Abstract
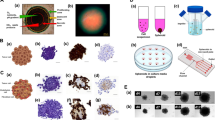
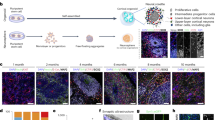
The iPSC-derived 3D models are considered to be a connective link between 2D culture and in vivo studies. However, the sensitivity of such 3D models is yet to be established. We assessed the sensitivity of the hiPSC-derived 3D spheroids against 2D cultures of neural progenitor cells. The sub-toxic dose of Sodium Arsenite (SA) was used to investigate the alterations in miRNA-proteins in both systems. Though SA exposure induced significant alterations in the proteins in both 2D and 3D systems, these proteins were uncommon except for 20 proteins. The number and magnitude of altered proteins were higher in the 2D system compared to 3D. The association of dysregulated miRNAs with the target proteins showed their involvement primarily in mitochondrial bioenergetics, oxidative and ER stress, transcription and translation mechanism, cytostructure, etc., in both culture systems. Further, the impact of dysregulated miRNAs and associated proteins on these functions and ultrastructural changes was compared in both culture systems. The ultrastructural studies revealed a similar pattern of mitochondrial damage, while the cellular bioenergetics studies confirm a significantly higher energy failure in the 2D system than to 3D. Such a higher magnitude of changes could be correlated with a higher amount of internalization of SA in 2D cultures than in 3D spheroids. Our findings demonstrate that a 2D culture system seems better responsive than a 3D spheroid system against SA exposure.
Graphical Abstract









Similar content being viewed by others
Data Availability
The raw data of proteomics are available on the CCMS-MassIVE (accession No. MassIVE MSV000092902) online data repository. The raw data are available in the repository of the Institute on its server, and with the corresponding author too.
References
Wang H, Brown PC, Chow ECY, Ewart L, Ferguson SS, Fitzpatrick S, Freedman BS, Guo GL et al (2021) 3D cell culture models: drug pharmacokinetics, safety assessment, and regulatory consideration. Clin Transl Sci 14(5):1659–1680. https://doi.org/10.1111/cts.13066
Pratumkaew P, Issaragrisil S, Luanpitpong S (2021) Induced pluripotent stem cells as a tool for modeling hematologic disorders and as a potential source for cell-based therapies. Cells 10(11):3250. https://doi.org/10.3390/cells10113250
Hogberg HT, Smirnova L (2022) The future of 3D brain cultures in developmental neurotoxicity testing. Front Toxicol 4:808620. https://doi.org/10.3389/ftox.2022.808620
Rowe RG, Daley GQ (2019) Induced pluripotent stem cells in disease modelling and drug discovery. Nat Rev Genet 20(7):377–388. https://doi.org/10.1038/s41576-019-0100-z
Jensen C, Teng Y (2020) Is it time to start transitioning from 2D to 3D cell culture? Front Mol Biosci 7:33. https://doi.org/10.3389/fmolb.2020.00033
Truskey GA (2018) Human microphysiological systems and organoids as in vitro models for toxicological studies. Front Public Health 6:185. https://doi.org/10.3389/fpubh.2018.00185
Lovett ML, Nieland TJF, Dingle YL, Kaplan DL (2020) Innovations in 3-dimensional tissue models of human brain physiology and diseases. Adv Funct Mater 30(44). https://doi.org/10.1002/adfm.201909146
Juarez-Moreno K, Chavez-Garcia D, Hirata G, Vazquez-Duhalt R (2022) Monolayer (2D) or spheroids (3D) cell cultures for nanotoxicological studies? Comparison of cytotoxicity and cell internalization of nanoparticles. Toxicol In Vitro 85:105461. https://doi.org/10.1016/j.tiv.2022.105461
Caipa Garcia AL, Arlt VM, Phillips DH (2022) Organoids for toxicology and genetic toxicology: applications with drugs and prospects for environmental carcinogenesis. Mutagenesis 37(2):143–154. https://doi.org/10.1093/mutage/geab023
Zhao C, Cai Z (2023) Mass spectrometry-based omics and imaging technique: a novel tool for molecular toxicology and health impacts. Rev Environ Contam Toxicol 261(1). https://doi.org/10.1007/s44169-023-00032-2
Promthep K, Nopparat C, Mukda S, Pannengpetch S, Wisomka P, Chantadul V, Phanchana M, Panmanee J (2022) Proteomic profiling reveals neuronal ion channel dysregulation and cellular responses to DNA damage-induced cell cycle arrest and senescence in human neuroblastoma SH-SY5Y cells exposed to cypermethrin. Neurotoxicology 93:71–83. https://doi.org/10.1016/j.neuro.2022.08.015
Srivastava AK, Yadav SS, Mishra S, Yadav SK, Parmar D, Yadav S (2020) A combined microRNA and proteome profiling to investigate the effect of ZnO nanoparticles on neuronal cells. Nanotoxicology 14(6):757–773. https://doi.org/10.1080/17435390.2020.1759726
Smirnova L, Seiler AEM, Luch A (2015) microRNA profiling as tool for developmental neurotoxicity testing (DNT). Curr Protoc Toxicol 64:20 29 21–20 29 22. https://doi.org/10.1002/0471140856.tx2009s64
Pamies D, Block K, Lau P, Gribaldo L, Pardo CA, Barreras P, Smirnova L, Wiersma D et al (2018) Rotenone exerts developmental neurotoxicity in a human brain spheroid model. Toxicol Appl Pharmacol 354:101–114. https://doi.org/10.1016/j.taap.2018.02.003
Du X, Luo L, Huang Q, Zhang J (2022) Cortex metabolome and proteome analysis reveals chronic arsenic exposure via drinking water induces developmental neurotoxicity through hnRNP L mediated mitochondrial dysfunction in male rats. Sci Total Environ 820:153325. https://doi.org/10.1016/j.scitotenv.2022.153325
Schultz L, Zurich MG, Culot M, da Costa A, Landry C, Bellwon P, Kristl T, Hormann K et al (2015) Evaluation of drug-induced neurotoxicity based on metabolomics, proteomics and electrical activity measurements in complementary CNS in vitro models. Toxicol In Vitro 30(1 Pt A):138–165. https://doi.org/10.1016/j.tiv.2015.05.016
Negi R, Srivastava A, Srivastava AK, Pandeya A, Vatsa P, Ansari UA, Pant AB (2023) Proteome architecture of human-induced pluripotent stem cell-derived three-dimensional organoids as a tool for early diagnosis of neuronal disorders. Indian J Pharmacol 55(2):108–118. https://doi.org/10.4103/ijp.ijp_56_23
Rajpurohit CS, Kumar V, Cheffer A, Oliveira D, Ulrich H, Okamoto OK, Zatz M, Ansari UA et al (2020) Mechanistic insights of astrocyte-mediated hyperactive autophagy and loss of motor neuron function in SOD1 L39R linked amyotrophic lateral sclerosis. Mol Neuro 57:4117–4133. https://doi.org/10.1007/s12035-020-02006-0
Roper SJ, Linke F, Scotting PJ, Coyle B (2021) 3D spheroid models of paediatric SHH medulloblastoma mimic tumour biology, drug response and metastatic dissemination. Sci Rep 11(1):4259. https://doi.org/10.1038/s41598-021-83809-6
Kumar V, Pandey A, Jahan S, Shukla RK, Kumar D, Srivastava A, Singh S, Rajpurohit CS et al (2016) Differential responses of Trans-Resveratrol on proliferation of neural progenitor cells and aged rat hippocampal neurogenesis. Sci Rep 6(1):28142. https://doi.org/10.1038/srep28142
Zhang Y, Xu S, Wu T, Hu K, Chen S, Xu A, Wu L (2018) Assessment of genotoxic effects by constructing a 3d cellular system with highly sensitive mutagenic human–hamster hybrid cells. Chem Res Toxicol 31(7):594–600. https://doi.org/10.1021/acs.chemrestox.8b00069
Yadav SK, Jauhari A, Singh N, Pandey A, Sarkar S, Pandey S, Garg RK, Parmar D et al (2023) Transcriptomics and proteomics approach for the identification of altered blood micrornas and plasma proteins in parkinson’s disease. Cel Mol Neuro 45:3527–3553. https://doi.org/10.1007/s10571-023-01362-4
Petyuk VA, Yu L, Olson HM, Yu F, Clair G, Qian WJ, Shulman JM, Bennett DA (2021) Proteomic profiling of the substantia nigra to identify determinants of lewy body pathology and dopaminergic neuronal loss. J Proteome Res 20(5):2266–2282. https://doi.org/10.1021/acs.jproteome.0c00747
Zhou Y, Zhou B, Pache L, Chang M, Khodabakhshi AH, Tanaseichuk O, Benner C, Chanda SK (2019) Metascape provides a biologist-oriented resource for the analysis of systems-level datasets. Nat Commun 10(1):1523. https://doi.org/10.1038/s41467-019-09234-6
Javed Z, Worley BL, Stump C, Shimko SS, Crawford LC, Mythreye K, Hempel N (2022) Optimization of extracellular flux assay to measure respiration of anchorage-independent tumor cell spheroids. Bio Protoc 12(4):e4321. https://doi.org/10.21769/BioProtoc.4321
Shi H, Rath EM, Lin RCY, Sarun KH, Clarke CJ, McCaughan BC, Ke H, Linton A et al (2022) 3-Dimensional mesothelioma spheroids provide closer to natural pathophysiological tumor microenvironment for drug response studies. Front Oncol 12:973576. https://doi.org/10.3389/fonc.2022.973576
Wang Q, Cohen JD, Yukawa T, Estrella H, Leonard C, Nunes J, Choi C, Lewis L et al (2022) Assessment of a 3D neural spheroid model to detect pharmaceutical-induced neurotoxicity. Altex 39(4):560–582. https://doi.org/10.14573/altex.2112221
Zhang Q, Nguyen AL, Shi S, Hill C, Wilder-Smith P, Krasieva TB, Le AD (2012) Three-dimensional spheroid culture of human gingiva-derived mesenchymal stem cells enhances mitigation of chemotherapy-induced oral mucositis. Stem Cells Dev 21(6):937–947. https://doi.org/10.1089/scd.2011.0252
Wolff A, Frank M, Staehlke S, Peters K (2022) A comparative study on the adipogenic differentiation of mesenchymal stem/stromal cells in 2D and 3D culture. Cells 11(8):1313. https://doi.org/10.3390/cells11081313
Regmi S, Raut PK, Pathak S, Shrestha P, Park P-H, Jeong J-HJA (2021) Enhanced viability and function of mesenchymal stromal cell spheroids is mediated via autophagy induction. Autophagy 17(10):2991–3010. https://doi.org/10.1080/15548627.2020.1850608
Thakur M, Rachamalla M, Niyogi S, Datusalia AK, Flora SJS (2021) Molecular mechanism of arsenic-induced neurotoxicity including neuronal dysfunctions. Int J Mol Sci 22(18). https://doi.org/10.3390/ijms221810077
Kapalczynska M, Kolenda T, Przybyla W, Zajaczkowska M, Teresiak A, Filas V, Ibbs M, Blizniak R et al (2018) 2D and 3D cell cultures - a comparison of different types of cancer cell cultures. Arch Med Sci 14(4):910–919. https://doi.org/10.5114/aoms.2016.63743
Angelopoulos I, Gakis G, Birmpas K, Kyrousi C, Habeos EE, Kaplani K, Lygerou Z, Habeos I et al (2022) Metabolic regulation of the neural stem cell fate: Unraveling new connections, establishing new concepts. Front Neurosci 16:1009125. https://doi.org/10.3389/fnins.2022.1009125
Raffaello A, Mammucari C, Gherardi G, Rizzuto R (2016) Calcium at the center of cell signaling: interplay between endoplasmic reticulum, mitochondria, and lysosomes. Trends Biochem Sci 41(12):1035–1049. https://doi.org/10.1016/j.tibs.2016.09.001
Braymer JJ, Lill R (2017) Iron-sulfur cluster biogenesis and trafficking in mitochondria. J Biol Chem 292(31):12754–12763. https://doi.org/10.1074/jbc.R117.787101
Chandravanshi LP, Gupta R, Shukla RK (2018) Developmental neurotoxicity of arsenic: involvement of oxidative stress and mitochondrial functions. Biol Trace Elem Res 186(1):185–198. https://doi.org/10.1007/s12011-018-1286-1
Hewitt T, Alural B, Tilak M, Wang J, Becke N, Chartley E, Perreault M, Haggarty SJ et al (2023) Bipolar disorder-iPSC derived neural progenitor cells exhibit dysregulation of store-operated Ca(2+) entry and accelerated differentiation. Mol Psychiatry 1–14. https://doi.org/10.1038/s41380-023-02152-6
Zheng Y, Shi Y, Tian C, Jiang C, Jin H, Chen J, Almasan A, Tang H et al (2004) Essential role of the voltage-dependent anion channel (VDAC) in mitochondrial permeability transition pore opening and cytochrome c release induced by arsenic trioxide. Oncogene 23(6):1239–1247. https://doi.org/10.1038/sj.onc.1207205
Chavan H, Christudoss P, Mickey K, Tessman R, Ni HM, Swerdlow R, Krishnamurthy P (2017) Arsenite effects on mitochondrial bioenergetics in human and mouse primary hepatocytes follow a nonlinear dose response. Oxid Med Cell Longev 2017:9251303. https://doi.org/10.1155/2017/9251303
Ding B, Ma X, Liu Y, Ni B, Lu S, Chen Y, Liu X, Zhang W (2023) Arsenic-Induced, mitochondria-mediated apoptosis is associated with decreased peroxisome proliferator-activated receptor γ coactivator α in rat brains. Toxics 11(7):576. https://doi.org/10.3390/toxics11070576
Lan J, Tang L, Wu S, Huang R, Zhong G, Jiang X, Tang Z, Hu L (2022) Curcumin alleviates arsenic-induced injury in duck skeletal muscle via regulating the PINK1/Parkin pathway and protecting mitochondrial function. Toxicol Appl Pharmacol 434:115820. https://doi.org/10.1016/j.taap.2021.115820
Zhong G, Hu T, Tang L, Li T, Wu S, Duan X, Pan J, Zhang H, Tang Z, Feng X (2022) Arsenic causes mitochondrial biogenesis obstacles by inhibiting the AMPK/PGC-1α signaling pathway and also induces apoptosis and dysregulated mitophagy in the duck liver. Ecotoxicol Environ Safy 230:113117. https://doi.org/10.1016/j.ecoenv.2021.113117
Chen Y, Liu X, Zhang Q, Wang H, Zhang R, Ge Y, Liang H, Li W et al (2023) Arsenic induced autophagy-dependent apoptosis in hippocampal neurons via AMPK/mTOR signaling pathway. Food Chem Toxicol 179:113954. https://doi.org/10.1016/j.fct.2023.113954
Cho J-H, Lee S, Jeon H, Kim AH, Lee W, Lee Y, Yang S, Yun J et al (2020) Tetrabromobisphenol A-induced apoptosis in neural stem cells through oxidative stress and mitochondrial dysfunction. Neurotox Res 38:74–85. https://doi.org/10.1007/s12640-020-00179-z
Ding B, Ma X, Liu Y, Ni B, Lu S, Chen Y, Liu X, Zhang W (2023) Arsenic-induced, mitochondria-mediated apoptosis is associated with decreased peroxisome proliferator-activated receptor γ coactivator α in rat brains. J Toxics 11(7):576. https://doi.org/10.3390/toxics11070576
Medda N, Patra R, Ghosh TK, Maiti S (2020) Neurotoxic mechanism of arsenic: synergistic effect of mitochondrial instability, oxidative stress, and hormonal-neurotransmitter impairment. Biol Trace Elem Res 198(1):8–15. https://doi.org/10.1007/s12011-020-02044-8
Zhang X, Yang H, Wang Y, Zhang J, Zhang H, Cao X, Hu T, Lin J et al (2023) Proteomic study on the mechanism of arsenic neurotoxicity in the rat cerebral cortex and the protective mechanism of dictyophora polysaccharides against arsenic neurotoxicity. ACS Chem Neuro. https://doi.org/10.1021/acschemneuro.3c00009
Jovanovic B, Eiermann N, Talwar D, Boulougouri M, Dick TP, Stoecklin G (2021) Thioredoxin 1 is required for stress granule assembly upon arsenite-induced oxidative stress. Food Chem Toxicol 156:112508. https://doi.org/10.1016/j.fct.2021.112508
Masjosthusmann S, Blum J, Bartmann K, Dolde X, Holzer AK, Stürzl LC, Keßel EH, Förster N, Dönmez A, Klose J (2020) Establishment of an a priori protocol for the implementation and interpretation of an in‐vitro testing battery for the assessment of developmental neurotoxicity. J EFSA Supporting Publications 17 (10):1938E
Yu JT, Liu Y, Dong P, Cheng RE, Ke SX, Chen KQ, Wang JJ, Shen ZS et al (2019) Up-regulation of antioxidative proteins TRX1, TXNL1 and TXNRD1 in the cortex of PTZ kindling seizure model mice. PLoS ONE 14(1):e0210670. https://doi.org/10.1371/journal.pone.0210670
Ali R, Alhaj Sulaiman A, Memon B, Pradhan S, Algethami M, Aouida M, McKay G, Madhusudan S et al (2023) Altered regulation of the glucose transporter GLUT3 in PRDX1 null cells caused hypersensitivity to arsenite. Cells 12(23):2682. https://doi.org/10.3390/cells12232682
Prakash C, Soni M, Kumar V (2016) Mitochondrial oxidative stress and dysfunction in arsenic neurotoxicity: a review. J Appl Toxicol 36(2):179–188. https://doi.org/10.1002/jat.3256
Kodron A, Mussulini BH, Pilecka I, Chacinska A (2021) The ubiquitin-proteasome system and its crosstalk with mitochondria as therapeutic targets in medicine. Pharmacol Res 163:105248. https://doi.org/10.1016/j.phrs.2020.105248
Alsayyah C, Ozturk O, Cavellini L, Belgareh-Touze N (1861) Cohen MM (2020) The regulation of mitochondrial homeostasis by the ubiquitin proteasome system. Biochim Biophys Acta Bioenerg 12:148302. https://doi.org/10.1016/j.bbabio.2020.148302
Tam LM, Wang Y (2020) Arsenic exposure and compromised protein quality control. Chem Res Toxicol 33(7):1594–1604. https://doi.org/10.1021/acs.chemrestox.0c00107
Dodson M, de la Vega MR, Harder B, Castro-Portuguez R, Rodrigues SD, Wong PK, Chapman E, Zhang DD (2018) Low-level arsenic causes proteotoxic stress and not oxidative stress. Toxicol Appl Pharmacol 341:106–113. https://doi.org/10.1016/j.taap.2018.01.014
Dodson M, Liu P, Jiang T, Ambrose AJ, Luo G, Rojo de la Vega M, Cholanians AB, Wong PK et al (2018) Increased O-GlcNAcylation of SNAP29 drives arsenic-induced autophagic dysfunction. Mol Cell Biol 38 (11). https://doi.org/10.1128/MCB.00595-17
Turakhiya A, Meyer SR, Marincola G, Bohm S, Vanselow JT, Schlosser A, Hofmann K, Buchberger A (2018) ZFAND1 recruits p97 and the 26S proteasome to promote the clearance of arsenite-induced stress granules. Mol Cell 70(5):906-919 e907. https://doi.org/10.1016/j.molcel.2018.04.021
Zhao P, Guo Y, Zhang W, Chai H, Xing H, Xing M (2017) Neurotoxicity induced by arsenic in Gallus Gallus: regulation of oxidative stress and heat shock protein response. Chemosphere 166:238–245. https://doi.org/10.1016/j.chemosphere.2016.09.060
Advani VM, Ivanov P (2019) Translational control under stress: reshaping the translatome. BioEssays 41(5):e1900009. https://doi.org/10.1002/bies.201900009
Low YH, Asi Y, Foti SC, Lashley T (2021) Heterogeneous nuclear ribonucleoproteins: implications in neurological diseases. Mol Neurobio 58(2):631–646. https://doi.org/10.1007/s12035-020-02137-4
Medda N, De SK, Maiti S (2021) Different mechanisms of arsenic related signaling in cellular proliferation, apoptosis and neo-plastic transformation. Ecotoxicol Environ Saf 208:111752. https://doi.org/10.1016/j.ecoenv.2020.111752
Pandey R, Rai V, Mishra J, Mandrah K, Kumar Roy S, Bandyopadhyay S (2017) From the cover: arsenic induces hippocampal neuronal apoptosis and cognitive impairments via an up-regulated BMP2/Smad-dependent reduced BDNF/TrkB signaling in rats. Toxicol Sci 159(1):137–158. https://doi.org/10.1093/toxsci/kfx124
He Z, Zhang Y, Zhang H, Zhou C, Ma Q, Deng P, Lu M, Mou Z et al (2021) NAC antagonizes arsenic-induced neurotoxicity through TMEM179 by inhibiting oxidative stress in Oli-neu cells. Ecotoxicol Environ Saf 223:112554. https://doi.org/10.1016/j.ecoenv.2021.112554
Chen G, Wang M, Ruan Z, Zhu L, Tang C (2021) Mesenchymal stem cell-derived exosomal miR-143-3p suppresses myocardial ischemia-reperfusion injury by regulating autophagy. Life Sci 280:119742. https://doi.org/10.1016/j.lfs.2021.119742
Malakootian M, Naeli P, Mowla SJ, Seidah NG (2022) Post-transcriptional effects of miRNAs on PCSK7 expression and function: miR-125a-5p, miR-143–3p, and miR-409–3p as negative regulators. Metabolites 12 (7). https://doi.org/10.3390/metabo12070588
Xu L, Chen W, Chen J, Jin Y, Ma W, Qi G, Sun X, Luo J et al (2022) PIWI-interacting RNA-23210 protects against acetaminophen-induced liver injury by targeting HNF1A and HNF4A. Biochem Pharmacol 197:114897. https://doi.org/10.1016/j.bcp.2021.114897
Acknowledgements
Dr Renu Negi is grateful to CSIR-UGC, New Delhi, for providing Research Fellowship. The authors are grateful to Ms Deepshikha Srivastava, Technical Officer, LC-MS/MS Laboratory, and Mr Jai Shankar, Technical Officer, Microscope Laboratory, Central Instrumentation Facility, CSIR-Indian Institute of Toxicology Research, Lucknow, India, for extending the operational support for LC-MS/MS during proteomics studies and Transmission Electron Microscopic study.
Funding
The research was funded by the Indian Council of Medical Research (ICMR), New Delhi, India (Grant Sanction number: 5/4–5/3/9/DHR/Neuro/2021/NCD-1 dated 30/03/2022).
Author information
Authors and Affiliations
Contributions
ABP has conceptualized the research work, designed the protocols, acquired the funds, project administration work, supervised throughout experimentations, and reviewed and edited the manuscript. RN, AS, and AKS performed the experiments, data curation, and analysis and prepared the original draft of the manuscript. PV and UAA helped with the data analysis writing and editing of the manuscript. BK and HS assisted in carrying out the parts of the Open Array and Real-Time PCR experiments. AP helps in the investigation, formal analysis, and visualization of the data.
Corresponding author
Ethics declarations
Ethical Approval
Not applicable.
Consent to Participate
Not applicable.
Consent for Publication
All the authors have read the contents of the manuscript and have provided their consent to publish the data.
Competing Interests
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Negi, R., Srivastava, A., Srivastava, A.K. et al. Proteomic-miRNA Biomics Profile Reveals 2D Cultures of Human iPSC-Derived Neural Progenitor Cells More Sensitive than 3D Spheroid System Against the Experimental Exposure to Arsenic. Mol Neurobiol (2024). https://doi.org/10.1007/s12035-024-03924-z
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s12035-024-03924-z