Abstract
Background
The outbreak of coronavirus disease 2019 (COVID-19) has emerged as a serious public health emergency of global concern. Angiotensin converting enzyme 2 (ACE2) peptidase domain is important for the cellular entry of severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2). Germline variants in ACE2 peptidase domain may influence the susceptibility for SARS-CoV-2 infection and disease severity in the host population. ACE2 genetic analysis among Caucasians showed inconclusive results. This is the first Asian study investigating the contribution of ACE2 germline variants to SARS-CoV-2 infection in Pakistani population.
Methods
In total, 442 individuals, including SARS-CoV-2-positive (n = 225) and SARS-CoV-2-negative (n = 217) were screened for germline variants in ACE2 peptidase domain (exons 2, 3, 9, and 10) using high resolution melting and denaturing high-performance liquid chromatography analyses followed by DNA sequencing of variant fragments. The identified variant was analyzed by in silico tools for potential effect on ACE2 protein.
Results
A missense variant, p.Lys26Arg, was identified in one SARS-CoV-2-positive (1/225; 0.4%) and three SARS-CoV-2-negative (3/217; 1.4%) individuals. No significant difference in the minor allele frequency of this variant was found among SARS-CoV-2-positive and SARS-CoV-2-negative individuals (1/313; 0.3% versus 3/328; 0.9%; P = 0.624), respectively. The SARS-CoV-2-positive patient carrying p.Lys26Arg showed mild COVID-19 disease symptoms. It was predicted as benign variant by in silico tool. No variant was detected in ACE2 residues important for binding of SARS-CoV-2 spike protein.
Conclusion
The p.Lys26Arg variant may have no association with SARS-CoV-2 susceptibility in Pakistani population. Whole ACE2 gene screening is warranted to clarify its role in SARS-CoV-2 infection.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
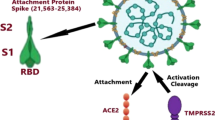
The outbreak of coronavirus disease 2019 (COVID-19) has emerged as a serious pandemic and accounted for 617.60 million cases and 6.53 million deaths worldwide, by October 8, 2022 (https://covid19.who.int/). A novel coronavirus-2019 (2019-nCoV), also named as severe acute respiratory syndrome coronavirus-2 (SARS-CoV-2) causes this disease [1]. SARS-CoV-2 infection primarily depends on the interaction of its spike protein with the host receptor, angiotensin converting enzyme 2 (ACE2). ACE2 is the cell membrane receptor of SARS-CoV-2 that mediates viral entry into cells [2,3,4].
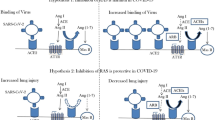
The human ACE2 gene (NM_001371415.1) is on the short arm of X chromosome (Xp22.2). It has 19 exons that encodes a protein of 805 amino acid residues and has two functional domains: N-terminal peptidase domain (residues 19 to 615) and C-terminal collectrin-like domain (residues 616 to 768) [5]. ACE2 is expressed in epithelial cells of the lung, intestine, kidney, blood vessels [6], and on the oral mucosa [7]. A recent structural analysis showed several key residues (K417, Y453, Q474, F486, Q498, T500, and N501) of the receptor binding motif in the SARS-CoV-2 spike protein receptor-binding domain (RBD) that interact with the ACE2 peptidase domain (residues Q24, D30, H34, Y41, Q42, M82, K353, and R357) [2, 5]. Another study revealed 17 residues of SARS-CoV-2 RBD contact 20 residues of the ACE2 (residues Q24, T27, F28, D30, K31, H34, E35, E37, D38, Y41, Q42, L79, M82, Y83, N330, K353, G354, D355, R357, and R393) that are encoded by ACE2 exon 2, 3, 9, and 10 [8]. Taken together, these studies showed that certain amino acid residues in the ACE2 peptidase domain are important for the recognition and cellular entry of SARS-CoV-2.
ACE2 receptor genetic variability may affect the virion entry into the host cell and thus the disease severity [9]. Cao and colleagues investigated the ACE2 in public databases and identified 32 coding variants that affect ACE2 protein [10]. The minor allele frequencies (MAF) of ACE2 variants differ in different populations, suggesting a variable influence [10]. Subsequently, several variants are reported to perturb the ACE2 stabilization, and interfere with interaction with viral spike protein [11]. The epidemiological data reported a disproportional spread of COVID-19 across various populations. It is transmissible in community settings. Various local clusters of COVID-19 are also reported [12]. However, in such clusters, not all individuals in close contact with confirmed COVID-19 patients acquired this infection [12]. This differential pattern of infection suggests distinct ACE2 genetic variants might influence SARS-CoV-2 susceptibility in different populations [13, 14]. These studies warrant ACE2 variants screening in original populations to predict the susceptibility and severity to COVID-19. Screening of ACE2 variants in Caucasian populations have reported controversial results regarding SARS-CoV-2 infection and COVID-19 disease outcomes [11, 15,16,17,18,19,20]. Since ACE2 genetic variability is unknown in COVID-19 patients in Asian populations, the present study aimed to investigate germline variants in peptidase domain (ACE2 exons 2, 3, 9, and 10; amino acids 1 to 115 and 301 to 433) of ACE2 in COVID-19 patients from Pakistan.
Materials and methods
Study subjects
This study includes 442 Pakistani individuals (≥ 18 years) who were tested for SARS-CoV-2 RNA using reverse transcription polymerase chain reaction (RT-PCR) at the Shaukat Khanum Memorial Cancer Hospital and Research Centre (SKM CH&RC), Lahore between June 07, 2020, and June 28, 2020. Of these, 225 SARS-CoV-2-positive and 217 SARS-CoV-2-negative individuals were retrospectively identified. The blood samples from these participants were collected simultaneously with the nasal swabs as per the National Institute of Health, Islamabad guidelines. Demographics and clinical data of all study participants were collected from the electronic medical records of SKM CH&RC. The Institutional Review Board of the SKM CH&RC approved the study (approval number IRB-20-15-A1) and granted the waiver of the written informed consent from study participants.
Molecular analyses
Genomic DNA was extracted from 3 ml to 5 ml of whole blood, as previously described [21]. Comprehensive screening for ACE2 (NM_001371415.1) exons 2, 3, 9, and 10 (coding for peptidase domain) was performed by High-Resolution Melting (HRM) Analysis using LightCycler 480-II (Roche Diagnostics, Indianapolis, IN, USA) or Denaturing High Performance Liquid Chromatography (DHPLC) analysis using Wave® DNA Fragment Analysis System (Transgenomic Inc., Omaha, NE, USA). Since the ACE2 is on X chromosome, hemizygous DNA of unknown male samples were mixed 1:1 with wildtype male DNA prior to the amplification, as previously described [22]. The primers were designed using Primer3 software freely available online (https://www.bioinformatics.nl/cgi-bin/primer3plus/primer3plus). Primer sequences, the setup of polymerase chain reaction (PCR), thermal cycling conditions, and the DHPLC running conditions are available upon request. Each sample revealing variants detected by either HRM or DHPLC analysis was directly sequenced using BigDye Terminator v.3.1 Cycle Sequencing Kit (Applied Biosystems, California, USA). The sequencing reactions were purified with BigDye XTerminator Purification Kit (Applied Biosystems, California, USA) and sequenced on an automated 3500 Genetic Analyzer (Applied Biosystems, California, USA) according to the manufacturer’s instructions. Bidirectional genomic DNA sequencing was performed to confirm the presence of a variant.
In silico analysis
A missense variant was analyzed for the potential effect on protein function using the default settings of web tool VarCards that annotates the coding variants using more than 60 genomic data sources, including 23 in silico algorithms, to comprehensively obtain predictive score and pathogenicity consequences of missense variants [23].
Statistical analysis
The quantitative variables (ages and vital signs) of the study participants were presented as medians/means along with ranges. The categorical variables (gender, travel and contact history, symptoms, comorbidities, radiological findings, disease severity, assessment outcomes, and MAF of c.77 A > G variant) were presented as frequencies and corresponding percentages. The quantitative and categorical variables data were analyzed using the independent t-test and Fisher’s exact test, respectively. All statistical test were two-sided, and the groups were considered as statistically different at P-value < 0.05. All statistical analyses were performed using IBM SPSS Statistics 20.0 software.
Results
In total, 442 individuals, comprising 225 SARS-CoV-2-positive and 217 SARS-CoV-2-negative, were included in this study. The median age at enrollment was 47.9 years (range 18.2–77.5) for male individuals (n = 243) and 47.0 years (range 18.0–76.0) for female individuals (n = 199). Compared to SARS-CoV-2-negative individuals, SARS-CoV-2-positive individuals were predominantly male (P = 0.013), reported a contact history with confirmed or suspected COVID-19 patients (P = 0.002), presented with COVID-19 disease symptoms (< 0.0001), had a high body temperature (P = 0.002), showed mild to severe COVID-19 disease (P < 0.0001), and had a longer duration of hospital stay (P = 0.013). The demographic and clinical characteristics of the study participants are presented in Table 1.
ACE2 exons 2, 3, 9, and 10 were screened for genetic variants using HRM and DHPLC, followed by DNA sequencing. No variants were detected in exon 3, 9, and 10 in SARS-CoV-2-positive and SARS-CoV-2-negative individuals. Only one missense variant in exon 2; c.77 A > G (p.Lys26Arg) (rs4646116) was identified in the cohort. Among females, it was identified as a heterozygote in one SARS-CoV-2-positive individual (1/88; 1.1%) and two SARS-CoV-2-negative individuals (2/111; 1.8%). Among males, it was identified as a hemizygous in one SARS-CoV-2-negative individual (1/106; 0.9%) but not among SARS-CoV-2-positive individuals (n = 137; 0%). There was no significant difference (P = 0.624) in the MAF of c.77 A > G among SARS-CoV-2-positive and SARS-CoV-2-negative (1/313; 0.3% vs. 3/328; 0.9%) individuals, respectively (Table 2). There was also no difference (P = 1.0) in the MAF of c.77 A > G between SARS-CoV-2-positive female and male individuals (1/176; 0.6% vs. 0/137; 0%), respectively (Table 2). It has been reported as a rare variant among South Asians (MAF = 0.13%; 25/19,044 (gnomAD database: https://gnomad.broadinstitute.org/variant/X-15618958-T-C?dataset=gnomad_r2_1 date last accessed: October 10, 2022). It was predicted as benign by all the 21 protein function prediction algorithms integrated in VarCards with a damaging score of 0. It has also been reported as a likely benign variant in the ClinVar database (https://www.ncbi.nlm.nih.gov/clinvar/variation/780203/?oq=rs4646116&m=NM_001371415.1(ACE2):c.77 A%3EG%20(p.Lys26Arg; date last accessed: October 10, 2022). Based on these findings, p.Lys26Arg was classified as likely benign.
The three SARS-CoV-2-negative individuals, including two females and one male harboring the p.Lys26Arg variant, remained asymptomatic for COVID-19 disease. The solitary SARS-CoV-2-positive female carrying the p.Lys26Arg variant was presented with mild COVID-19 disease symptoms of fever, cough, and sore throat.
Discussion
The clinical presentation of COVID-19 disease is highly variable, ranging from asymptomatic infection to severe form of pneumonia and respiratory or multi-organ failure. This is intriguing that genetic variability may influence the susceptibility to SARS-CoV-2 infection and its severity. The relevance of germline ACE2 variants as a screening target for the risk assessment of SARS-CoV-2 infection is of interest. Most of such studies have investigated the association between ACE2 variants and SARS-CoV-2 infection or COVID-19 disease severity using bioinformatics approaches while utilizing the frequencies of ACE2 genetic variants from public databases [10, 14, 24,25,26,27,28]. Only two European studies from Germany [17] and Spain [29] investigated the contribution of ACE2 variants to SARS-CoV-2 susceptibility among SARS-CoV-2-positive and SARS-CoV-2-negative individuals. To our knowledge, this is the first Asian study to screen ACE2 germline variants in the peptidase domain in the Pakistani population.
No germline variant was detected in any of the 20 ACE2 residues, previously inferred for binding of SARS-CoV-2 spike protein. This is consistent with previous reports [11], 16, 18, 20, 29, 30]. Variants in most of these residues are absent or present with a low frequency (MAF < 0.0006) in the general population (gnomAD database). These findings highlight that coding variants in the ACE2 peptidase domain are very rare and may not confer an increased susceptibility or resistance to SARS-CoV-2 infection.
In the present study, only one ACE2 missense variant c.77 A > G (p.Lys26Arg) was identified with a frequency of 0.44% (1/225) in SARS-CoV-2-positive and 1.38% (3/217) in SARS-CoV-2-negative Pakistani individuals. The allele frequency of c.77 A > G among SARS-CoV-2-positive (MAF = 0.3%) and SARS-CoV-2-negative (MAF = 0.9%) individuals was not significantly different (P = 0.624). In consistent with our results, this variant has been reported in a German population with a frequency of 0.34% (1/297) in SARS-CoV-2-positive and 1.18% (3/253) in SARS-CoV-2-negative individuals [17]. Recently, a high prevalence of this variant (10/164; 6.1%) was reported of in SARS-CoV-2-positive patients from Serbia [15]. However, it was not detected in SARS-CoV-2-positive patients from Italy [11, 18], Spain [29], and Russia [20]. This variant has been reported in gnomAD with MAF of 0.388% and 0.131% in the general population and South Asians, respectively. In our study the solitary SARS-CoV-2-positive patient carrying p.Lys26Arg variant showed mild COVID-19 disease symptoms in agreement with previous reports from Serbia [30] and Germany [17]. Collectively, these findings suggest that the p.Lys26Arg variant may neither be associated with increased SARS-CoV-2 infection nor depict a protective effect.
In the current study, p.Lys26Arg was predicted as a benign variant using VarCards in silico analysis tool. Consistently, this variant has been previously classified as a benign variant using various in silico analysis tools including SIFT, MUTTASTER, PROVEAN, PolyPhen-2, CADD, and REVEL [24, 31, 32]. Other studies, based on structural modeling and molecular docking simulations suggested that the p.Lys26Arg increased binding affinity of SARS-CoV-2 spike protein to the ACE2 receptor and showed increased SARS-CoV-2 infection as compared to the wildtype ACE2 [9, 14, 24, 26, 33, 34]. However, this is contradictory to the p.Lys26Arg frequencies in public databases or previous studies on SARS-CoV-2-negative and SARS-CoV-2-positive individuals [17, 29]. Other computational chemistry reports showed that p.Lys26Arg decreased the SARS-CoV-2/ACE2 electrostatic attraction and binding affinity to the host receptor [25, 27, 28]. Further, functional assays and screening of the p.Lys26Arg variant in larger populations of diverse ethnicities and geographical origin may address this discrepancy.
We restricted our analysis to the ACE2 peptidase domain variants screening, which have been established as RBD contact residues for SARS-CoV-2. Previous studies investigated the few ACE2 variants [17, 19, 35,36,37] or screened the complete gene [11, 20, 29, 38,39,40] and reported conflicting results. No association was found in Spanish [29, 39], Turkish [41], Russian [20], and British [40] populations. In contrast, different ACE2 variants were identified in Saudi [36], German [17], Italian [35], Spanish [19], and Polish [37] populations, that were linked with genetic susceptibility to SARS-CoV-2 or severity of the COVID-19 disease. However, most of these variants were in the noncoding region of the ACE2. Previously, it has been shown that noncoding variants may perturb the ACE2 gene activity [18]. These reports suggest that ACE2 variants, particularly in noncoding regions, may have an impact on infection and disease severity of COVID-19 among diverse populations. Complete ACE2 gene screening of in larger populations would be helpful for the clear understanding of its genetic association with SARS-CoV-2 infection and the course of COVID-19 disease.
Strength of the current study includes the size of study population that was well calculated through Online Sample Size Estimator (http://osse.bii.a-star.edu.sg/calculation1.php), complying by scientific standards for sample size. Another strength is that all study participants were tested by RT-PCR and stratified as SARS-CoV-2-positive or SARS-CoV-2-negative individuals. Whereas, previous studies reported the prevalence of ACE2 variants in SARS-CoV-2-positive individuals in comparison with healthy controls who were not tested for SARS-CoV-2 [11, 18, 20, 35, 36, 38, 39]. Limitation of this study includes the screening of only partial coding sequence (exons 2, 3, 9, and 10) of ACE2 peptidase domain. Variant screening was performed using HRM and DHPLC assay with variant detection sensitivity < 100%, that may have missed some of the ACE2 variants. Functional variants in other coding region of ACE2 may influence COVID-19 disease occurrence and severity in the Pakistani population.
Conclusion
In summary, this is the first study investigating the germline ACE2 variants in SARS-CoV-2-positive and SARS-CoV-2-negative individuals from Pakistan. No variants were detected in any of the 20 ACE2 residues, previously reported for binding of SARS-CoV-2 spike protein. The only identified missense variant, (p.Lys26Arg) may not be associated with increased susceptibility or protective effect for SARS-CoV-2 infection in the Pakistani population. Screening of the complete ACE2 gene is warranted to clarify its role in SARS-CoV-2 infection in the Pakistani population.
Data Availability
The datasets generated and/or analyzed during current study are available from the corresponding author on reasonable request.
Abbreviations
- ACE2:
-
Angiotensin converting enzyme 2.
- COVID 19:
-
Coronavirus disease 2019.
- DHPLC:
-
Denaturing High-performance Liquid Chromatography.
- HRM:
-
High Resolution Melting.
- MAF:
-
Minor allele frequency.
- PCR:
-
Polymerase chain reaction.
- PD domain:
-
Peptidase domain.
- RBD:
-
Receptor binding domain.
- SARS-CoV-2:
-
Severe acute respiratory syndrome coronavirus 2.
- WHO:
-
World Health Organization.
References
Tan W, Zhao X, Ma X, Wang W, Niu P, Xu W, Gao GF, Wu G (2020) A novel coronavirus genome identified in a cluster of pneumonia cases - Wuhan, China 2019–2020. China CDC Wkly 2:61–62
Hoffmann M, Kleine-Weber H, Schroeder S, Krüger N, Herrler T, Erichsen S, Schiergens TS, Herrler G, Wu NH, Nitsche A, Müller MA, Drosten C, Pöhlmann S (2020) SARS-CoV-2 cell entry depends on ACE2 and TMPRSS2 and is blocked by a clinically proven protease inhibitor. Cell 181:271–280e278. https://doi.org/10.1016/j.cell.2020.02.052
Lippi G, Lavie CJ, Henry BM, Sanchis-Gomar F (2020) Do genetic polymorphisms in angiotensin converting enzyme 2 (ACE2) gene play a role in coronavirus disease 2019 (COVID-19)? Clin Chem Lab Med 58:1415–1422. https://doi.org/10.1515/cclm-2020-0727
Lu R, Zhao X, Li J, Niu P, Yang B, Wu H, Wang W, Song H, Huang B, Zhu N, Bi Y, Ma X, Zhan F, Wang L, Hu T, Zhou H, Hu Z, Zhou W, Zhao L, Chen J, Meng Y, Wang J, Lin Y, Yuan J, Xie Z, Ma J, Liu WJ, Wang D, Xu W, Holmes EC, Gao GF, Wu G, Chen W, Shi W, Tan W (2020) Genomic characterisation and epidemiology of 2019 novel coronavirus: implications for virus origins and receptor binding. Lancet 395:565–574. https://doi.org/10.1016/s0140-6736(20)30251-8
Yan R, Zhang Y (2020) Structural basis for the recognition of SARS-CoV-2 by full-length human ACE2. Science 367:1444–1448. https://doi.org/10.1126/science.abb2762
Hamming I, Timens W, Bulthuis ML, Lely AT, Navis G, van Goor H (2004) Tissue distribution of ACE2 protein, the functional receptor for SARS coronavirus. A first step in understanding SARS pathogenesis. J Pathol 203:631–637. https://doi.org/10.1002/path.1570
Xu H, Zhong L, Deng J, Peng J, Dan H, Zeng X, Li T, Chen Q (2020) High expression of ACE2 receptor of 2019-nCoV on the epithelial cells of oral mucosa. Int J Oral Sci 12:8. https://doi.org/10.1038/s41368-020-0074-x
Lan J, Ge J, Yu J (2020) Structure of the SARS-CoV-2 spike receptor-binding domain bound to the ACE2 receptor. Nature 581:215–220. https://doi.org/10.1038/s41586-020-2180-5
Senapati S, Banerjee P, Bhagavatula S, Kushwaha PP, Kumar S (2021) Contributions of human ACE2 and TMPRSS2 in determining host-pathogen interaction of COVID-19. J Genet 100. https://doi.org/10.1007/s12041-021-01262-w
Cao Y, Li L, Feng Z, Wan S, Huang P, Sun X, Wen F, Huang X, Ning G, Wang W (2020) Comparative genetic analysis of the novel coronavirus (2019-nCoV/SARS-CoV-2) receptor ACE2 in different populations. Cell Discov 6:11. https://doi.org/10.1038/s41421-020-0147-1
Benetti E, Tita R, Spiga O, Ciolfi A (2020) ACE2 gene variants may underlie interindividual variability and susceptibility to COVID-19 in the italian population. Eur J Hum Genet 28:1602–1614. https://doi.org/10.1038/s41431-020-0691-z
Pung R, Chiew CJ, Young BE, Chin S, Chen MI, Clapham HE, Cook AR, Maurer-Stroh S, Toh M, Poh C, Low M, Lum J, Koh VTJ, Mak TM, Cui L, Lin R, Heng D, Leo YS, Lye DC, Lee VJM (2020) Investigation of three clusters of COVID-19 in Singapore: implications for surveillance and response measures. Lancet 395:1039–1046. https://doi.org/10.1016/s0140-6736(20)30528-6
Napoli PE, Nioi M (2020) Global spread of Coronavirus Disease 2019 and Malaria: an Epidemiological Paradox in the early stage of a pandemic. J Clin Med 9. https://doi.org/10.3390/jcm9041138
Suryamohan K, Diwanji D, Stawiski EW, Gupta R, Miersch S, Liu J, Chen C (2021) Human ACE2 receptor polymorphisms and altered susceptibility to SARS-CoV-2. Commun Biol 4:475. https://doi.org/10.1038/s42003-021-02030-3
Ercegovac M, Asanin M, Savic-Radojevic A (2022) Antioxidant Genetic Profile modifies probability of developing neurological sequelae in Long-COVID. https://doi.org/10.3390/antiox11050954. Antioxidants 11
Khayat AS, de Assumpção PP, Meireles Khayat BC (2020) ACE2 polymorphisms as potential players in COVID-19 outcome. PLoS ONE 15:e0243887. https://doi.org/10.1371/journal.pone.0243887
Möhlendick B, Schönfelder K, Breuckmann K, Elsner C, Babel N, Balfanz P, Dahl E, Dreher M, Fistera D, Herbstreit F, Hölzer B, Koch M, Kohnle M, Marx N, Risse J, Schmidt K, Skrzypczyk S, Sutharsan S, Taube C, Westhoff TH, Jöckel KH, Dittmer U, Siffert W, Kribben A (2021) ACE2 polymorphism and susceptibility for SARS-CoV-2 infection and severity of COVID-19. Pharmacogenet Genomics 31:165–171. https://doi.org/10.1097/fpc.0000000000000436
Novelli A, Biancolella M, Borgiani P, Cocciadiferro D, Colona VL, D’Apice MR, Rogliani P, Zaffina S, Leonardis F, Campana A, Raponi M, Andreoni M, Grelli S, Novelli G (2020) Analysis of ACE2 genetic variants in 131 italian SARS-CoV-2-positive patients. Hum Genomics 14:29. https://doi.org/10.1186/s40246-020-00279-z
Sabater Molina M, Nicolás Rocamora E, Bendicho AI, Vázquez EG, Zorio E, Rodriguez FD, Gil Ortuño C, Rodríguez AI, Sánchez-López AJ, Jara Rubio R, Moreno-Docón A, Marcos PJ, García Pavía P, Villa RB (2022) Polymorphisms in ACE, ACE2, AGTR1 genes and severity of COVID-19 disease. PLoS ONE 17:e0263140. https://doi.org/10.1371/journal.pone.0263140
Shikov AE, Barbitoff YA, Glotov AS, Danilova MM, Tonyan ZN, Nasykhova YA, Mikhailova AA, Bespalova ON, Kalinin RS, Mirzorustamova AM, Kogan IY, Baranov VS, Chernov AN, Pavlovich DM, Azarenko SV, Fedyakov MA, Tsay VV, Eismont YA, Romanova OV, Hobotnikov DN, Vologzhanin DA, Mosenko SV, Ponomareva TA, Talts YA, Anisenkova AU, Lisovets DG, Sarana AM, Urazov SP, Scherbak SG, Glotov OS (2020) Analysis of the Spectrum of ACE2 Variation suggests a possible influence of Rare and Common Variants on susceptibility to COVID-19 and Severity of Outcome. Front Genet 11:551220. https://doi.org/10.3389/fgene.2020.551220
Rashid MU, Muzaffar M, Khan FA, Kabisch M, Muhammad N, Faiz S, Loya A, Hamann U (2015) Association between the BsmI polymorphism in the vitamin D receptor gene and breast Cancer risk: results from a pakistani case-control study. PLoS ONE 10:e0141562. https://doi.org/10.1371/journal.pone.0141562
Erali M, Wittwer CT (2010) High resolution melting analysis for gene scanning. Methods 50:250–261. https://doi.org/10.1016/j.ymeth.2010.01.013
Li J, Shi L, Zhang K, Zhang Y, Hu S, Zhao T, Teng H, Li X, Jiang Y, Ji L, Sun Z (2018) VarCards: an integrated genetic and clinical database for coding variants in the human genome. Nucleic Acids Res 46:D1039–d1048. https://doi.org/10.1093/nar/gkx1039
Al-Mulla F, Mohammad A, Al Madhoun A, Haddad D, Ali H, Eaaswarkhanth M, John SE, Nizam R, Channanath A, Abu-Farha M, Ahmad R, Abubaker J, Thanaraj TA (2021) ACE2 and FURIN variants are potential predictors of SARS-CoV-2 outcome: a time to implement precision medicine against COVID-19. Heliyon 7:e06133. https://doi.org/10.1016/j.heliyon.2021.e06133
Ali F, Elserafy M, Alkordi MH, Amin M (2020) ACE2 coding variants in different populations and their potential impact on SARS-CoV-2 binding affinity. Biochem Biophys Rep 24:100798. https://doi.org/10.1016/j.bbrep.2020.100798
Calcagnile M, Forgez P, Iannelli A, Bucci C, Alifano M, Alifano P (2021) Molecular docking simulation reveals ACE2 polymorphisms that may increase the affinity of ACE2 with the SARS-CoV-2 spike protein. Biochimie 180:143–148. https://doi.org/10.1016/j.biochi.2020.11.004
Lanjanian H, Moazzam-Jazi M (2021) SARS-CoV-2 infection susceptibility influenced by ACE2 genetic polymorphisms: insights from Tehran Cardio-Metabolic Genetic Study. Sci Rep 11:1529. https://doi.org/10.1038/s41598-020-80325-x
Li Q, Guan X, Wu P, Wang X, Zhou L, Tong Y, Ren R, Leung KSM, Lau EHY, Wong JY, Xing X, Xiang N, Wu Y, Li C, Chen Q, Li D, Liu T, Zhao J, Liu M, Tu W, Chen C, Jin L, Yang R, Wang Q, Zhou S, Wang R, Liu H, Luo Y, Liu Y, Shao G, Li H, Tao Z, Yang Y, Deng Z, Liu B, Ma Z, Zhang Y, Shi G, Lam TTY, Wu JT, Gao GF, Cowling BJ (2020) Early Transmission Dynamics in Wuhan, China, of Novel Coronavirus-Infected Pneumonia. N Engl J Med 382:1199–1207. https://doi.org/10.1056/NEJMoa2001316
Torre-Fuentes L, Matías-Guiu J, Hernández-Lorenzo L, Montero-Escribano P, Pytel V, Porta-Etessam J, Gómez-Pinedo U (2021) ACE2, TMPRSS2, and furin variants and SARS-CoV-2 infection in Madrid, Spain. J Med Virol 93:863–869. https://doi.org/10.1002/jmv.26319
Djukic T, Stevanovic G, Coric V (2022) GSTO1, GSTO2 and ACE2 polymorphisms modify susceptibility to developing COVID-19. J Pers Med 12. https://doi.org/10.3390/jpm12030458
Badawi S, Mohamed FE, Alkhofash NR, John A, Ali A, Ali BR (2022) Characterization of ACE2 naturally occurring missense variants: impact on subcellular localization and trafficking. Hum Genomics 16:35. https://doi.org/10.1186/s40246-022-00411-1
Hussain M, Jabeen N, Raza F, Shabbir S, Baig AA, Amanullah A, Aziz B (2020) Structural variations in human ACE2 may influence its binding with SARS-CoV-2 spike protein. J Med Virol 92:1580–1586. https://doi.org/10.1002/jmv.25832
Othman H, Bouslama Z, Brandenburg JT, da Rocha J, Hamdi Y, Ghedira K, Srairi-Abid N, Hazelhurst S (2020) Interaction of the spike protein RBD from SARS-CoV-2 with ACE2: similarity with SARS-CoV, hot-spot analysis and effect of the receptor polymorphism. Biochem Biophys Res Commun 527:702–708. https://doi.org/10.1016/j.bbrc.2020.05.028
Wang J, Xu X, Zhou X, Chen P, Liang H, Li X, Zhong W, Hao P (2020) Molecular simulation of SARS-CoV-2 spike protein binding to pangolin ACE2 or human ACE2 natural variants reveals altered susceptibility to infection. J Gen Virol 101:921–924. https://doi.org/10.1099/jgv.0.001452
Cafiero C, Rosapepe F, Palmirotta R (2021) Angiotensin system polymorphisms’ in SARS-CoV-2 positive patients: Assessment between symptomatic and asymptomatic patients: a pilot study. Pharmgenomics Pers Med 14:621–629. https://doi.org/10.2147/pgpm.s303666
Mir MM, Mir R, Alghamdi MAA, Alsayed BA (2021) Strong Association of Angiotensin converting Enzyme-2 gene Insertion/Deletion polymorphism with susceptibility to SARS-CoV-2, Hypertension, Coronary Artery Disease and COVID-19 Disease Mortality. J Pers Med 11. https://doi.org/10.3390/jpm11111098
Sienko J, Marczak I (2022) Association of ACE2 Gene Variants with the severity of COVID-19 Disease-A prospective observational study. Int J Environ Res Public Health 19. https://doi.org/10.3390/ijerph191912622
Ravikanth V, Sasikala M, Naveen V, Latha SS, Parsa KVL, Vijayasarathy K, Amanchy R, Avanthi S, Govardhan B, Rakesh K, Kumari DS, Srikaran B, Rao GV, Reddy DN (2021) A variant in TMPRSS2 is associated with decreased disease severity in COVID-19. Meta Gene 29:100930. https://doi.org/10.1016/j.mgene.2021.100930
Gómez J, Albaiceta GM, García-Clemente M, López-Larrea C, Amado-Rodríguez L, Lopez-Alonso I, Hermida T, Enriquez AI, Herrero P, Melón S, Alvarez-Argüelles ME, Boga JA, Rojo-Alba S, Cuesta-Llavona E, Alvarez V, Lorca R, Coto E (2020) Angiotensin-converting enzymes (ACE, ACE2) gene variants and COVID-19 outcome. Gene 762:145102. https://doi.org/10.1016/j.gene.2020.145102
Zhou P, Yang XL, Wang XG, Hu B, Zhang L, Zhang W, Si HR, Zhu Y, Li B, Huang CL, Chen HD, Chen J, Luo Y, Guo H, Jiang RD, Liu MQ, Chen Y, Shen XR, Wang X, Zheng XS, Zhao K, Chen QJ, Deng F, Liu LL, Yan B, Zhan FX, Wang YY, Xiao GF, Shi ZL (2020) A pneumonia outbreak associated with a new coronavirus of probable bat origin. Nature 579:270–273. https://doi.org/10.1038/s41586-020-2012-7
Karakaş Çelik S, Çakmak Genç G (2021) Polymorphisms of ACE (I/D) and ACE2 receptor gene (Rs2106809, Rs2285666) are not related to the clinical course of COVID-19: a case study. J Med Virol 93:5947–5952. https://doi.org/10.1002/jmv.27160
Acknowledgements
The authors are grateful to all study subjects for their participation. The authors thank Dr. Faisal Sultan for critical reading of the manuscript.
Funding
This study was supported by the Shaukat Khanum Memorial Cancer Hospital and Research Centre, grant number IRB-20-15-A1. MUR has received this research grant.
Author information
Authors and Affiliations
Contributions
Conceptualization: NM, MUR; Methodology: RS, US, NM; Formal analysis and investigation: NM, MUR, RS, US, KS, UH, AR; Writing – original draft preparation: HN, AA; Writing – review and editing: NM, MUR; Funding acquisition: MUR; Supervision: MUR. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors have no relevant financial or non-financial interests to disclose.
Ethical approval
All procedures performed in studies involving human participants were in accordance with the ethical standards of the institutional and/or national research committee and with the 1964 Helsinki declaration and its later amendments or comparable ethical standards. The study was approved by the Institutional Review Board (IRB) of SKM CH&RC (IRB approval number IRB-20-15-A1, date July 24, 2020).
Consent to participate
Owing to the retrospective study, the IRB granted the waiver of the written informed consent from study participants.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Muhammad, N., Naeemi, H., Azeem, A. et al. Genetic analysis of ACE2 peptidase domain in SARS-CoV-2-positive and SARS-CoV-2-negative individuals from Pakistan. Mol Biol Rep 50, 4309–4316 (2023). https://doi.org/10.1007/s11033-023-08315-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11033-023-08315-6