Abstract
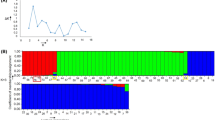
Radish (Raphanus sativus L.) is a major vegetable cultivated worldwide. As a member of the family Brassicaceae, this species has diverse morphological characteristics in root. Radish cultivars have been classified based on morphological traits, including root shape and color. Despite its economic importance in Asia, genomic research in radish is less well developed relative to Brassica rapa, a close relative of radish. In this study, we developed genomic simple sequence repeat (SSR) markers using a SSR-enriched library and investigated genetic diversity in a collection of 144 radish cultivars. A total of 237 primer pairs for SSRs were designed and 184 (77.6%) primer pairs produced PCR amplicons. Of these, we selected 27 SSR markers (14.7%) based on polymorphism in a subset of 11 cultivars and then used to assess genetic relationships in the germplasm panel. For these markers, the number of alleles per marker ranged from 2 to 18 with an average of 7.77 alleles and the polymorphism information content (PIC) values ranged from 0.491 to 0.906. The estimates of pairwise F st revealed significant genetic differentiation between the five market classes of 135 radish cultivars (74 big, 15 small, 29 young, 12 Altari, and 5 processing). Clustering analysis using NTSYS-pc and STRUCTURE software also found that the 74 big radishes were divided into 5–7 clusters. In addition, all 27 SSR markers were able to differentiate 64 big radish cultivars based on the UPGMA dendrogram and each of the 23 markers independently identified 1 to 17 big radish cultivars. These results suggest that cultivated radishes have different patterns of genetic variation and breeding practices should be a driving force for the genetic differentiation between and within market classes. The SSR markers developed in this study will be a useful resource for genetic study and protection of plant breeder’s intellectual property right through cultivar identification.
Similar content being viewed by others
Literature Cited
Anderson, J.A., G.A. Churchill, J.E. Autrique, S.D. Tanksley, and M.E. Sorrells. 1993. Optimizing parental selection for genetic-linkage maps. Genome 36:181–186.
Blair, M.W., M.C. Giraldo, H.F. Buendia, E. Tovar, M.C. Duque, and S.E. Beebe. 2006. Microsatellite marker diversity in common bean (Phaseolus vulgaris L.). Theor. Appl. Genet. 113:100–109.
Clapham, A.R., T.G. Tutin, and E.F. Warburg. 1962. Flora of the British Isles. Cambridge University Press, Cambridge, UK.
Cloutier, S., Z. Niu, R. Datla, and S. Duguid. 2009. Development and analysis of EST-SSRs for flax (Linum usitatissimum L.). Theor. Appl. Genet. 119:53–63.
The Tomato Genome Consortium. 2012. The tomato genome sequence provides insights into fleshy fruit evolution. Nature 485:635–641.
Crisp, D. 1995. Radish, Raphanus sativus (Cruciferae), p.86–89 In: J. Smarttand N.W. Simmonds (eds.). Evolution of crop plants. Longman Scientific & Technical, Harlow, UK.
De Candolle, A. 1886. Origin of cultivated plants. 2nd ed. Hafner Publishing Company, New York, NY, USA.
Dieringer, D. and C. Schlotterer. 2003. MICROSATELLITEANALYSER (MSA): A platform independent analysis tool for large microsatellite data sets. Mol. Ecol. Notes 3:167–169.
Evanno, G., S. Regnaut, and J. Goudet. 2005. Detecting the number of clusters of individuals using the software STRUCTURE: A simulation study. Mol. Ecol. 14:2611–2620.
Hamilton J.P., S. Sim, K. Stoffel, A. Van Deynze, C.R. Buell, and D.M. Francis. 2012. Single nucleotide polymorphism discovery in cultivated tomato via sequencing by synthesis. Plant Genome-US 5:17–29.
Henslow, G. 1898. The history of the radish. Gard.Chron. 23:389.
tHuang, S.W., R.Q. Li, Z.H. Zhang, L. Li, X.F. Gu, W. Fan, W.J. Lucas, X.W. Wang, B.Y. Xie, P.X. Ni, Y.Y. Ren, H.M. Zhu, J. Li, K. Lin, W.W. Jin, Z.J. Fei, G.C. Li, J. Staub, A. Kilian, E.a.G. Van Der Vossen, Y. Wu, J. Guo, J. He, Z.Q. Jia, Y. Ren, G. Tian, Y. Lu, J. Ruan, W.B. Qian, M.W. Wang, Q.F. Huang, B. Li, Z.L. Xuan, J.J. Cao, Asan, Z.G. Wu, J.B. Zhang, Q.L. Cai, Y.Q. Bai, B.W. Zhao, Y.H. Han, Y. Li, X.F. Li, S.H. Wang, Q.X. Shi, S.Q. Liu, W.K. Cho, J.Y. Kim, Y. Xu, K. Heller-Uszynska, H. Miao, Z.C. Cheng, S.P. Zhang, J. Wu, Y.H. Yang, H.X. Kang, M. Li, H.Q. Liang, X.L. Ren, Z.B. Shi, M. Wen, M. Jian, H.L. Yang, G.J. Zhang, Z.T. Yang, R. Chen, S.F. Liu, J.W. Li, L.J. Ma, H. Liu, Y. Zhou, J. Zhao, X.D. Fang, G.Q. Li, L. Fang, Y.R. Li, D.Y. Liu, H.K. Zheng, Y. Zhang, N. Qin, Z. Li, G.H. Yang, S. Yang, L. Bolund, K. Kristiansen, H.C. Zheng, S.C. Li, X.Q. Zhang, H.M. Yang, J. Wang, R.F. Sun, B.X. Zhang, S.Z. Jiang, J. Wang, Y.C. Du, and S.G. Li. 2009. The genome of the cucumber, Cucumis sativus L. Nat. Genet. 41:1275–1281.
Jarne, P. and P.J.L. Lagoda. 1996. Microsatellites, from molecules to populations and back. Trends Ecol. Evol. 11:424–429.
Jiang, L.N., L.J. Wang, L.W. Liu, X.W. Zhu, L.L. Zhai, and Y.Q. Cong. 2012. Development and characterization of cDNA library based novel EST-SSR marker in radish (Raphanus sativus L.). Sci. Hortic. 140:164–172.
Johnston, J.S., A.E. Pepper, A.E. Hall, Z.J. Chen, G. Hodnett, J. Drabek, R. Lopez, and H.J. Price. 2005. Evolution of genome size in Brassicaceae. Ann. Bot. 95:229–235.
Kaneko, Y., C. Kimizuka-Takagi, S.W. Bang, and Y. Matsuzawa. 2007. Radish, p. 141–160. In:C. Kole (ed.).Genome mapping and molecular breeding in plants: Vegetables. Springer, Berlin Heidelberg, Germany.
Kim, S., M. Park, S.I. Yeom, Y.M. Kim, J.M. Lee, H.A. Lee, E. Seo, J. Choi, K. Cheong, K.T. Kim, K. Jung, G.W. Lee, S.K. Oh, C. Bae, S.B. Kim, H.Y. Lee, S.Y. Kim, M.S. Kim, B.C. Kang, Y.D. Jo, H.B. Yang, H.J. Jeong, W.H. Kang, J.K. Kwon, C. Shin, J.Y. Lim, J.H. Park, J.H. Huh, J.S. Kim, B.D. Kim, O. Cohen, I. Paran, M.C. Suh, S.B. Lee, Y.K. Kim, Y. Shin, S.J. Noh, J. Park, Y.S. Seo, S.Y. Kwon, H.A. Kim, J.M. Park, H.J. Kim, S.B. Choi, P.W. Bosland, G. Reeves, S.H. Jo, B.W. Lee, H.T. Cho, H.S. Choi, M.S. Lee, Y. Yu, Y. Do Choi, B.S. Park, A. Van Deynze, H. Ashrafi, T. Hill, W.T. Kim, H.S. Pai, H.K. Ahn, I. Yeam, J.J. Giovannoni, J.K.C. Rose, I. Sorensen, S.J. Lee, R.W. Kim, I.Y. Choi, B.S. Choi, J.S. Lim, Y.H. Lee, and D. Choi. 2014. Genome sequence of the hot pepper provides insights into the evolution of pungency in Capsicum species. Nat. Genet. 46:270–278.
Kitamura, S. 1958. Varieties of radish and their transition, p. 1–19. In:I. Nishiyama(ed.). Japanese radish. The Japan Science Society Press, Tokyo, Japan.
Kitashiba, H., F. Li, H. Hirakawa, T. Kawanabe, Z.W. Zou, Y. Hasegawa, K. Tonosaki, S. Shirasawa, A. Fukushima, S. Yokoi, Y. Takahata, T. Kakizaki, M. Ishida, S. Okamoto, K. Sakamoto, K. Shirasawa, S. Tabata, and T. Nishio. 2014. Draft sequences of the radish (Raphanus sativus L.) genome. DNA Res. 21:481–490.
Kobabe, G. 1959. Naturliche Einkreuzung von Hederich (Raphanus raphanistrum L.) in Radies (Raphanus sativus var. radicula DC.) und das Verhalten von Knollenform und Farbe in den nachforgenden F-und R-Generationen. Zeitschrift Fur Pflanzenzuchtung 42:1–10.
Koressaar, T. and M. Remm. 2007. Enhancements and modifications of primer design program Primer3. Bioinformatics 23:1289–1291.
Kwon, Y.S., S.G. Park, and S.I. Yi. 2009. Assessment of genetic variation among commercial tomato (Solanum lycopersicum L.) varieties using SSR markers and morphological characteristics. Genes Genom. 31:1–10.
Kwon, Y.S., Y.H. Oh, S.I. Yi, H.Y. Kim, J.M. An, S.G. Yang, S.H. Ok, and J.S. Shin. 2010. Informative SSR markers for commercial variety discrimination in watermelon (Citrullus lanatus). GenesGenom. 32:115–122.
Nakatsuji, R., T. Hashida, N. Matsumoto, M. Tsuro, N. Kubo, and M. Hirai. 2011. Development of genomic and EST-SSR markers in radish (Raphanus sativus L.). Breed. Sci. 61:413–419.
Nei, M. 1978. Estimation of average heterozygosity and genetic distance from a small number of individuals. Genetics 89:583–590.
Pritchard, J.K., M. Stephens, and P. Donnelly. 2000. Inference of population structure using multilocus genotype data. Genetics 155:945–959.
Ramchiary, N., V.D. Nguyen, X.N. Li, C.P. Hong, V. Dhandapani, S.R. Choi, G. Yu, Z.Y. Piao, and Y.P. Lim. 2011. Genic microsatellite markers in Brassica rapa: Development, characterization, mapping, and their utility in other cultivated and wild Brassica relatives. DNA Res. 18:305–320.
Rohlf, F.J. 2008}. NTSYSpc: Numerical taxonomy system, ver.2.20. Exeter Publishing Ltd., Setauket, NY, USA.
Shirasawa, K., M. Oyama, H. Hirakawa, S. Sato, S. Tabata, T. Fujioka, C. Kimizuka-Takagi, S. Sasamoto, A. Watanabe, M. Kato, Y. Kishida, M. Kohara, C. Takahashi, H. Tsuruoka, T. Wada, T. Sakai, and S. Isobe. 2011. An EST-SSR linkage map of Raphanus sativus and comparative genomics of the Brassicaceae. DNA Res. 18: 221–232.
Sim, S., G. Durstewitz, J. Plieske, R. Wieseke, M.W. Ganal, A. Van Deynze, J.P. Hamilton, C.R. Buell, M. Causse, S. Wijeratne, and D.M. Francis. 2012. Development of a large SNP genotyping array and generation of high-density genetic maps in tomato. PLoS ONE 7:e40563.
Sim, S., J.K. Yu, Y.K. Jo, M.E. Sorrells, and G. Jung. 2009. Transferability of cereal EST-SSR markers to ryegrass. Genome 52:431–437.
Varshney, R.K., A. Graner, and M.E. Sorrells. 2005. Genic microsatellite markers in plants: Features and applications. Trends Biotechnol. 23:48–55.
Wang, S.F., X.F. Wang, Q.W. He, X.X. Liu, W.L. Xu, L.B. Li, J.W. Gao, and F.D. Wang. 2012. Transcriptome analysis of the roots at early and late seedling stages using Illumina paired-end sequencing and development of EST-SSR markers in radish. Plant Cell Rep. 31:1437–1447.
Wang, X.W., H.Z. Wang, J. Wang, R.F. Sun, J. Wu, S.Y. Liu, Y.Q. Bai, J.H. Mun, I. Bancroft, F. Cheng, S.W. Huang, X.X. Li, W. Hua, J.Y. Wang, X.Y. Wang, M. Freeling, J.C. Pires, A.H. Paterson, B. Chalhoub, B. Wang, A. Hayward, A.G. Sharpe, B.S. Park, B. Weisshaar, B.H. Liu, B. Li, B. Liu, C.B. Tong, C. Song, C. Duran, C.F. Peng, C.Y. Geng, C.S. Koh, C.Y. Lin, D. Edwards, D.S. Mu, D. Shen, E. Soumpourou, F. Li, F. Fraser, G. Conant, G. Lassalle, G.J. King, G. Bonnema, H.B. Tang, H.P. Wang, H. Belcram, H.L. Zhou, H. Hirakawa, H. Abe, H. Guo, H. Wang, H.Z. Jin, I.A.P. Parkin, J. Batley, J.S. Kim, J. Just, J.W. Li, J.H. Xu, J. Deng, J.A. Kim, J.P. Li, J.Y. Yu, J.L. Meng, J.P. Wang, J.M. Min, J. Poulain, J. Wang, K. Hatakeyama, K. Wu, L. Wang, L. Fang, M. Trick, M.G. Links, M.X. Zhao, M.N. Jin, N. Ramchiary, N. Drou, P.J. Berkman, Q.L. Cai, Q.F. Huang, R.Q. Li, S. Tabata, S.F. Cheng, S. Zhang, S.J. Zhang, S.M. Huang, S. Sato, S.L. Sun, S.J. Kwon, S.R. Choi, T.H. Lee, W. Fan, X. Zhao, X. Tan, X. Xu, Y. Wang, Y. Qiu, Y. Yin, Y.R. Li, Y.C. Du, Y.C. Liao, Y. Lim, Y. Narusaka, Y.P. Wang, Z.Y. Wang, Z.Y. Li, Z.W. Wang, Z.Y. Xiong, and Z.H. Zhang. 2011. The genome of the mesopolyploid crop species Brassica rapa. Nat. Genet. 43:1035–1039.
Weir, B.S. and C.C. Cockerham. 1984. Estimating F-statistics for the analysis of population-structure. Evolution 38:1358–1370.
Yagi, M., T. Kimura, T. Yamamoto, S. Isobe, S. Tabata, and T. Onozaki. 2012. QTL analysis for resistance to bacterial wilt (Burkholderia caryophylli) in carnation (Dianthus caryophyllus) using an SSR-based genetic linkage map. Mol. Breed. 30:495–509.
Yamagishi, H. 2004. Assessment of cytoplasmic polymorphisms by PCR-RFLP of the mitochondrial orfB region in wild and cultivated radishes. Plant Breed. 123:141–144.
Yamagishi, H. and T. Terachi. 2003. Multiple origins of cultivated radishes as evidenced by a comparison of the structural variations in mitochondrial DNA of Raphanus. Genome 46:89–94.
Yamane, K., N. Lu, and O. Ohnishi. 2005. Chloroplast DNA variations of cultivated radish and its wild relatives. Plant Sci. 168:627–634.
Yamane, K., N. Lu, and O. Ohnishi. 2009. Multiple origins and high genetic diversity of cultivated radish inferred from polymorphism in chloroplast simple sequence repeats. Breed. Sci. 59:55–65.
Yi, S.I., K.M. Bae, Y.S. Kwon, and I.H. Cho. 2006. Development of oriental melon (Cucumis melo L.)derived SSR markers using a PCR-based method and polymorphic application for the genotyping of commercial lines. Kor. J. Genet. 28:317–324.
Author information
Authors and Affiliations
Corresponding author
Additional information
These authors contributed equally to this work.
Electronic supplementary material
Rights and permissions
About this article
Cite this article
Bae, KM., Sim, SC., Hong, JH. et al. Development of genomic SSR markers and genetic diversity analysis in cultivated radish (Raphanus sativus L.). Hortic. Environ. Biotechnol. 56, 216–224 (2015). https://doi.org/10.1007/s13580-015-0089-y
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13580-015-0089-y