Abstract
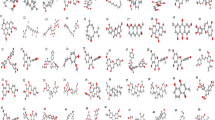
Since N-cadherin protein plays a remarkable role in cancer metastasis and tumor growth and progression, finding new effective inhibitors of this protein can be of high importance in cancer treatment. Nevertheless, few molecules have been introduced to inhibit N-cadherin protein to date. In this work, in order to find and present potent inhibitors, 3358 FDA-approved small molecules were docked against N-cadherin protein. All complexes with binding energy − 9 to − 8 kcal/mol were selected for protein-ligand interaction analysis. In the following, Tanimoto coefficient (Tc) was calculated for those molecules that established appropriate interactions with N-cadherin in order to compute the similarity score between them. Afterwards, molecular dynamics simulation and free energy calculations were done to estimate the stability and ability of the chosen ligands in complex with the target protein. Finally, seven small molecules among 3358 FDA-approved were suggested as potential inhibitors of N-cadherin protein.












Similar content being viewed by others
References
Gumbiner BM (2005) Regulation of cadherin-mediated adhesion in morphogenesis. Nat Rev Mol Cell Biol 6(8):622–634
Hatta K, Okada TS, Takeichi M (1985) A monoclonal antibody disrupting calcium-dependent cell-cell adhesion of brain tissues: possible role of its target antigen in animal pattern formation. Proc Natl Acad Sci U S A 82(9):2789–2793
Grunwald GB et al (1981) Enzymatic dissection of embryonic cell adhesive mechanisms. II. Developmental regulation of an endogenous adhesive system in the chick neural retina. Dev Biol 86(2):327–338
Cook JH, Lilien J (1982) The accessibility of certain proteins on embryonic chick neural retina cells to iodination and tryptic removal is altered by calcium. J Cell Sci 55:85–103
Takeichi M (1995) Morphogenetic roles of classic cadherins. Curr Opin Cell Biol 7(5):619–627
Shan WS et al (2000) Functional cis-heterodimers of N- and R-cadherins. J Cell Biol 148(3):579–590
Burden-Gulley SM et al (2009) Novel peptide mimetic small molecules of the HAV motif in N-cadherin inhibit N-cadherin-mediated neurite outgrowth and cell adhesion. Peptides 30(12):2380–2387
Perotti A et al (2009) Clinical and pharmacological phase I evaluation of Exherin (ADH-1), a selective anti-N-cadherin peptide in patients with N-cadherin-expressing solid tumours. Ann Oncol 20(4):741–745
Yeung KT, Yang J (2017) Epithelial-mesenchymal transition in tumor metastasis. Mol Oncol 11(1):28–39
Casal JI, Bartolome RA, N-Cadherin B (2019) Relevance of cadherins 5, 6 and 17 in cancer progression and metastasis. Int J Mol Sci 20(13)
Nalla AK et al (2011) N-cadherin mediates angiogenesis by regulating monocyte chemoattractant protein-1 expression via PI3K/Akt signaling in prostate cancer cells. Exp Cell Res 317(17):2512–2521
Zheng X et al (2015) Epithelial-to-mesenchymal transition is dispensable for metastasis but induces chemoresistance in pancreatic cancer. Nature 527(7579):525–530
Fischer KR et al (2015) Epithelial-to-mesenchymal transition is not required for lung metastasis but contributes to chemoresistance. Nature 527(7579):472–476
Blaschuk OW (2015) N-cadherin antagonists as oncology therapeutics. Philos Trans R Soc Lond Ser B Biol Sci 370(1661):20140039
Craik DJ et al (2013) The future of peptide-based drugs. Chem Biol Drug Des 81(1):136–147
Sliwoski G et al (2014) Computational methods in drug discovery. Pharmacol Rev 66(1):334–395
Baldi A (2010) Computational approaches for drug design and discovery: an overview. System Rev Pharm 1(1):99
Huang H-J et al (2010) Current developments of computer-aided drug design. J Taiwan Inst Chem Eng 41(6):623–635
Joseph-McCarthy D (1999) Computational approaches to structure-based ligand design. Pharmacol Ther 84(2):179–191
Eslami M, et al (2018) Deep analysis of N-cadherin/ADH-1 interaction: a computational survey. J Biomol Struct Dyn 1–19
Lionta E et al (2014) Structure-based virtual screening for drug discovery: principles, applications and recent advances. Curr Top Med Chem 14(16):1923–1938
Walters WP, Stahl MT, Murcko MA (1998) Virtual screening—an overview. Drug Discov Today 3(4):160–178
Shoichet BK (2004) Virtual screening of chemical libraries. Nature 432(7019):862
Dallakyan S, Olson AJ (2015) Small-molecule library screening by docking with PyRx, Chem Biol 243–250. Springer
Trott O, Olson AJ (2010) AutoDock Vina: improving the speed and accuracy of docking with a new scoring function, efficient optimization, and multithreading. J Comput Chem 31(2):455–461
Abraham M, et al. GROMACS User Manual Version. 2014;5(2)
Paissoni C et al (2015) GMXPBSA 2.1: a GROMACS tool to perform MM/PBSA and computational alanine scanning. Comput Phys Commun 186:105–107
Biovia DS (2017) Discovery studio modeling environment. Release
Pettersen EF et al (2004) UCSF Chimera—a visualization system for exploratory research and analysis. J Comput Chem 25(13):1605–1612
Hajiebrahimi A, Ghasemi Y, Sakhteman A (2017) FLIP: an assisting software in structure based drug design using fingerprint of protein-ligand interaction profiles. J Mol Graph Model 78:234–244
Durrant JD, McCammon JA (2011) Molecular dynamics simulations and drug discovery. BMC Biol 9(1):71
Kabsch W, Sander C (1983) Dictionary of protein secondary structure: pattern recognition of hydrogen-bonded and geometrical features. Biopolymers 22(12):2577–2637
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Khajeh, S., Eslami, M., Nezafat, N. et al. Surveying FDA-approved drugs as new potential inhibitors of N-cadherin protein: a virtual screening approach. Struct Chem 31, 2355–2369 (2020). https://doi.org/10.1007/s11224-020-01595-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11224-020-01595-9