Abstract
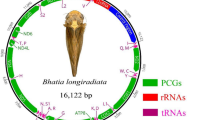
The complete mitochondrial genome of Cucullaea labiata (Arcoida: Cucullaeidae) was firstly determined in this study in order to better understand the phylogenetic relationship between Cucullaeidae and Arcidae. The C. labiata mitochondrial genome was 25,845 bp in size and contained 12 protein-coding genes, 2 rRNA and 22 tRNA genes. The number and the location of the tRNA genes were different from three Arcidae species (Scapharca broughtonii, Scapharca kagoshimensis and Tegillarca granosa). Gene arrangement also differed dramatically. The length of the non-coding regions was 10,559 bp, in which the largest one (6057 bp) included eight point nine copies of a 659 bp repeat motif. The number of repeated sequences was different in different individuals, similar to the findings from the mitochondrial genome of S. broughtonii and Placopecten magellanicus. One intron was found in cox1 gene both in CL_98 and in CL_99 individuals of C. labiata. The reason why mitochondrial introns are retained so scarcely in bivalve taxa needs further research. Phylogenetic analyses based on 12 concatenated amino acid sequences of protein-coding genes supported Cucullaeidae was the sister group of Arcidae.



Similar content being viewed by others
References
Abascal F, Zardoya R, Posada D (2005) ProtTest: selection of bestfit models of protein evolution. Bioinformatics 21:2104–2105
Benson G (1999) Tandem repeats finder: a program to analyze DNA sequences. Nucleic Acids Res 27:573–580
Boore JL (1999) Animal mitochondrial genomes. Nucleic Acids Res 27:1767–1780
Boore JL, Brown WM (2000) Mitochondrial genomes of Galathealinum, Helobdella, and Platynereis: sequence and gene arrangement comparisons indicate that Pogonophora is not a phylum and Annelida and Arthropoda are not sister taxa. Mol Biol Evol 17:87–106
Boss KJ (1982) Mollusca. In: Parker SP (ed) Synopsis and classification of living organisms, vol 1. McGraw-Hill, New York, p 1111
Burland TG (2000) DNASTAR’s Lasergene sequence analysis software. Methods Mol Biol 132:71–91
Castresana J (2000) Selection of conserved blocks from multiple alignments for their use in phylogenetic analysis. Mol Biol Evol 17:540–552
Drummond AJ, Rambaut A (2007) BEAST: Bayesian evolutionary analysis by sampling trees. BMC Evol Biol 7:214
Endo K, Noguchi Y, Ueshima R, Jacobs HT (2005) Novel repetitive structures, deviant protein-encoding sequences and unidentified ORFs in the mitochondrial genome of the brachiopod Lingula anatina. J Mol Evol 61:36–53
Feng YW, Li Q, Kong LF (2015) Molecular phylogeny of Arcoidea with emphasis on Arcidae species (Bivalvia: Pteriomorphia) along the coast of China: challenges to current classification of arcoids. Mol Phylogenet Evol 85:189–196
Fukami H, Chen CA, Chiou CY, Knowlton N (2007) Novel group I introns encoding a putative homing endonuclease in the mitochondrial cox1 gene of scleractinian corals. J Mol Evol 64:591–600
Hall TA (1999) BioEdit: a user-friendly biological sequence alignment editor and analysis program for Windows 95/98/NT. Nucleic Acids Symp Ser 95–98
Hofmann K, Stoffel W (1993) TMbase—a database of membrane spanning proteins segments. Biol Chem Hoppe-Seyler 374:166
Käll L, Krogh A, Sonnhammer ELL (2004) A combined transmembrane topology and signal peptide prediction method. J Mol Biol 338:1027–1036
Lang BF, Laforest MJ, Burger G (2007) Mitochondrial introns: a critical view. Trends Genet 23:119–125
Li Q, Park C, Kijima A (2002) Isolation and characterization of microsatellite loci in the Pacific abalone, Haliotis discus hannai. J Shellfish Res 21:811–815
Liu YG, Kurokawa T, Sekino M, Tanabe T, Watanabe K (2013) Complete mitochondrial DNA sequence of the ark shell Scapharca broughtonii: an ultra-large metazoan mitochondrial genome. Comp Biochem Physiol D 8:72–81
Lowe TM, Eddy SR (1997) tRNAscan-SE: a program for improved detection of transfer RNA genes in genomic sequence. Nucleic Acids Res 25:955–964
Lunt DH, Whipple LE, Hyman BC (1998) Mitochondrial DNA variable number tandem repeats (VNTRs): utility and problems in molecular ecology. Mol Ecol 7:1441–1455
Matsumoto M (2003) Phylogenetic analysis of the subclass Pteriomorphia (Bivalvia) from mtDNA COI sequences. Mol Phylogenet Evol 27:429–440
Meng X, Zhao N, Shen X, Hao J, Liang M, Zhu X, Cheng HL, Yan BL, Liu Z (2012) Complete mitochondrial genome of Coelomactra antiquata (Mollusca: Bivalvia): the first representative from the family Mactridae with novel gene order and unusual tandem repeats. Comp Biochem Physiol D 7:175–179
Oliver PG, Holmes AM (2006) The Arcoidea (Mollusca: Bivalvia): a review of the current phenetic-based systematics. Zool J Linn Soc 148:237–251
Palumbi SR (1996) Nucleic acids II: the polymerase chain reaction. In: Hillis D, Moritz C (eds) Molecular systematics. Sinauer, Sunderland, pp 205–247
Plazzi F, Ceregato A, Taviani M, Passamonti M (2011) A molecular phylogeny of bivalve mollusks: ancient radiations and divergences as revealed by mitochondrial genes. PLoS ONE 6:e27147
Pons JL, Labesse G (2009) @TOME-2: a new pipeline for comparative modeling of protein-ligand complexes. Nucleic Acids Res 37:W485–W491
Ronquist F, Huelsenbeck JP (2003) MrBayes 3: Bayesian phylogenetic inference under mixed models. Bioinformatics 19:1572–1574
Rot C, Goldfarb I, Ilan M, Huchon D (2006) Putative cross-kingdom horizontal gene transfer in sponge (Porifera) mitochondria. BMC Evol Biol 6:71
Shadel GS, Clayton DA (1997) Mitochondrial DNA maintenance in vertebrates. Annu Rev Biochem 66:409–435
Sharma PP, González VL, Kawauchi GY, Andrade SCS, Guzmán A, Collins TM, Glover EA, Harper EM, Healy JM, Mikkelsen PM et al (2012) Phylogenetic analysis of four nuclear protein-encoding genes largely corroborates the traditional classification of Bivalvia (Mollusca). Mol Phylogenet Evol 65:64–74
Smith DR, Snyder M (2007) Complete mitochondrial DNA sequence of the scallop Placopecten magellanicus: evidence of transposition leading to an uncharacteristically large mitochondrial genome. J Mol Evol 65:380–391
Snyder M, Fraser AR, LaRoche J, Gartner-Kepkay KE, Zouros E (1987) A typical mitochondrial DNA from the deep-sea scallop Placopecten magellanicus. Proc Natl Acad Sci USA 84:7595–7599
Steiner G, Hammer S (2000) Molecular phylogeny of the Bivalvia inferred from 18 S rDNA sequences with particular reference to the Pteriomorphia. Geol Soc Lond Spec Publ 177:11–29
Sun SE, Kong LF, Yu H, Li Q (2014) The complete mitochondrial genome of Scapharca kagoshimensis (Bivalvia: Arcidae). Mitochondrial DNA 1–2
Sun SE, Kong LF, Yu H, Li Q (2015) The complete mitochondrial DNA of Tegillarca granosa and comparative mitogenomic analyses of three Arcidae species. Gene 557:61–70
Waller TR (1998) Origin of the molluscan class Bivalvia and a phylogeny of major groups. Bivalves 1(4):5
Wang X, Lavrov DV (2008) Seventeen new complete mtDNA sequences reveal extensive mitochondrial genome evolution within the Demospongiae. PLoS ONE 3:e2723
Wolstenholme DR (1992) Animal mitochondrial DNA: structure and evolution. Int Rev Cytol 141:173–216
Wu XY, Xu XD, Yu ZN, Wei ZP, Xia JJ (2010) Comparison of seven Crassostrea mitogenomes and phylogenetic analyses. Mol Phylogenet Evol 57:448–454
Wyman SK, Jansen RK, Boore JL (2004) Automatic annotation of organellar genomes with DOGMA. Bioinformatics 20:3252–3255
Xu KF, Kanno M, Yu H, Li Q, Kijima A (2011) Complete mitochondrial DNA sequence and phylogenetic analysis of Zhikong scallop Chlamys farreri (Bivalvia: Pectinidae). Mol Biol Rep 38:3067–3074
Yamanoue Y, Miya M, Matsuura K, Yagishita N, Mabuchi K, Sakai H (2007) Phylogenetic position of tetraodontiform fishes within the higher teleosts: Bayesian inferences based on 44 whole mitochondrial genome sequences. Mol Phylogenet Evol 45:89–101
Yu ZN, Wei ZP, Kong XY, Shi W (2008) Complete mitochondrial DNA sequence of oyster Crassostrea hongkongensis—a case of tandem duplication-random loss for genome rearrangement in Crassostrea? BMC Genomics 9:477
Yuan Y, Li Q, Yu H, Kong LF (2012) The complete mitochondrial genomes of six heterodont bivalves (Tellinoidea and Solenoidea): variable gene arrangements and phylogenetic implications. PLoS ONE 7:e32353
Zdobnov EM, Apweiler R (2001) InterProScan—an integration platform for the signature—recognition methods in InterPro. Bioinformatics 17:847–848
Acknowledgements
This study was supported by research grants from the National Natural Science Foundation of China (41276138), and Fundamental Research Funds for the Central Universities.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
Yanwei Feng declares that he/she has no conflict of interest. Qi Li declares that he/she has no conflict of interest. Hong Yu declares that he/she has no conflict of interest. Lingfeng Kong declares that he/she has no conflict of interest.
Ethical approval
The research was conducted in the absence of any ethical issue on aquatic animal research.
Rights and permissions
About this article
Cite this article
Feng, Y., Li, Q., Yu, H. et al. Complete mitochondrial genome sequence of Cucullaea labiata (Arcoida: Cucullaeidae) and phylogenetic implications. Genes Genom 39, 867–875 (2017). https://doi.org/10.1007/s13258-017-0548-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13258-017-0548-1