellular locations have been mainly regarded as hide-outs for these receptors representing a strategy for cancer cells to resist TRAIL-mediated apoptosis. Recently, a novel function of intracellular TRAIL-R2 has been revealed. When present in the nuclei of tumor cells, TRAIL-R2 inhibits the processing of the primary let-7 miRNA (pri-let-7) via interaction with accessory proteins of the Microprocessor complex. The nuclear TRAIL-R2-driven decrease in mature let-7 enhances the malignancy of cancer cells. This finding represents a new example of nuclear activity of typically plasma membrane-located cytokine and growth factor receptors. Furthermore, this extends the list of nucleic acid targets of the cell surface receptors by pri-miRNA in addition to DNA and mRNA. Here we review the diverse functions of TRAIL-R2 depending on its intracellular localization and we particularly discuss the nuclear TRAIL-R2 (nTRAIL-R2) function in the context of known nuclear activities of other normally plasma membrane-localized receptors.
Similar content being viewed by others
Facts
-
Plasma membrane-located TRAIL-R1/R2 induce both cell death and non-apoptotic signaling pathways.
-
Tumor cells show elevated intracellular levels of TRAIL-R1/R2.
-
Nuclear TRAIL-R2 interacts with the Microprocessor complex, inhibits let-7 maturation and enhances tumor cell proliferation.
Open Questions
-
What is the mechanism of nuclear import/export of TRAIL-R2 and how is this regulated?
-
Which miRNAs, beside let-7, are regulated by nTRAIL-R2?
-
What are additional specific functions of nTRAIL-R2? Is there any impact of other TRAIL-Rs (TRAIL-R1, -R3 and -R4) on nuclear localization and function of TRAIL-R2?
-
Do TRAIL-R1, TRAIL-R3 and TRAIL-R4 possess independent specific nuclear activities?
-
Is there a specific function of cytoplasmic TRAIL death receptors?
The death ligand TNF-related apoptosis-inducing ligand (TRAIL), a member of the TNF cytokine superfamily, interacts with five different receptors, plasma membrane-expressed TRAIL-R1-4 and a soluble receptor, osteoprotegerin (OPG), all members of the TNF receptor superfamily.1, 2, 3, 4, 5, 6, 7, 8 Two TRAIL receptors, TRAIL-R1/DR4 and TRAIL-R2/DR5/TRICK/Killer, contain a cytoplasmic death domain (DD) and are hence able to induce apoptosis.1, 2, 3 Therefore, they are also referred to as the TRAIL death receptors.
TRAIL has governed particular interest because of its apparent tumor cell-specific apoptosis-inducing capacity.9, 10 Consequently, TRAIL and agonistic TRAIL death receptor-specific antibodies are regarded as promising therapeutic agents for treatment of various malignancies and are currently under clinical investigation.11, 12, 13 However, many tumor cells are resistant to TRAIL-mediated apoptosis and, furthermore, TRAIL death receptors may induce pro-inflammatory, invasion- and metastasis-promoting non-apoptotic signaling pathways under certain conditions.14, 15 These observations point to the necessity to improve TRAIL-R1/R2-targeting therapies by combination with molecules sensitizing to cell death and at the same time inhibiting potentially harmful non-apoptotic signaling.
Overexpression of TRAIL-R1 and/or TRAIL-R2 has been demonstrated for many tumor entities (Table 1).16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29 Importantly, these studies detected mainly an intracellular localization of the receptors, suggesting that intracellular TRAIL-R1 and TRAIL-R2 may provide a growth advantage or other benefit for tumor cells. Indeed, correlation of the expression status with clinical parameters disclosed prevalent prognostic significance of high death receptor expression, identifying predominantly TRAIL-R2 and occasionally also TRAIL-R1 as negative prognostic markers (Table 1). For example, high TRAIL-R2 expression correlated with higher tumor grade, positive nodal status and shortened survival of breast cancer patients.20, 21 Strong TRAIL-R2 expression also negatively correlated with disease-free survival of patients suffering from metastatic HNSCC.23 For stage I and II NSCLC, high TRAIL-R2 expression correlated with poorly differentiated tumors and significantly reduced patient survival.27 Similarly, poorly differentiated areas of stage III NSCLC showed high expression of TRAIL-R1 and TRAIL-R2, but only TRAIL-R2 positivity was associated with an increased risk of death.28 Moreover, pancreatic cancer cells with stem cell characteristics also express TRAIL-R2 at increased levels.30 Correspondingly, high intracellular expression of TRAIL-R2 has been demonstrated for stem cells at the base of the intestinal crypts in the small intestine.31
Reports of positive correlation between receptor expression and better prognosis are occasionally also found. Thus, TRAIL-R1 was identified as an independent prognostic marker for a more favorable course of colorectal carcinoma.18, 19 In bladder cancer, higher levels of either TRAIL-R1 or TRAIL-R2 correlated with longer recurrence-free survival.29 In glioblastoma multiforme, high expression of both TRAIL-R1 and TRAIL-R2, of note mainly expressed at the cell surface, was found to be associated with longer overall survival of the respective patients.22
So far, mainly the overall tissue expression level of TRAIL receptors and occasionally also their presence at the cell surface were taken into account when estimating their values as a prognostic marker for cancer. However, recent discoveries on TRAIL-R2 functions revealed that even though an enormous body of information on death receptors has been gathered, our knowledge of the cellular functions of these receptors is still incomplete.16 The authors report that nuclear TRAIL-R2 (nTRAIL-R2) is engaged in microRNA (miRNA) maturation, thus performing functions entirely different from its classical role as a plasma membrane-located signal transducer. With this finding, the frequently observed high intracellular expression of TRAIL-Rs in tumors gains importance and demands careful re-evaluation.
Here we review the known functions of human TRAIL death receptors with respect to their subcellular compartmentalization. The novel nuclear functions of TRAIL-R2 will be discussed in comparison to nuclear functions of other conventionally plasma membrane-located cytokine and growth factor receptors.
TRAIL Receptors and Their Classical Functions at the Plasma Membrane
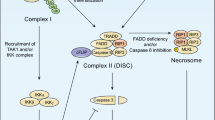
The best understood function of TRAIL-R1 and TRAIL-R2 is the induction of apoptosis (reviewed in Gonzalvez and Ashkenazi32 and see Figure 1). Upon TRAIL ligation, TRAIL death receptors cluster into higher multimeric complexes and gain the capability of assembling a DISC (death-inducing signaling complex) at their intracellular DD. Through this domain, TRAIL-R1 and TRAIL-R2 recruit the adaptor protein FADD (Fas-associated protein with death domain) that in turn, via death effector domain (DED), recruits the inactive pro-forms of the initiator caspases pro-caspase-8 and -10. Proximity-induced self-cleavage of pro-caspases within the DISC leads to their activation and subsequent dissociation from the membrane-localized receptor complexes.
Functions of plasma membrane-localized TRAIL death receptors. Following binding of their ligand, TRAIL death receptors TRAIL-R1 and TRAIL-R2 are able to induce cell death (apoptosis or necroptosis). Alternatively, these receptors may also enhance cell proliferation, inflammation, migration and invasion via activation of multiple ‘non-death-inducing’ signal transduction pathways. LMP, lysosomal membrane permeabilization; MOMP, mitochondrial outer membrane permeabilization
Besides death receptors, TRAIL can also bind to two additional plasma membrane-bound receptors, TRAIL-R3 and TRAIL-R4. TRAIL-R3 is a glycosylphosphatidylinositol-anchored receptor lacking transmembrane and cytoplasmatic domains and is therefore not capable of inducing intracellular signaling.5 TRAIL-R4 is more homologous to the TRAIL death receptors, but because of a truncated DD, it is unable to induce apoptosis.4, 6
Efficiency of the initiator caspase activation may be influenced at the DISC level, for example, by variation of its composition. It has been shown that the complex formation between death receptors TRAIL-R1 and TRAIL-R2 and the apoptosis-incompetent receptors, TRAIL-R3 or TRAIL-R4, leads to impaired DISC function.33, 34 In addition, recruitment of the proteolytically inactive caspase homolog cFLIP (cellular FLICE-like inhibitory protein) represents an apoptosis-inhibiting mechanism.35, 36
In some cells, activated initiator caspases may directly activate the effector caspases −3, −6 and −7, which in turn cleave a broad range of protein substrates and execute cell death. Such cells have been classified as the so-called type I cells with respect to apoptosis induction.37 In contrast, type II cells use the loop way via engagement of the intrinsic apoptosis pathway to amplify the death signal generated at the DISC. In these cells, caspase-8-mediated cleavage of the BH3-only protein Bid generates its truncated form (tBid) that, by association with Bax and Bak, leads to the mitochondrial outer membrane permeabilization (MOMP) and to the release of pro-apoptotic mitochondrial proteins like cytochrome C, Smac/Diablo (second mitochondria-derived activator of caspase/direct IAP binding protein with low pI) and HtrA2/Omi (high temperature requirement protein A2/Omi) into the cytosol. Cytosolic cytochrome C together with ATP and Apaf-1 (apoptotic protease activating factor-1) form a second caspase-inducing platform, the Apoptosome, that mediates the activation of caspase-9. Caspase-9, in turn, acts as an initiator caspase activating the effector caspases.
An alternative TRAIL-induced apoptosis pathway has been described in hepatocytes (Figure 1).38 In these cells, TRAIL-R2 is internalized upon TRAIL stimulation by endocytosis, targeted via vesicular transport to lysosomes that become permeabilized and release cathepsin B, subsequently promoting cell demise. Lysosomal membrane permeabilization (LMP) as a precondition for the execution of apoptosis in these cells requires the PACS-2 (phosphofurin acidic cluster sorting protein 2)-mediated recruitment of Bim and Bax to lysosomal membranes.39
Under conditions when caspases are inhibited, plasma membrane-expressed TRAIL death receptors may also induce an alternative death pathway known as necroptosis. For this pathway, a pivotal role of the kinases RIP-1 (receptor-interacting protein-1), RIP-3 as well as pseudo-kinase MLKL (mixed lineage kinase domain-like) has been shown,40, 41 but the exact molecular mechanisms of TRAIL-induced necroptosis are still not fully understood.
Over the past years, it has become increasingly evident that TRAIL-R1 and TRAIL-R2, besides death signaling pathways, can also activate multiple non-cell death signal transduction pathways like NF-κB (nuclear factor κ-light-chain-enhancer of activated B cells), PKC (protein kinase C), PI3K (phosphatidylinositide 3-kinases), AKT/PKB, Src (Rous sarcoma virus oncogene cellular homolog) and TAK1 (transforming growth factor-β activated kinase 1), as well as different members of the MAP-kinase family like ERK1/ERK2 (extracellular regulated kinases), JNK (c-Jun N-terminal kinase) and p38 (Figure 1).14, 42, 43, 44, 45, 46, 47, 48, 49, 50, 51
Regarding the molecular determinants for TRAIL-mediated activation of cell death or non-apoptotic pathways, the engagement of different protein complexes has been proposed. In this respect, non-apoptotic pathways have been suggested to be activated within a secondary signaling complexes formed downstream of the DISC.52 Besides FADD and pro-capase-8, the secondary signaling complex also include specific adaptor proteins like RIP1, TRAF2 and Nemo.
Different localizations of aggregated TRAIL receptors within specific plasma membrane domains have also been proposed to determine the formation and activity of ligand-induced TRAIL-R1/R2 signaling complexes. Hence, aggregated TRAIL receptors localized in lipid rafts were shown to be involved in the apoptotic signaling, whereas those localized outside lipid rafts form complexes capable of non-apoptotic signaling.53
Alternative to the MAPKs activation via secondary protein complexes, their stimulation via effector caspases was described in prostate cancer cells. Here, two different proteolytic fragments of Mst1 (mammalian sterile 20-like kinase 1) generated upon cleavage by caspases-3 or -7 selectively activate ERK or JNK and p38, respectively.54
This multiplicity and heterogeneity of TRAIL-induced pathways, as briefly outlined, is responsible for the observed broad cellular effects of this cytokine ranging from the induction of cell death, establishment of a death-resistant phenotype and expression of pro-inflammatory molecules promoting cell proliferation, migration and invasion (Figure 1). Particularly in apoptosis-resistant tumor cells, TRAIL-induced pro-inflammatory pathways may result in enhanced invasion and metastasis in vivo.42, 55, 56, 57 A comprehensive overview of TRAIL-induced non-apoptotic signaling pathways and their functional relevance in different cellular systems is provided in the recent review by Azijli et al.15
Cytoplasmic Locations and Functions of TRAIL Death Receptors
In addition to their plasma membrane expression, TRAIL-R1 and TRAIL-R2 are also often found in the cytoplasm of several cell types (Tables 1 and 2).16, 17, 18, 19, 20, 21, 22, 27, 28, 58, 59, 60, 61, 62, 63 However, in most cases the cytoplasmic localization of TRAIL death receptors was not resolved in detail. Thus, it still remained unclear whether the intracellular staining in cancer cells observed by immunohistochemistry reflects localization of TRAIL receptors in the membranes of organelles within the secretory and/or endocytotic pathway or whether a non-membraneous cytosolic localization of the receptors was also detected in these cases. The presence of TRAIL-R1 and TRAIL-R2 in soluble cytosolic fractions of diverse tumor cells was demonstrated recently.16 As the subcellular fractionation was performed without the use of any detergent, it can be concluded that these receptors are not anchored in intracellular membranes, but are present in the non-membraneous cytosolic compartment. The function of these soluble TRAIL death receptors remains to be shown.
In colon carcinoma, the cytoplasmic staining of TRAIL death receptors was interpreted as the presence of these receptors in the trans-Golgi network,64 a finding previously described for melanoma cells.65 The study on apoptosis-resistant breast cancer cells demonstrated the presence of TRAIL receptors in autophagosomes rather than the plasma membrane.66 Treatment of these cells with different inhibitors of autophagy led to a re-appearance of both TRAIL death receptors on the cell surface and sensitized these cells toward TRAIL-mediated death. A correlation of apoptosis resistance with intracellular presence of death receptors in breast cancer cells was also proposed in another study.59 Finally, as internalization of TRAIL death receptors in response to TRAIL treatment has been demonstrated in cell lines, the cytoplasmic localization of TRAIL receptors in tumor tissue could be caused by an autocrine or paracrine activation of their internalization by stroma and/or tumor cell-derived TRAIL. So far, however, this remains speculation and needs further investigation. Summing up, despite the variety of intracellular TRAIL death receptor locations, the frequently observed loss of TRAIL death receptors at the cell surface in tumors seems to be a general principle for the acquisition of TRAIL resistance in cancer cells. In addition, high cytoplasmic levels of these receptors as well as their presence in soluble form suggest specific pro-tumoral functions in this location.
TRAIL Death Receptors in the Nucleus
Few relatively recent studies show the localization of TRAIL receptors in the nucleus (Tables 1 and 2).16, 27, 31, 59, 65, 67, 68, 69 An early report on the melanoma cell line, Mel-FH, demonstrated TRAIL-R1 as predominantly expressed in the nucleus, where it colocalized with TRAIL-R3 and TRAIL-R4.65 Nuclear presence of TRAIL-R1 and TRAIL-R2 has also been shown for normal and psoriatic skin, with nTRAIL-R2 being more strongly expressed in psoriatic lesions and in the dermis of psoriatic skin compared with healthy skin.70 In the same study, CD4+ and CD8+ cells of the inflammatory infiltrate showed exclusive nuclear staining for TRAIL-R1. Nuclear presence of TRAIL-R2 was also described for nonmalignant, normal lung bronchial epithelial cells in basal nuclei.27 However, staining of these cell nuclei was in general weak and restricted to a small number of cells.
Only very few studies have addressed the functional relevance of nTRAIL-R2. Again, as for the presence of TRAIL-R1/R2 in the cytoplasm, the nuclear sequestration has been proposed as an apoptosis-resistance mechanism.67, 69 Trophoblast cells of the human placenta were found resistant to TRAIL-mediated apoptosis and expressed high levels of TRAIL-R2 that was mainly localized in the nucleus.67 Following stimulation with TNFα, TRAIL-R2 relocated to the plasma membrane and the cells became TRAIL sensitive. Another study demonstrated that inhibition of the nuclear import of TRAIL-R2 sensitized HeLa and HepG2 cells to TRAIL-induced apoptosis.69
Two very recent studies provide hints for surprizing and entirely novel functions of nTRAIL-R. In one report, the pattern of intracellular distribution of TRAIL-R during the development of the mouse CNS was studied. The authors demonstrated that the expression of TRAIL-R was not only restricted to proliferating zones but, importantly, was mainly located in the cell nuclei, suggesting a specific, proliferation-associated function of nTRAIL-R.31 Another study investigating human nuclear TRAIL receptors finally uncovered specific functions of nTRAIL-R2 connecting this receptor to the process of miRNA maturation (Figure 2).16 Briefly, in several different tumor cells, nTRAIL-R2 interacts with the core Microprocessor complex Drosha/DGCR8 and with several of its accessory proteins, p68, NF45 and hnRNPA1.16 Through this interaction, nTRAIL-R2 inhibits maturation of the miRNA let-7 and consequently increases the levels of let-7 targets Lin28B and HMGA2. HMGA2 and Lin28B proteins are highly expressed in embryonic tissues, downregulated in differentiated tissues and, because of low expression of let-7, re-expressed in tumors, where their levels positively correlate with tumor progression.71, 72, 73 In this scenario, inhibition of let-7-maturation by nTRAIL-R2 enhances the malignancy of tumor cells. In line with this notion, it was observed that TRAIL-R2 was expressed at higher levels in cell nuclei of PDAC tissue than in nuclei of peritumoral normal duct cells. Moreover, high nTRAIL-R2 correlated with high nuclear HMGA2 expression in the same tissue and with worse prognosis of patients with early-stage PDAC.16 Furthermore, knockdown of TRAIL-R2 resulted in decreased growth of experimental orthotopic tumors in mice, and progressive decrease of TRAIL-R2 expression as well as its exclusion from the nuclei accompanied the differentiation of pancreatic tumor cells into three-dimensional duct-like structures in vitro.16 The correlation of nuclear localization of TRAIL receptors and survival of cancer patients was analyzed, to the best of our knowledge, in only one additional report. In a study on NSCLC tumors from patients treated with chemotherapy, all four TRAIL receptors were detected in the nuclei and cytoplasm.68 In these patients, however, high nuclear and cytoplasmic expression of TRAIL-R2 correlated with improved survival. However, it is well known that chemotherapeutic agents upregulate TRAIL-R2 in cells with wild-type p53. Unfortunately, the p53 status was not analyzed in this study and it may be possible that the observed positive correlation simply reflects the better prognosis of patients with wild-type p53. In agreement with this interpretation, other studies on untreated lung cancer patients,27, 28 as well as patients suffering from other cancers, have instead found negative correlations between high overall expression of TRAIL-R2 and the survival time of the patients.
Nuclear functions of TRAIL-R2. Nuclear TRAIL-R2 interacts with the core Microprocessor complex and its accessory proteins. This inhibits maturation of the let-7 miRNA and results in an increase of the let-7 targets HMGA2 and Lin28B, finally enhancing cell proliferation. The mechanism of the TRAIL-R2 entrance into the nucleus is still not known. The importin-β-mediated transport through nuclear pores or an alternative pathway through Sec61 translocons known to be responsible for the nuclear import of other plasma membrane receptors are possible
The nuclear presence of TRAIL death receptors represents a new example of plasma membrane receptors that can localize to the nucleus, where they elicit functions distinct from their canonical signal transduction pathways that originate at the plasma membrane. Most prominently, the receptor tyrosine kinases (RTKs), and among them the members of the epidermal growth factor receptor (EGFR) family, have been found to shuttle to the nucleus in response to ligand binding (reviewed by Carpenter and Liao74 and Wang et al.75). Here, via interaction with classical transcription factors or with RNA helicase A (RHA), nuclear EGFRs activate the expression of several target genes.76, 77, 78, 79, 80, 81 In addition, via interaction with RNA-polymerase I, EGFR2 increases rRNA expression and cellular translation.82 EGFRs may also phosphorylate nuclear proteins such as PCNA,83 and DNA-dependent protein kinase (DNA-PK),84, 85, 86 thus influencing a variety of other nuclear processes like DNA replication and DNA damage repair.
Other RTKs found in the nucleus are fibroblast growth factor receptors (FGF-R1, -R2 and -R3; reviewed by Bryant and Stow87) vascular endothelial growth factor receptors (VEGF-R1, see Cai et al.88 and VEGF-R2, see Shay-Salit et al.89) insulin receptor,90 cMet,91 TrkA,92 growth hormone receptor93 and receptor tyrosine kinase-like orphan receptor 1 (Ror1).94 Particularly well understood are functions of nuclear FGF-R1 that, via its interaction with 90-kDa ribosomal S6 kinase (RSK1), activates the transcription factor CREB, thus regulating the transcription of genes involved in proliferation and differentiation.95 As RSK1 phosphorylates several transcription factors as well as histone H3, the ability to activate RSK1 enables nuclear FGF-R1 to regulate a plethora of biological processes.
Besides RTKs, other types of cell surface transmembrane receptors have also been described as being localized in the nucleus. Type I transforming growth factor β receptor (TβR1) can localize to the nucleus following TGF-β treatment,96 where it can interact with hnRNPA1, thus stimulating the alternative splicing of pre-mRNA for EGFR1.97 It is noteworthy that hnRNP A1, besides its known regulatory role in splicing, was also described as an accessory protein of the Microprocessor complex interacting with nTRAIL-R2 in this context.16 However, in contrast to the findings for TRAIL-R2, no association of TβR1 with miRNA maturation has been described till now.
CD40 and the B cell-activating factor receptor (BAFF-R), both members of the TNF receptor superfamily, also localize to the nucleus in normal and malignant B cells, in addition to their plasma membrane expression. In the nucleus, these receptors interact with c-Rel and enhance transcription of several NF-κB target genes.98, 99 Nuclear BAFF-R, in addition, interacts with IKK-β and histone H3, leading to increased IKK-β-mediated phosphorylation of histone H3.99
The surprising role of nTRAIL-R2 in regulating let-7-maturation strongly demands future research on the mechanisms of intracellular trafficking of this receptor. Regarding the mechanism by which full-length plasma membrane receptors passage to the nucleus, the intracellular trafficking of RTKs might serve as a model. Following ligand binding at the cell surface, EGFRs are internalized via endocytosis, moved by retrograde transport in endosomal vesicles into the ER and subsequently translocated through the ER membrane into the cytosol by action of the Sec61 translocon.100 Free cytosolic full-length receptors can then be bound at their tripartite nuclear localization motif (NLS) by β-importin and finally imported into the nucleus through the nuclear pore complex.101 An alternative model for the entry of full-length EGFR family members into the nucleus involving a passage of these receptors into the nucleoplasm through a Sec61 translocon in the inner nuclear membrane has also been proposed.102 For FGF-R1 it was shown that soluble cytoplasmic and nuclear forms and, on the other hand, plasma membrane receptors represent separated pools of different origin and with distinct functions.103 It has been proposed that FGF-R1 is synthesized at ER-attached polyribosomes and either enters Golgi, becomes extensively glycosylated and reaches the plasma membrane or, in non-glycosylated form, exits pre-Golgi vesicles and translocates as soluble cytoplasmic form complexed with RSK1 into the nucleus via nuclear pores.103 The origin of the soluble form of nTRAIL-R2 as well as the mode of its nuclear translocation is currently completely unknown. The receptors may either originate from the plasma membrane or from the non-membraneous soluble cytoplasmic receptor pool. The latter pathway seems to be more likely, as neither inhibition of endogenous secreted or plasma membrane expressed TRAIL via neutralizing antibodies nor the treatment of cells with recombinant TRAIL influenced the nuclear levels of TRAIL-R2.16 Two nuclear localization sequences (NLS) have been identified in TRAIL-R2. It has been reported recently that mutation of these sequences or knockdown of importin-β resulted in a diminished nuclear presence of TRAIL-R2 and, in parallel, enhanced expression at the plasma membrane.69 In contrast to this study, experiments performed in our laboratory using the same NLS-mutated TRAIL-R2 form did not reveal any impact of these NLS sequences on the nuclear import of the recombinant TRAIL-R2 in Panc1 and MDA-MB-231 cells (A. Trauzold, unpublished observations). However, we cannot rule out that the overexpressed NLS-mutated TRAIL-R2 interacts with endogenous TRAIL death receptors and utilize them as Trojan horses for nuclear import in these cells. Further studies are necessary to clarify the exact mechanism of the translocation of TRAIL-R2 to the nucleus.
Another important question concerns the mechanism(s) by which nTRAIL-R2 inhibits the Drosha-mediated processing activity toward pri-let-7. Drosha and DGCR8 constitute the core of the Microprocessor complex that associates with multiple accessory proteins as hnRNPA1, p68, KSRP, NF45 and NF90. The core complex is capable of pri-miRNA processing in the absence of cofactors, whereas the associated factors in the larger complex may facilitate or alter the selectivity of nuclear miRNA processing.104 Accordingly, pri-let-7 is processed in a complex manner involving stimulatory factors (p68 and KSRP) as well as inhibitors (hnRNPA1, NF45 and NF90; Figure 3).105, 106, 107, 108, 109 It seems likely that nTRAIL-R2 modulates the pri-let-7 processing by interaction with NF45, hnRNPA1 and p68.16 This may be facilitated by either sequestration of activators or stabilization of an inhibitory complex.
Role of nuclear p68, KSRP, hnRNPA1 and NF45/NF90 in pri-let-7-processing. Processing of pri-let-7 by the Microprocessor complex Drosha/DGCR8 is subjected to complex regulation by positive and negative factors. P68, as a more general factor, may function as a scaffolding protein, bringing Drosha and accessory proteins together.106 In addition, p68 can also unwind let-7 miRNA duplex, facilitating the loading of pri-miRNA into Microprocessor complex.105 KSRP binds specifically to the loop region of pri-let-7 promoting Drosha-mediated processing, whereas binding of hnRNPA1 to this region not only prevents the recruitment of KSRP, but also inhibits pri-let-7 processing.107, 108 Similarly, NF45 and NF90, acting mainly as a dimer, can specifically bind to pri-let7 and inhibit its maturation to pre-let-7109
It can be assumed that via its interaction with aforementioned proteins, nTRAIL-R2 might also influence the maturation of other miRNAs in addition to let-7. In this respect it is worth mentioning that p68 has been implicated in the recognition of a subset of pri-miRNAs targeting these for Drosha-mediated processing.110 Analysis of global changes of miRNA levels in cells with knockdown of TRAIL-R2 indicates the existence of further TRAIL-R2-regulated miRNA, besides the members of the let-7-family being most prominently represented.16 However, the array results have to be independently verified and, in addition, the question of whether these miRNAs are regulated by nTRAIL-R2 or by plasma membrane-expressed TRAIL-R2 remains to be answered.
Further investigations should also tackle the structural basis for the interactions of nTRAIL-R2 with proteins of the Microprocessor complex. As the core components Drosha and DGCR8 were found to interact with TRAIL-R2 in an indirect manner, attempts should be made to delineate the physical contact sites between TRAIL-R2 and the accessory proteins, that is, hnRNPA1, p68, NF45 and NF90. Insights into the structural constraints of the interaction sites might allow the design of pharmacological inhibitors of these interactions, opening opportunities for targeted therapeutic interventions in the future.
Conclusions and Future Prospects
In spite of more than a decade of research on TRAIL death receptors, until very recently only their canonical signaling functions at the plasma membrane or within the endosomal/lysosomal compartment have been investigated. However, a number of reports also pointed to the intracellular presence of TRAIL-R1 and TRAIL-R2. In intracellular compartments, TRAIL death receptors are likely to neither contribute to canonical apoptosis signaling nor to non-apoptotic signal transduction, regardless of whether they are soluble within cytosol, trapped within the trans-Golgi network, in autophagosomes or are present in the nucleus. The elevated intracellular expression of TRAIL death receptors in cancer and the emerging evidence for their compartmentalization-dependent functions make clear that it is not sufficient to determine the general receptor expression in cells or tissues in a given experimental context. Instead, it is important to perform differentiated analyses of the subcellular distribution of receptors for the understanding of both the molecular biology and the disease-related relevance of TRAIL death receptors, especially when conclusions on the impact of these receptors for disease prognosis are drawn.
Now, a first study has demonstrated a functional impact of the nuclear localization of TRAIL death receptors in cancer cells. TRAIL-R2 was found to associate with components of the large Microprocessor complex inhibiting the processing of pri-let-7 miRNA. This strongly augments the malignant potential of these tumor cells. Such an effect of nTRAIL-R2 on miRNA processing is clearly distinct from the known nuclear functions of other cell surface receptors as EGFR family members, other RTKs or cytokine receptors that can modulate DNA-associated processes like replication, transcription or DNA damage repair and RNA splicing.
Thus, these novel findings will stimulate the search for unknown TRAIL-R2-specific nuclear functions, among them regulation of other miRNAs and possibly also RNA- and DNA-targeting mechanisms. This will further broaden the range of possibly affected nuclear regulatory processes of plasma membrane receptors that emerge as important regulators in the cell nucleus. The findings on a specific function of nTRAIL-R2 should also promote the search for possible unknown nuclear functions of other members of the death receptor family. In this respect, the capability of TRAIL-R1 and TRAIL-R4 to specifically interact with TRAIL-R2 in the nucleus also raises the question of the capacity of these receptors to modulate the nuclear functions of TRAIL-R2. Finally, these recent insights into TRAIL receptor biology might open new perspectives for anti-cancer therapies targeting processes regulated by nTRAIL-R2.
Abbreviations
- APAF-1:
-
apoptotic protease activating factor-1
- DED:
-
death effector domain
- DISC:
-
death-inducing signaling complex
- ERK1:
-
extracellular regulated kinase-1
- FADD:
-
Fas-associated protein with death domain
- cFLIP:
-
cellular flice-like inhibitory protein
- HtrA2/Omi:
-
high temperature requirement protein A2/Omi
- JNK:
-
c-Jun N-terminal kinase
- LMP:
-
lysosomal membrane permeabilization
- miRNA:
-
microRNA
- MLKL:
-
mixed lineage kinase domain-like
- MOMP:
-
mitochondrial outer membrane permeabilization
- NF-κB:
-
nuclear factor κ-light-chain-enhancer of activated B cells
- OPG:
-
osteoprotegerin
- PACS-2:
-
phosphofurin acidic cluster sorting protein-2
- PKB:
-
protein kinase B
- PKC:
-
protein kinase C
- RIP-1:
-
receptor-interacting protein-1
- Smac/Diablo:
-
second mitochondria-derived activator of caspase/direct
- IAP:
-
binding protein with low pI
- TRAIL:
-
TNF-related apoptosis-inducing ligand
References
Pan G, O'Rourke K, Chinnaiyan AM, Gentz R, Ebner R, Ni J et al. The receptor for the cytotoxic ligand TRAIL. Science 1997; 276: 111–113.
Walczak H, Degli-Esposti MA, Johnson RS, Smolak PJ, Waugh JY, Boiani N et al. TRAIL-R2: a novel apoptosis-mediating receptor for TRAIL. EMBO J 1997; 16: 5386–5397.
Screaton GR, Mongkolsapaya J, Xu XN, Cowper AE, McMichael AJ, Bell JI . TRICK2, a new alternatively spliced receptor that transduces the cytotoxic signal from TRAIL. Curr Biol 1997; 7: 693–696.
Pan G, Ni J, Wei YF, Yu G, Gentz R, Dixit VM . An antagonist decoy receptor and a death domain-containing receptor for TRAIL. Science 1997; 277: 815–818.
Degli-Esposti MA, Smolak PJ, Walczak H, Waugh J, Huang CP, DuBose RF et al. Cloning and characterization of TRAIL-R3, a novel member of the emerging TRAIL receptor family. J Exp Med 1997; 186: 1165–1170.
Degli-Esposti MA, Dougall WC, Smolak PJ, Waugh JY, Smith CA, Goodwin RG . The novel receptor TRAIL-R4 induces NF-kappaB and protects against TRAIL-mediated apoptosis, yet retains an incomplete death domain. Immunity 1997; 7: 813–820.
Schneider P, Thome M, Burns K, Bodmer JL, Hofmann K, Kataoka T et al. TRAIL receptors 1 (DR4) and 2 (DR5) signal FADD-dependent apoptosis and activate NF-kappaB. Immunity 1997; 7: 831–836.
Emery JG, McDonnell P, Burke MB, Deen KC, Lyn S, Silverman C et al. Osteoprotegerin is a receptor for the cytotoxic ligand TRAIL. J Biol Chem 1998; 273: 14363–14367.
Ashkenazi A, Pai RC, Fong S, Leung S, Lawrence DA, Marsters SA et al. Safety and antitumor activity of recombinant soluble Apo2 ligand. J Clin Invest 1999; 104: 155–162.
Walczak H, Miller RE, Ariail K, Gliniak B, Griffith TS, Kubin M et al. Tumoricidal activity of tumor necrosis factor-related apoptosis-inducing ligand in vivo. Nat Med 1999; 5: 157–163.
Dimberg LY, Anderson CK, Camidge R, Behbakht K, Thorburn A, Ford HL . On the TRAIL to successful cancer therapy? Predicting and counteracting resistance against TRAIL-based therapeutics. Oncogene 2013; 32: 1341–1350.
Micheau O, Shirley S, Dufour F . Death receptors as targets in cancer. Br J Pharmacol 2013; 169: 1723–1744.
Wiezorek J, Holland P, Graves J . Death receptor agonists as a targeted therapy for cancer. Clin Cancer Res 2010; 16: 1701–1708.
Röder C, Trauzold A, Kalthoff H . Impact of death receptor signaling on the malignancy of pancreatic ductal adenocarcinoma. Eur J Cell Biol 2011; 90: 450–455.
Azijli K, Weyhenmeyer B, Peters GJ, de Jong S, FAE Kruyt . Non-canonical kinase signaling by the death ligand TRAIL in cancer cells: discord in the death receptor family. Cell Death Differ 2013; 20: 858–868.
Haselmann V, Kurz A, Bertsch U, Hübner S, Olempska-Müller M, Fritsch J et al. Nuclear death receptor TRAIL-R2 inhibits maturation of Let-7 and promotes proliferation of pancreatic and other tumor cells. Gastroenterology 2014; 146: 278–290.
Gallmeier E, Bader DC, Kriegl L, Berezowska S, Seeliger H, Goke B et al. Loss of TRAIL-receptors is a recurrent feature in pancreatic cancer and determines the prognosis of patients with no nodal metastasis after surgery. PLoS One 2013; 8: e56760.
Bavi P, Prabhakaran SE, Abubaker J, Qadri Z, George T, Al-Sanea N et al. Prognostic significance of TRAIL death receptors in Middle Eastern colorectal carcinomas and their correlation to oncogenic KRAS alterations. Mol Cancer 2010; 9: 203.
Sträter J, Hinz U, Walczak H, Mechtersheimer G, Koretz K, Herfarth C et al. Expression of TRAIL and TRAIL receptors in colon carcinoma: TRAIL-R1 is an independent prognostic parameter. Clin Cancer Res 2002; 8: 3734–3740.
Ganten TM, Sykora J, Koschny R, Batke E, Aulmann S, Mansmann U et al. Prognostic significance of tumour necrosis factor-related apoptosis-inducing ligand (TRAIL) receptor expression in patients with breast cancer. J Mol Med 2009; 87: 995–1007.
McCarthy MM, Sznol M, DiVito KA, Camp RL, Rimm DL, Kluger HM . Evaluating the expression and prognostic value of TRAIL-R1 and TRAIL-R2 in breast cancer. Clin Cancer Res 2005; 11: 5188–5194.
Kuijlen JM, Mooij JJ, Platteel I, Hoving EW, van der Graaf WT, Span MM et al. TRAIL-receptor expression is an independent prognostic factor for survival in patients with a primary glioblastoma multiforme. J Neurooncol 2006; 78: 161–171.
Elrod HA, Fan S, Muller S, Chen GZ, Pan L, Tighiouart M et al. Analysis of death receptor 5 and caspase-8 expression in primary and metastatic head and neck squamous cell carcinoma and their prognostic impact. PLoS One 2010; 5: e12178.
Maduro JH, Noordhuis MG, ten Hoor KA, Pras E, Arts HJ, Eijsink JJ et al. The prognostic value of TRAIL and its death receptors in cervical cancer. Int J Radiat Oncol Biol Phys 2009; 75: 203–211.
Dong HP, Kleinberg L, Silins I, Florenes VA, Trope CG, Risberg B et al. Death receptor expression is associated with poor response to chemotherapy and shorter survival in metastatic ovarian carcinoma. Cancer 2008; 112: 84–93.
Vigneswaran N, Baucum DC, Wu J, Lou Y, Bouquot J, Muller S et al. Repression of tumor necrosis factor-related apoptosis-inducing ligand (TRAIL) but not its receptors during oral cancer progression. BMC Cancer 2007; 7: 108.
Cooper WA, Kohonen-Corish MR, Zhuang L, McCaughan B, Kennedy C, Screaton G et al. Role and prognostic significance of tumor necrosis factor-related apoptosis-inducing ligand death receptor DR5 in nonsmall-cell lung cancer and precursor lesions. Cancer 2008; 113: 135–142.
Spierings DC, de Vries EG, Timens W, Groen HJ, Boezen HM, de Jong S . Expression of TRAIL and TRAIL death receptors in stage III non-small cell lung cancer tumors. Clin Cancer Res 2003; 9: 3397–3405.
Li Y, Jin X, Li J, Jin X, Yu J, Sun X et al. Expression of TRAIL, DR4, and DR5 in bladder cancer: correlation with response to adjuvant therapy and implications of prognosis. Urology 2012; 79: e967–915.
Rajeshkumar NV, Rasheed ZA, Garcia-Garcia E, Lopez-Rios F, Fujiwara K, Matsui WH et al. A combination of DR5 agonistic monoclonal antibody with gemcitabine targets pancreatic cancer stem cells and results in long-term disease control in human pancreatic cancer model. Mol Cancer Ther 2010; 9: 2582–2592.
Niu Y, Li Y, Zang J, Huang H, Deng J, Cui Z et al. Death receptor 5 and neuroproliferation. Cell Mol Neurobiol 2012; 32: 255–265.
Gonzalvez F, Ashkenazi A . New insights into apoptosis signaling by Apo2L/TRAIL. Oncogene 2010; 29: 4752–4765.
Merino D, Lalaoui N, Morizot A, Schneider P, Solary E, Micheau O . Differential inhibition of TRAIL-mediated DR5-DISC formation by decoy receptors 1 and 2. Mol Cell Biol 2006; 26: 7046–7055.
Neumann S, Hasenauer J, Pollak N, Scheurich P . Dominant negative effects of TNF-related apoptosis-inducing ligand (TRAIL) receptor 4 on TRAIL receptor 1 signaling by formation of heteromeric complexes. J Biol Chem 2014; 289: 16576–16587.
Haag C, Stadel D, Zhou S, Bachem MG, Moller P, Debatin KM et al. Identification of c-FLIP(L) and c-FLIP(S) as critical regulators of death receptor-induced apoptosis in pancreatic cancer cells. Gut 2011; 60: 225–237.
Kavuri SM, Geserick P, Berg D, Dimitrova DP, Feoktistova M, Siegmund D et al. Cellular FLICE-inhibitory protein (cFLIP) isoforms block CD95- and TRAIL death receptor-induced gene induction irrespective of processing of caspase-8 or cFLIP in the death-inducing signaling complex. J Biol Chem 2011; 286: 16631–16646.
Scaffidi C, Fulda S, Srinivasan A, Friesen C, Li F, Tomaselli KJ et al. Two CD95 (APO-1/Fas) signaling pathways. Embo J 1998; 17: 1675–1687.
Akazawa Y, Mott JL, Bronk SF, Werneburg NW, Kahraman A, Guicciardi ME et al. Death receptor 5 internalization is required for lysosomal permeabilization by TRAIL in malignant liver cell lines. Gastroenterology 2009; 136: 2365–2376.
Werneburg NW, Bronk SF, Guicciardi ME, Thomas L, Dikeakos JD, Thomas G et al. Tumor necrosis factor-related apoptosis-inducing ligand (TRAIL) protein-induced lysosomal translocation of proapoptotic effectors is mediated by phosphofurin acidic cluster sorting protein-2 (PACS-2). J Biol Chem 2012; 287: 24427–24437.
Jouan-Lanhouet S, Arshad MI, Piquet-Pellorce C, Martin-Chouly C, Le Moigne-Muller G, Van Herreweghe F et al. TRAIL induces necroptosis involving RIPK1/RIPK3-dependent PARP-1 activation. Cell Death Differ 2012; 19: 2003–2014.
Voigt S, Philipp S, Davarnia P, Winoto-Morbach S, Roder C, Arenz C et al. TRAIL-induced programmed necrosis as a novel approach to eliminate tumor cells. BMC Cancer 2014; 14: 74.
Azijli K, Yuvaraj S, Peppelenbosch MP, Wurdinger T, Dekker H, Joore J et al. Kinome profiling of non-canonical TRAIL signaling reveals RIP1-Src-STAT3-dependent invasion in resistant non-small cell lung cancer cells. J Cell Sci 2012; 125: 4651–4661.
Jeremias I, Kupatt C, Baumann B, Herr I, Wirth T, Debatin KM . Inhibition of nuclear factor kappaB activation attenuates apoptosis resistance in lymphoid cells. Blood 1998; 91: 4624–4631.
Trauzold A, Wermann H, Arlt A, Schutze S, Schafer H, Oestern S et al. CD95 and TRAIL receptor-mediated activation of protein kinase C and NF-kappa B contributes to apoptosis resistance in ductal pancreatic adenocarcinoma cells. Oncogene 2001; 20: 4258–4269.
Ehrhardt H, Fulda S, Schmid I, Hiscott J, Debatin KM, Jeremias I . TRAIL induced survival and proliferation in cancer cells resistant towards TRAIL-induced apoptosis mediated by NF-kappaB. Oncogene 2003; 22: 3842–3852.
Grunert M, Gottschalk K, Kapahnke J, Gundisch S, Kieser A, Jeremias I . The adaptor protein FADD and the initiator caspase-8 mediate activation of NF-kappaB by TRAIL. Cell Death Dis 2012; 3: e414.
Secchiero P, Melloni E, Heikinheimo M, Mannisto S, Di Pietro R, Iacone A et al. TRAIL regulates normal erythroid maturation through an ERK-dependent pathway. Blood 2004; 103: 517–522.
Siegmund D, Klose S, Zhou D, Baumann B, Roder C, Kalthoff H et al. Role of caspases in CD95L- and TRAIL-induced non-apoptotic signalling in pancreatic tumour cells. Cell Signal 2007; 19: 1172–1184.
Zauli G, Sancilio S, Cataldi A, Sabatini N, Bosco D, Di Pietro R . PI-3K/Akt and NF-kappaB/IkappaBalpha pathways are activated in Jurkat T cells in response to TRAIL treatment. J Cell Physiol 2005; 202: 900–911.
Choo MK, Kawasaki N, Singhirunnusorn P, Koizumi K, Sato S, Akira S et al. Blockade of transforming growth factor-beta-activated kinase 1 activity enhances TRAIL-induced apoptosis through activation of a caspase cascade. Mol Cancer Ther 2006; 5: 2970–2976.
Xu J, Zhou JY, Wei WZ, Wu GS . Activation of the Akt survival pathway contributes to TRAIL resistance in cancer cells. PLoS One 2010; 5: e10226.
Varfolomeev E, Maecker H, Sharp D, Lawrence D, Renz M, Vucic D et al. Molecular determinants of kinase pathway activation by Apo2 ligand/tumor necrosis factor-related apoptosis-inducing ligand. J Biol Chem 2005; 280: 40599–40608.
Song JH, Tse MC, Bellail A, Phuphanich S, Khuri F, Kneteman NM et al. Lipid rafts and nonrafts mediate tumor necrosis factor related apoptosis-inducing ligand induced apoptotic and nonapoptotic signals in non small cell lung carcinoma cells. Cancer Res 2007; 67: 6946–6955.
Song JJ, Lee YJ . Differential cleavage of Mst1 by caspase-7/-3 is responsible for TRAIL-induced activation of the MAPK superfamily. Cell Signal 2008; 20: 892–906.
Trauzold A, Siegmund D, Schniewind B, Sipos B, Egberts J, Zorenkov D et al. TRAIL promotes metastasis of human pancreatic ductal adenocarcinoma. Oncogene 2006; 25: 7434–7439.
Ishimura N, Isomoto H, Bronk SF, Gores GJ . Trail induces cell migration and invasion in apoptosis-resistant cholangiocarcinoma cells. Am J Physiol Gastrointest Liver Physiol 2006; 290: G129–G136.
Hoogwater FJ, Nijkamp MW, Smakman N, Steller EJ, Emmink BL, Westendorp BF et al. Oncogenic K-Ras turns death receptors into metastasis-promoting receptors in human and mouse colorectal cancer cells. Gastroenterology 2010; 138: 2357–2367.
Granci V, Bibeau F, Kramar A, Boissière-Michot F, Thézénas S, Thirion A et al. Prognostic significance of TRAIL-R1 and TRAIL-R3 expression in metastatic colorectal carcinomas. Eur J Cancer 2008; 44: 2312–2318.
Chen JJ, JH-C Shen, Rivera Rosado L, Zhang Y, Di X, Zhang B . Mislocalization of death receptors correlates with cellular resistance to their cognate ligands in human breast cancer cells. Oncotarget 2012; 3: 833–842.
Sanlioglu AD, Korcum AF, Pestereli E, Erdogan G, Karaveli S, Savas B et al. TRAIL death receptor-4 expression positively correlates with the tumor grade in breast cancer patients with invasive ductal carcinoma. Int J Radiat Oncol Biol Phys 2007; 69: 716–723.
Macher-Goeppinger S, Aulmann S, Tagscherer KE, Wagener N, Haferkamp A, Penzel R et al. Prognostic value of tumor necrosis factor-related apoptosis-inducing ligand (TRAIL) and TRAIL receptors in renal cell cancer. Clin Cancer Res 2009; 15: 650–659.
Zhuang L, Lee CS, Scolyer RA, McCarthy SW, Zhang XD, Thompson JF et al. Progression in melanoma is associated with decreased expression of death receptors for tumor necrosis factor-related apoptosis-inducing ligand. Hum Pathol 2006; 37: 1286–1294.
Kriegl L, Jung A, Engel J, Jackstadt R, Gerbes AL, Gallmeier E et al. Expression, cellular distribution, and prognostic relevance of TRAIL receptors in hepatocellular carcinoma. Clin Cancer Res 2010; 16: 5529–5538.
Oikonomou E, Kosmidou V, Katseli A, Kothonidis K, Mourtzoukou D, Kontogeorgos G et al. TRAIL receptor upregulation and the implication of KRAS/BRAF mutations in human colon cancer tumors. Int J Cancer 2009; 125: 2127–2135.
Zhang XD, Franco AV, Nguyen T, Gray CP, Hersey P . Differential localization and regulation of death and decoy receptors for TNF-related apoptosis-inducing ligand (TRAIL) in human melanoma cells. J Immunol 2000; 164: 3961–3970.
Di X, Zhang G, Zhang Y, Takeda K, Rosado LA, Zhang B . Accumulation of autophagosomes in breast cancer cells induces TRAIL resistance through downregulation of surface expression of death receptors 4 and 5. Oncotarget 2013; 4: 1349–1364.
Bai X, Williams JL, Greenwood SL, Baker PN, Aplin JD, Crocker IP . A placental protective role for trophoblast-derived TNF-related apoptosis-inducing ligand (TRAIL). Placenta 2009; 30: 855–860.
Leithner K, Stacher E, Wurm R, Ploner F, Quehenberger F, Wohlkoenig C et al. Nuclear and cytoplasmic death receptor 5 as prognostic factors in patients with non-small cell lung cancer treated with chemotherapy. Lung Cancer 2009; 65: 98–104.
Kojima Y, Nakayama M, Nishina T, Nakano H, Koyanagi M, Takeda K et al. Importin beta1 protein-mediated nuclear localization of death receptor 5 (DR5) limits DR5/tumor necrosis factor (TNF)-related apoptosis-inducing ligand (TRAIL)-induced cell death of human tumor cells. J Biol Chem 2011; 286: 43383–43393.
Peternel S, Prpic-Massari L, Manestar-Blazic T, Brajac I, Kastelan M . Increased expression of TRAIL and its death receptors DR4 and DR5 in plaque psoriasis. Arch Dermatol Res 2011; 303: 389–397.
Lee YS, Dutta A . The tumor suppressor microRNA let-7 represses the HMGA2 oncogene. Genes Dev 2007; 21: 1025–1030.
Park SM, Shell S, Radjabi AR, Schickel R, Feig C, Boyerinas B et al. Let-7 prevents early cancer progression by suppressing expression of the embryonic gene HMGA2. Cell Cycle 2007; 6: 2585–2590.
Rybak A, Fuchs H, Smirnova L, Brandt C, Pohl EE, Nitsch R et al. A feedback loop comprising lin-28 and let-7 controls pre-let-7 maturation during neural stem-cell commitment. Nat Cell Biol 2008; 10: 987–993.
Carpenter G, Liao HJ . Trafficking of receptor tyrosine kinases to the nucleus. Exp Cell Res 2009; 315: 1556–1566.
Wang YN, Hung MC . Nuclear functions and subcellular trafficking mechanisms of the epidermal growth factor receptor family. Cell Biosci 2012; 2: 13.
Hung LY, Tseng JT, Lee YC, Xia W, Wang YN, Wu ML et al. Nuclear epidermal growth factor receptor (EGFR) interacts with signal transducer and activator of transcription 5 (STAT5) in activating Aurora-A gene expression. Nucleic Acids Res 2008; 36: 4337–4351.
Lo HW, Cao X, Zhu H, Ali-Osman F . Cyclooxygenase-2 is a novel transcriptional target of the nuclear EGFR-STAT3 and EGFRvIII-STAT3 signaling axes. Mol Cancer Res 2010; 8: 232–245.
Lo HW, Hsu SC, Ali-Seyed M, Gunduz M, Xia W, Wei Y et al. Nuclear interaction of EGFR and STAT3 in the activation of the iNOS/NO pathway. Cancer Cell 2005; 7: 575–589.
Jaganathan S, Yue P, Paladino DC, Bogdanovic J, Huo Q, Turkson J . A functional nuclear epidermal growth factor receptor, SRC and Stat3 heteromeric complex in pancreatic cancer cells. PLoS One 2011; 6: e19605.
Hanada N, Lo HW, Day CP, Pan Y, Nakajima Y, Hung MC . Co-regulation of B-Myb expression by E2F1 and EGF receptor. Mol Carcinog 2006; 45: 10–17.
Huo L, Wang YN, Xia W, Hsu SC, Lai CC, Li LY et al. RNA helicase A is a DNA-binding partner for EGFR-mediated transcriptional activation in the nucleus. Proc Natl Acad Sci USA 2010; 107: 16125–16130.
Li LY, Chen H, Hsieh YH, Wang YN, Chu HJ, Chen YH et al. Nuclear ErbB2 enhances translation and cell growth by activating transcription of ribosomal RNA genes. Cancer Res 2011; 71: 4269–4279.
Wang SC, Nakajima Y, Yu YL, Xia W, Chen CT, Yang CC et al. Tyrosine phosphorylation controls PCNA function through protein stability. Nat Cell Biol 2006; 8: 1359–1368.
Dittmann K, Mayer C, Fehrenbacher B, Schaller M, Raju U, Milas L et al. Radiation-induced epidermal growth factor receptor nuclear import is linked to activation of DNA-dependent protein kinase. J Biol Chem 2005; 280: 31182–31189.
Hsu SC, Miller SA, Wang Y, Hung MC . Nuclear EGFR is required for cisplatin resistance and DNA repair. Am J Transl Res 2009; 1: 249–258.
Liccardi G, Hartley JA, Hochhauser D . EGFR nuclear translocation modulates DNA repair following cisplatin and ionizing radiation treatment. Cancer Res 2011; 71: 1103–1114.
Bryant DM, Stow JL . Nuclear translocation of cell-surface receptors: lessons from fibroblast growth factor. Traffic 2005; 6: 947–953.
Cai J, Jiang WG, Grant MB, Boulton M . Pigment epithelium-derived factor inhibits angiogenesis via regulated intracellular proteolysis of vascular endothelial growth factor receptor 1. J Biol Chem 2006; 281: 3604–3613.
Shay-Salit A, Shushy M, Wolfovitz E, Yahav H, Breviario F, Dejana E et al. VEGF receptor 2 and the adherens junction as a mechanical transducer in vascular endothelial cells. Proc Natl Acad Sci USA 2002; 99: 9462–9467.
Kasuga K, Kaneko H, Nishizawa M, Onodera O, Ikeuchi T . Generation of intracellular domain of insulin receptor tyrosine kinase by gamma-secretase. Biochem Biophys Res Commun 2007; 360: 90–96.
Gomes DA, Rodrigues MA, Leite MF, Gomez MV, Varnai P, Balla T et al. c-Met must translocate to the nucleus to initiate calcium signals. J Biol Chem 2008; 283: 4344–4351.
Bonacchi A, Taddei ML, Petrai I, Efsen E, Defranco R, Nosi D et al. Nuclear localization of TRK-A in liver cells. Histol Histopathol 2008; 23: 327–340.
Conway-Campbell BL, Wooh JW, Brooks AJ, Gordon D, Brown RJ, Lichanska AM et al. Nuclear targeting of the growth hormone receptor results in dysregulation of cell proliferation and tumorigenesis. Proc Natl Acad Sci USA 2007; 104: 13331–13336.
Tseng HC, Lyu PC, Lin WC . Nuclear localization of orphan receptor protein kinase (Ror1) is mediated through the juxtamembrane domain. BMC Cell Biol 2010; 11: 48.
Hu Y, Fang X, Dunham SM, Prada C, Stachowiak EK, Stachowiak MK . 90-kDa ribosomal S6 kinase is a direct target for the nuclear fibroblast growth factor receptor 1 (FGFR1): role in FGFR1 signaling. J Biol Chem 2004; 279: 29325–29335.
Zwaagstra JC, Guimond A, O'Connor-McCourt MD . Predominant intracellular localization of the type I transforming growth factor-β receptor and increased nuclear accumulation after growth arrest. Exp Cell Res 2000; 258: 121–134.
Chandra M, Zang S, Li H, Zimmerman LJ, Champer J, Tsuyada A et al. Nuclear translocation of type I transforming growth factor β receptor confers a novel function in RNA processing. Mol Cell Biol 2012; 32: 2183–2195.
Zhou HJ, Pham LV, Tamayo AT, Lin-Lee YC, Fu L, Yoshimura LC et al. Nuclear CD40 interacts with c-Rel and enhances proliferation in aggressive B-cell lymphoma. Blood 2007; 110: 2121–2127.
Fu LC, Lin-Lee YC, Pham LV, Tamayo AT, Yoshimura LC, Ford RJ . BAFF-R promotes cell proliferation and survival through interaction with IKK beta and NF-kappa B/c-Rel in the nucleus of normal and neoplastic B-lymphoid cells. Blood 2009; 113: 4627–4636.
Liao HJ, Carpenter G . Role of the Sec61 translocon in EGF receptor trafficking to the nucleus and gene expression. Mol Biol Cell 2007; 18: 1064–1072.
Giri DK, Ali-Seyed M, Li LY, Lee DF, Ling P, Bartholomeusz G et al. Endosomal transport of ErbB-2: mechanism for nuclear entry of the cell surface receptor. Mol Cell Biol 2005; 25: 11005–11018.
Wang YN, Yamaguchi H, Huo L, Du Y, Lee HJ, Lee HH et al. The translocon Sec61beta localized in the inner nuclear membrane transports membrane-embedded EGF receptor to the nucleus. J Biol Chem 2010; 285: 38720–38729.
Dunham-Ems SM, Pudavar HE, Myers JM, Maher PA, Prasad PN, Stachowiak MK . Factors controlling fibroblast growth factor receptor-1's cytoplasmic trafficking and its regulation as revealed by FRAP analysis. Mol Biol Cell 2006; 17: 2223–2235.
Gregory RI, Yan KP, Amuthan G, Chendrimada T, Doratotaj B, Cooch N et al. The Microprocessor complex mediates the genesis of microRNAs. Nature 2004; 432: 235–240.
Salzman DW, Shubert-Coleman J, Furneaux H . P68 RNA helicase unwinds the human let-7 microRNA precursor duplex and is required for let-7-directed silencing of gene expression. J Biol Chem 2007; 282: 32773–32779.
Suzuki HI, Yamagata K, Sugimoto K, Iwamoto T, Kato S, Miyazono K . Modulation of microRNA processing by p53. Nature 2009; 460: 529–533.
Trabucchi M, Briata P, Garcia-Mayoral M, Haase AD, Filipowicz W, Ramos A et al. The RNA-binding protein KSRP promotes the biogenesis of a subset of microRNAs. Nature 2009; 459: 1010–1014.
Michlewski G, Caceres JF . Antagonistic role of hnRNP A1 and KSRP in the regulation of let-7a biogenesis. Nat Struct Mol Biol 2010; 17: 1011–1018.
Sakamoto S, Aoki K, Higuchi T, Todaka H, Morisawa K, Tamaki N et al. The NF90-NF45 complex functions as a negative regulator in the microRNA processing pathway. Mol Cell Biol 2009; 29: 3754–3769.
Fukuda T, Yamagata K, Fujiyama S, Matsumoto T, Koshida I, Yoshimura K et al. DEAD-box RNA helicase subunits of the Drosha complex are required for processing of rRNA and a subset of microRNAs. Nat Cell Biol 2007; 9: 604–611.
Acknowledgements
Our research was supported by the Deutsche Forschungsgemeinschaft (PCC-Kiel/project TR 1063/2-1; SKELMET FOR 1586/ project TR 1063/3-1). Investigation of clinical specimens was supported by the P2N Biobank of the Medical Faculty Kiel (BMBF grant 01EY1103).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Edited by H-U Simon
Rights and permissions
Cell Death and Disease is an open-access journal published by Nature Publishing Group. This work is licensed under a Creative Commons Attribution-NonCommercial-ShareAlike 3.0 Unported License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-sa/3.0/
About this article
Cite this article
Bertsch, U., Röder, C., Kalthoff, H. et al. Compartmentalization of TNF-related apoptosis-inducing ligand (TRAIL) death receptor functions: emerging role of nuclear TRAIL-R2. Cell Death Dis 5, e1390 (2014). https://doi.org/10.1038/cddis.2014.351
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/cddis.2014.351
- Springer Nature Limited
This article is cited by
-
Death receptor 5 is required for intestinal stem cell activity during intestinal epithelial renewal at homoeostasis
Cell Death & Disease (2024)
-
Postdiagnosis circulating osteoprotegerin and TRAIL concentrations and survival and recurrence after a breast cancer diagnosis: results from the MARIE patient cohort
Breast Cancer Research (2023)
-
Proteomic profiling of longitudinal changes in kidney function among middle-aged and older men and women: the KORA S4/F4/FF4 study
BMC Medicine (2023)
-
Harnessing TRAIL-induced cell death for cancer therapy: a long walk with thrilling discoveries
Cell Death & Differentiation (2023)
-
Nodular fasciitis: a comprehensive, time-correlated investigation of 17 cases
Modern Pathology (2021)