Abstract
Background
The major activity of β-amylase (BMY) is the production of maltose by the hydrolytic degradation of starch. BMY is found to be produced by some plants and few microorganisms only. The industrial importance of the enzyme warrants its application in a larger scale with the help of genetic engineering, for which the regulatory mechanism is to be clearly understood.
Results and Conclusion
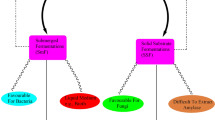
In plants, the activities of BMY are regulated by various environmental stimuli including stress of drought, cold and heat. In vascular plant, Arabidopsis sp. the enzyme is coded by nine BAM genes, whereas in most bacteria, BMY enzymes are coded by the spoII gene family. The activities of these genes are in turn controlled by various compounds. Production and inhibition of the microbial BMY is regulated by the activation and inactivation of various BAM genes. Various types of transcriptional regulators associated with the plant- BMYs regulate the production of BMY enzyme. The enhancement in the expression of such genes reflects evolutionary significance. Bacterial genes, on the other hand, as exemplified by Bacillus sp and Clostridium sp, clearly depict the importance of a single regulatory gene, the absence or mutation of which totally abolishes the BMY activity.



Similar content being viewed by others
References
Gopinath SC, Anbu P, Arshad MK, Lakshmipriya T, Voon CH, Hashim U, Chinni SV (2017) Biotechnological processes in microbial amylase production. Biomed Res Int 2017:1272193
Ray RR, Nanda G (1996) Microbial β-amylases: biosynthesis, characteristics, and industrial applications. Crit Rev Microbiol 22(3):181–199
Raveendran S, Parameswaran B, Ummalyma SB, Abraham A, Mathew AK, Madhavan A, Rebello S, Pandey A (2018) Applications of microbial enzymes in food industry. Food Technol Biotechnol 56(1):16–30
Vajravijayan S, Pletnev S, Mani N, Pletneva N, Nandhagopal N, Gunasekaran K (2018) Structural insights on starch hydrolysis by plant β-amylase and its evolutionary relationship with bacterial enzymes. Int J Biol Macromol 113:329–337
Robinson PK (2015) Enzymes: principles and biotechnological applications. Essays Biochem 59:1–41
Kaplan F, Guy CL (2004) beta-Amylase induction and the protective role of maltose during temperature shock. Plant Physiol 135(3):1674–1684
Kossmann J, Lloyd J (2000) Understanding and influencing starch biochemistry. Crit Rev Biochem Mol Biol 35:141–196
Das R, Kayastha AM (2019) β-amylase: general properties, mechanism and panorama of applications by immobilization on nano-structures. In: Husain Q, Ullah M (eds) Biocatalysis. Springer, Cham
Husain Q, Ullah MF (eds) (2019) Biocatalysis. Springer, Cham. https://doi.org/10.1007/978-3-030-25023-2
Uozumi N, Matsuda T, Tsukagoshi N, Udaka S (1991) Structural and functional roles of cysteine residues of Bacillus polymyxa beta-amylase. Biochemistry 30(18):4594–4599
Kaplan F, Guy CL (2005) RNA interference of Arabidopsis beta-amylase prevents maltose accumulation upon cold shock and increases sensitivity of PSII photochemical efficiency to freezing stress. Plant J 44(5):730–743
Swanson MA (1948) Studies on the structure of polysaccharides IV. Relation of the iodine color to the structure. J Biol Chem 172(2):825–837
Cleveland F, Kerr R (1948) The action of beta-amylase on corn amylose. Cereal Chem 25(2):133–139
French D, Levine ML, Pazur J, Norberg E (1950) Studies on the schardinger dextrins. IV. The action of soy bean beta amylase on amyloheptaose. J Am Chem Soc 72(4):1746–1748
Thoma JA, Koshland DE (1960) Stereochemistry of enzyme, substrate, and products during beta-amylase action. J Biol Chem 235:2511–2517
Smith SM, Fulton DC, Chia T, Thorneycroft D, Chapple A, Dunstan H, Hylton C, Zeeman SC, Smith AM (2004) Diurnal changes in the transcriptome encoding enzymes of starch metabolism provide evidence for both transcriptional and post-transcriptional regulation of starch metabolism in Arabidopsis leaves. Plant Physiol 136(1):2687–2699
Eglinton JK, Langridge P, Evans DE (1998) Thermostability variation in alleles of barley betaamylase. J Cereal Sci 28(3):301–309
Fulton DC, Stettler M, Mettler T, Vaughan CK, Li J, Francisco P (2008) Beta-AMYLASE4, a noncatalytic protein required for starch breakdown, acts upstream of three active beta-amylases in Arabidopsis chloroplasts. Plant Cell 20:1040–1058
Sparla F, Costa A, Lo Schiavo F, Pupillo P, Trost P (2006) Redox regulation of a novel plastid-targeted β-amylase. Plant Physiol 141:840–850
Li J, Zhou W, Francisco P, Wong R, Zhang D, Smith SM (2017) Inhibition of Arabidopsis chloroplast β-amylase BAM3 by maltotriose suggests a mechanism for the control of transitory leaf starch mobilisation. PLoS ONE 12:e0172504
Saika H, Nakazono M, Ikeda A, Yamaguchi J, Masaki S (2005) Kanekatsu MA transposon-induced spontaneous mutation results in low β-amylase content in rice. Plant Sci 169:239–244
Hirano T, Takahashi Y, Fukayama H, Michiyama H (2011) Identification of two plastid-targeted β-amylases in rice. Plant Prod Sci 14:318–324
Laby RJ, Kim D, Gibson SI (2001) The ram1 mutant of Arabidopsis exhibits severely decreased βamylase activity. Plant Physiol 127(4):1798–1807
Lee JS, Wittchen KD, Stahl C, Strey J, Meinhardt F (2001) Cloning, expression, and carbon catabolite repression of the bamM gene encoding β-amylase of Bacillus megaterium DSM319. Appl Microbiol Biotechnol 56:205–211
Amore A, Giacobbe S, Faraco V (2013) Regulation of cellulase and hemicellulase gene expression in fungi. Curr Genomics 14(4):230–249
Ray RR, Jana SC, Nanda G (1996) Induction and carbon catabolite repression in the biosynthesis of beta-amylase by Bacillus megaterium B6. Biochem Mol Biol Int 38(2):223–230
Hyun HH, Zeikus JG (1985) General biochemical characterization of thermostable extracellular P-amylase of Clostridium thermosulfirogenes. Appl Environ Microbiol 49:1162
Chatterjee BS, Ghosh A, Das A (1992) Starch digestion and adsorption by P-amylase of Emericella nidulans (Aspergillus nidulans). J Appl Bacteriol 12:208
Mizukami M, Yamagata H, Sakaguchi K, Udaka S (1992) Efficient production of thermostable Clostridium thermosulfurogenes P-amylase by Bacillus brevis. J Ferment Bioeng 73:112
Nanmori T, Numata Y, Shinke R (1987) Isolation and characterization of a Bacillus cereus mutant strain hyperproductive of exo p-amylase. Appl Environ Microbiol 53:768
Koaze Y, Nakajima Y, Hidaka H, Niwa T, Yoshida K, Ito J, Niida T, Shomura T, Ueda M (1975) U.S. Patent 3,368-464
Glen PF, Fox LW (2020) Barley: current understanding of ‘omics data on quality. In: Beddows C (ed) Reference module in food science. Elsevier, Amsterdam
Gimbi DM, Kitabatake N (2002) Changes in alpha-and beta-amylase activities during seed germination of African finger millet. Int J Food Sci Nutr 53(6):481–488. https://doi.org/10.1080/09637480220164361
Mita S, Suzuki-Fujii K, Nakamura K (1995) Sugar-inducible expression of a gene for beta-amylase in Arabidopsis thaliana. Plant Physiol 107(3):895–904. https://doi.org/10.1104/pp.107.3.895
Erkkilä MJ, Leah R, Ahokas H, Cameron-Mills V (1998) Allele-dependent barley grain beta-amylase activity. Plant Physiol 117(2):679–685
Yue C, Cao H, Lin H et al (2019) Expression patterns of alpha-amylase and beta-amylase genes provide insights into the molecular mechanisms underlying the responses of tea plants (Camellia sinensis) to stress and postharvest processing treatments. Planta 250:281–298
Görke B, Stülke J (2008) Carbon catabolite repression in bacteria: many ways to make the most out of nutrients. Nat Rev Microbiol 6(8):613–624
Hueck CJ, Hillen W (1995) Catabolite repression in Bacillus subtilis: a global regulatory mechanism for the gram-positive bacteria? Mol Microbiol 15(3):395–401
Wittchen KD, Meinhardt F (1995) Inactivation of the major extracellular protease from Bacillus megaterium DSM319 by gene replacement. Appl Microbiol Biotechnol 42:871–877
Maeo K, Tomiya T, Hayashi K, Akaike M, Morikami A, Ishiguro S, Nakamura K (2001) Sugar-responsible elements in the promoter of a gene for beta-amylase of sweet potato. Plant Mol Biol 46(5):627–637
Zhang H, Liu J, Hou J, Yao Y, Lin Y, Ou Y, Song B, Xie C (2014) The potato amylase inhibitor gene SbAI regulates cold-induced sweetening in potato tubers by modulating amylase activity. Plant Biotechnol J 12(7):984–993
Henkin TM, Grundy FJ, Nicholson WL, Chambliss GH (1991) Catabolite repression of α amylase gene expression in Bacillus subtilis involves a trans-acting gene product homologous to the Escherichia coli lacl and galR repressors. Mol Microbiol 5:575–584. https://doi.org/10.1111/j.1365-2958.1991.tb00728.x
Fujita Y, Miwa Y, Galinier A, Deutscher J (1995) Specific recognition of the Bacillus subtilis gnt cis-acting catabolite-responsive element by a protein complex formed between CcpA and seryl-phosphorylated HPr. Mol Microbial 17:953–960
Kinoshita T, Caño-Delgado A, Seto H, Hiranuma S, Fujioka S, Yoshida S, Chory J (2005) Binding of brassinosteroids to the extracellular domain of plant receptor kinase BRI1. Nature 433:167–171
Reinhold H, Soyk S, Šimková K, Hostettler C, Marafino J, Mainiero S, Vaughan CK, Monroe JD, Zeeman SC (2011) b-amylase–like proteins function as transcription factors in Arabidopsis, controlling shoot growth and development. Plant Cell 23:1391–1403
Özcan D, SİpahİoĞlu HM (2020) Simultaneous production of alpha and beta amylase enzymes using separate gene bearing recombinant vectors in the same Escherichia coli cells. Turk J Biol Turk biyoloji dergisi 44(4):201–207. https://doi.org/10.3906/biy-2001-71
Ali M, Ishqi HM, Husain Q (2020) Enzyme engineering: reshaping the biocatalytic functions. Biotechnol Bioeng. https://doi.org/10.1002/bit.27329
Nipkow A, Shen GJ, Zeikus JG (1989) Continuous production of thermostable P-amylase with Clostridium thermosulfurogenes: effect of culture conditions and metabolite levels on enzyme synthesis and activity. Appl Environ Microbiol 55:689
Huang CJ, Chen YL, Hsu WH (1988) Thermostable P-amylase production by Bacillus circulans H-107. J Chin Agric Chem SOC 26:457
Sandhu DK, Vilkhu KS, Soni SK (1987) Production of a-amylase by Saccharomyces jibulig era (syn. Endomycopsis jibulig era). J Ferment Technol 65:387
Duan X, Shen Z, Zhang X, Wang Y, Huang Y (2019) Production of recombinant beta-amylase of Bacillus aryabhattai. Prep Biochem Biotechnol 49:88
Monroe JD, Storm AR (2018) Review: the Arabidopsis β-amylase (BAM) gene family: diversity of form and function. Plant Sci 276:163–170
Author information
Authors and Affiliations
Contributions
All the authors contributed equally in drafting the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors don’t have any conflict of interest.
Ethical approval
Not Applicable.
Consent to participate
Not Applicable.
Consent to publish
All the authors have consent to publish.
Informed consent
All the authors have the consent.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Nag, M., Lahiri, D., Garai, S. et al. Regulation of β-amylase synthesis: a brief overview. Mol Biol Rep 48, 6503–6511 (2021). https://doi.org/10.1007/s11033-021-06613-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11033-021-06613-5