Abstract
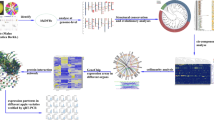
With complete sequencing of the ‘Golden Delicious’ genome, the transcriptome, proteome and metabolome have been fused with the genome. However, to date, no studies have evaluated the importance of systematic functions based on systematic biology in apples. In this study, we summarized the history of genetic maps of apples, and introduced the functions of gene families from the perspective of genomics; the functions included resisting abiotic and biotic stress, hormone response, fruit development and quality, epitomizing the development of other omics and relationships between functions. Furthermore, we established a model for gene structure and functional analyses, as well as of the protein interaction network, for apple. These results provide detailed systematic information for biological basis studies in apple as well as apple breeding, laying a good foundation for systematic biology.


Similar content being viewed by others
Abbreviations
- AFLP:
-
Amplified fragment length polymorphism
- RAPD:
-
Random amplified polymorphic DNA
- SSR:
-
Simple sequence repeat
- SCAR:
-
Sequence characterized amplified regions
- QTL:
-
Quantitative trait locus
- SNP:
-
Single nucleotide polymorphism
- CAPS:
-
Cleaved amplified polymorphism sequences
- SSCP:
-
Single-strand conformation polymorphism
- PDS:
-
Phytoene desaturase
- IAA:
-
Indole-3-acetic acid
- GA:
-
Gibberellin
- JA:
-
Jasmonic acid
- Ea :
-
Erwinia amylovora
- LG:
-
Linkage group
- GDR:
-
Genome database for Rosaceae
- PPI:
-
Protein–protein Interactions
- GO:
-
Gene ontology
- 1-MCP:
-
1-methylcyclopropene
References
Antanaviciute L, Fernández-Fernández F, Jansen J, Banchi E, Evans KM, Viola R, Velasco R, Dunwell JM, Troggio M, Sargent DJ (2012) Development of a dense SNP-based linkage map of an apple rootstock progeny using the Malus Infinium whole genome genotyping array. BMC Genom 13:203
Bai Y, Dougherty L, Xu K (2014) Towards an improved apple reference transcriptome using RNA-sEq. Mol Genet Genom 289:427–438
Barad S, Sela N, Kumar D, Kumar-Dubey A, Glam-Matana N, Sherman A, Prusky D (2016) Fungal and host transcriptome analysis of pH-regulated genes during colonization of apple fruits by Penicillium expansum. BMC Genom 17:1
Bianco L, Cestaro A, Sargent DJ, Banchi E, Derdak S, Di Guardo M, Salvi S, Jansen J, Viola R, Gut I (2014) Development and validation of a 20 K single nucleotide polymorphism (SNP) whole genome genotyping array for apple (Malus × domestica Borkh). PLoS ONE 9:e110377
Bocsanczy A, Phillips J, Korban S, Dardick C, Bassett C (2011) Apple (Malus x domestica) transcriptome in response to the compatible pathogen Erwinia amylovora and the incompatible pathogen pseudomonas syringae. Acta Hortic 2011:237–241
Brendel V (2007) Gene Structure Annotation at PlantGDB. Methods Mol Biol 406:521–533
Buron-Moles G, Wisniewski M, Viñas I, Teixidó N, Usall J, Droby S, Torres R (2015) Characterizing the proteome and oxi-proteome of apple in response to a host (Penicillium expansum) and a non-host (Penicillium digitatum) pathogen. J Proteomics 114:136–151
Bus VG, Bassett HC, Bowatte D, Chagné D, Ranatunga CA, Ulluwishewa D, Wiedow C, Gardiner SE (2010) Genome mapping of an apple scab, a powdery mildew and a woolly apple aphid resistance gene from open-pollinated mildew immune selection. Tree Genet Genomes 6:477–487
Buts K, Carpentier S, Hertog M, Cordewener J, America T, Nicolai B (2011) A systems biology approach to browning in apple. Influence of Harvest Date on the Proteome
Buts K, Hertog ML, Ho QT, America AH, Cordewener J, Vercammen J, Carpentier SC, Nicolai B (2016) Influence of pre-harvest calcium, potassium and triazole application on the proteome of apple at harvest. J Sci Food Agric. doi: 10.1002/jsfa.7664
Cao Z-H, Zhang S-Z, Wang R-K, Zhang R-F, Hao Y-J (2013) Genome wide analysis of the apple MYB transcription factor family allows the identification of MdoMYB121 gene confering abiotic stress tolerance in plants. PLoS ONE 8:e69955
Chagné D, Crowhurst RN, Troggio M, Davey MW, Gilmore B, Lawley C, Vanderzande S, Hellens RP, Kumar S, Cestaro A (2012) Genome-wide SNP detection, validation, and development of an 8 K SNP array for apple. PLoS ONE 7:e31745
Chikwambi Z (2013) A transcriptome analysis of apple (Malus x domestica Borkh.) cv ‘Golden Delicious’ fruit during fruit growth and development. University of the Western Cape
Choi C (2013) Development of novel ssr (simple sequence repeat) markers and construction of genetic map in apple for Korean Apple genome project. In plant and animal genome XXI conference. Plant and Animal Genome
Chua HN, Sung WK, Wong l (2010) Exploiting indirect neighbours and topological weight to predict protein function from protein–protein interactions. Bioinformatics 22:452–452
Conesa A, Gotz S (2008) Blast2GO: A comprehensive suite for functional analysis in plant genomics. Int J Plant Genomics 2008: 619832
Conner PJ, Brown SK, Weeden NF (1997) Randomly amplified polymorphic DNA-based genetic linkage maps of three apple cultivars. J Am Soc Hortic Sci 122:350–359
Cornille A, Gladieux P, Smulders MJ, Roldán-Ruiz I, Laurens F, Le Cam B, Nersesyan A, Clavel J, Olonova M, Feugey L (2012) New insight into the history of domesticated apple: secondary contribution of the European wild apple to the genome of cultivated varieties. PLoS Genet 8:e1002703
Cuthbertson D, Andrews PK, Reganold JP, Davies NM, Lange BM (2012) Utility of metabolomics toward assessing the metabolic basis of quality traits in apple fruit with an emphasis on antioxidants. J Agric Food Chem 60:8552–8560
Ding YD, Chang JW, Guo J, Chen D, Li S, Xu Q, Deng XX, Cheng YJ, Chen LL (2014) Prediction and functional analysis of the sweet orange protein-protein interaction network. BMC Plant Biol 14:213
Ehlers M, Mack CI, Weinert CH, Egert B, Trierweiler B, Baab G, Kulling SE (2016) Comparison of conventional and new apple cultivars using GC-based metabolomics. Berichte aus dem Julius Kühn-Institut 183: 83–84
El-Sharkawy I, Liang D, Xu K (2015) Transcriptome analysis of an apple (Malus × domestica) yellow fruit somatic mutation identifies a gene network module highly associated with anthocyanin and epigenetic regulation. J Exp Bot 66:7359–7376
Evans K, Patocchi A, Rezzonico F, Mathis F, Durel C, Fernández-Fernández F, Boudichevskaia A, Dunemann F, Stankiewicz-Kosyl M, Gianfranceschi L (2011) Genotyping of pedigreed apple breeding material with a genome-covering set of SSRs: trueness-to-type of cultivars and their parentages. Mole Breed 28:535–547
Farneti B, Busatto N, Khomenko I, Cappellin L, Gutierrez S, Spinelli F, Costa F (2015) Untargeted metabolomics investigation of volatile compounds involved in the development of apple superficial scald by PTR-ToF-MS. Metabolomics 11:341–349
Fernández-Fernández F, Antanaviciute L, van Dyk M, Tobutt K, Evans K, Rees D, Dunwell J, Sargent D (2012) A genetic linkage map of an apple rootstock progeny anchored to the Malus genome sequence. Tree Genet Genomes 8:991–1002
Gabaldón T, Koonin EV (2013) Functional and evolutionary implications of gene orthology. Nat Rev Genet 14:360–366
Gardner KM (2013) Genome-wide survey of genetic diversity in apple using genotyping-by-sequencing. In plant and animal genome XXI Conference. Plant and Animal Genome
Gianfranceschi L, Seglias N, Tarchini R, Komjanc M, Gessler C (1998) Simple sequence repeats for the genetic analysis of apple. Theor Appl Genet 96:1069–1076
Giovannoni J (2010) Harvesting the apple genome. Nat Genet 42:822
Girardi CL, Rombaldi CV, Dal Cero J, Nobile PM, Laurens F, Bouzayen M, Quecini V (2013) Genome-wide analysis of the AP2/ERF superfamily in apple and transcriptional evidence of ERF involvement in scab pathogenesis. Sci Hortic 151:112–121
Goodstein DM, Shu S, Howson R, Neupane R, Hayes RD, Fazo J, Mitros T, Dirks W, Hellsten U, Putnam N, Rokhsar DS (2012) Phytozome: a comparative platform for green plant genomics. Nucleic Acids Res 40:D1178–D1186
Gutierrez RA, Shasha DE, Coruzzi GM (2005) Systems biology for the virtual plant. Plant Physiol 138:550–554
Han D-S, Kim H-S, Jang W-H, Lee S-D, Suh J-K (2004) PreSPI: a domain combination based prediction system for protein–protein interaction. Nucleic Acids Res 32:6312–6320
Han Y, Zheng D, Vimolmangkang S, Khan MA, Beever JE, Korban SS (2011) Integration of physical and genetic maps in apple confirms whole-genome and segmental duplications in the apple genome. J Exp Bot 62:5117–5130
Hatoum D, Hertog ML, Geeraerd AH, Nicolai BM (2016) Effect of browning related pre-and postharvest factors on the ‘Braeburn’apple metabolome during CA storage. Postharvest Biol Technol 111:106–116
Hemmat M, Weedon N, Manganaris A, Lawson D (1994) Molecular marker linkage map for apple. J Hered 85:4–11
Igarashi M, Abe Y, Hatsuyama Y, Ueda T, Fukasawa-Akada T, Kon T, Kudo T, Sato T, Suzuki M (2008) Linkage maps of the apple (Malus × domestica Borkh.) cvs ‘Ralls Janet’and ‘Delicious’ include newly developed EST markers. Mol Breed 22:95–118
Ireland HS, Guillen F, Bowen J, Tacken EJ, Putterill J, Schaffer RJ, Johnston JW (2012) Mining the apple genome reveals a family of nine ethylene receptor genes. Postharvest Biol Technol 72:42–46
Ji X-H, Zhang R, Wang N, Yang L, Chen X-S (2015) Transcriptome profiling reveals auxin suppressed anthocyanin biosynthesis in red-fleshed apple callus (Malus sieversii f. niedzwetzkyana). Plant Cell Tissue Organ Culture (PCTOC 123:389–404
Johnson FT, Zhu Y (2015) Transcriptome changes in apple peel tissues during CO2 injury symptom development under controlled atmosphere storage regimens. Horticu Res 2:15061
Jung S, Staton M, Lee T, Blenda A, Svancara R, Abbott A, Main D (2008a) GDR (genome database for Rosaceae): integrated web-database for Rosaceae genomics and genetics data. Nucleic Acids Res 36:D1034–D1040
Jung S, Staton M, Lee T, Blenda A, Svancara R, Abbott A, Main D (2008b) GDR (genome database for Rosaceae): integrated web-database for Rosaceae genomics and genetics data. Nucleic Acids Res 36:D1034–D1040
Ke X, Yin Z, Song N, Dai Q, Voegele RT, Liu Y, Wang H, Gao X, Kang Z, Huang L (2014) Transcriptome profiling to identify genes involved in pathogenicity of Valsa mali on apple tree. Fungal Genet Biol 68:31–38
Keller-Przybyłkowicz S, Korbin MU (2013) The history of mapping the apple genome. Folia Hortic 25:161–168.
Kenis K, Keulemans J (2005) Genetic linkage maps of two apple cultivars (Malus × domestica Borkh.) based on AFLP and microsatellite markers. Mol Breed 15:205–219
Khan MA, Han Y, Zhao YF, Troggio M, Korban SS (2012) A multi-population consensus genetic map reveals inconsistent marker order among maps likely attributed to structural variations in the apple genome. PLoS ONE 7:e47864
Kitano H (2002) Systems biology: a brief overview. Science 295:1662–1664
Knight H, Trewavas AJ, Knight MR (1997) Calcium signalling in Arabidopsis thaliana responding to drought and salinity. Plant J 12:1067–1078
Kovtun Y, Chiu W-L, Tena G, Sheen J (2000) Functional analysis of oxidative stress-activated mitogen-activated protein kinase cascade in plants. Proc Nat Acad Sci USA 97:2940–2945
Krost C, Petersen R, Schmidt E, Braun P (2012) Deep sequencing of the shoot apical meristem transcriptome of columnar apple trees (Malus × domestica). In II International Symposium of Biotechnology Fruit Species 1048:103–106
Kumar S, Molloy C, Muñoz P, Daetwyler H, Chagné D, Volz R (2015a) Genome-enabled estimates of additive and nonadditive genetic variances and Prediction of apple phenotypes across environments. G3: Genes| Genomes|. Genet 5:2711–2718
Kumar S, Rowan D, Hunt M, Chagné D, Whitworth C, Souleyre E (2015b) Genome-wide scans reveal genetic architecture of apple flavour volatiles. Mol Breed 35:1–16
Kunihisa M, Moriya S, Abe K, Okada K, Haji T, Hayashi T, Kanamori H (2016) Genomic dissection of a ‘Fuji’ apple cultivar: re-sequencing, SNP marker development, definition of haplotypes, and QTL detection. Breed Sci 66(4):499–515
Lashbrooke J, Aharoni A, Costa F (2015) Genome investigation suggests MdSHN3, an APETALA2-domain transcription factor gene, to be a positive regulator of apple fruit cuticle formation and an inhibitor of russet development. J Exp Bot 66:6579–6589
Lee J, Rudell DR, Davies PJ, Watkins CB (2012) Metabolic changes in 1-methylcyclopropene (1-MCP)-treated ‘Empire’apple fruit during storage. Metabolomics 8:742–753
Li Y, Wu B, Yu Y, Yang G, Wu C, Zheng C (2011) Genome-wide analysis of the RING finger gene family in apple. Mol Genet Genom 286:81–94
Li J, Hou H, Li X, Xiang J, Yin X, Gao H, Zheng Y, Bassett CL, Wang X (2013a) Genome-wide identification and analysis of the SBP-box family genes in apple (Malus × domestica Borkh.). Plant Physiol Biochem 70:100–114
Li Q, Liu J, Tan D, Allan AC, Jiang Y, Xu X, Han Z, Kong J (2013b) A genome-wide expression profile of salt-responsive genes in the apple rootstock Malus zumi. Int J Mol Sci 14:21053–21070
Li T, Tan D, Yang X, Wang A (2013c) Exploring the apple genome reveals six ACC synthase genes expressed during fruit ripening. Sci Hortic 157:119–123
Li X, Guo R, Li J, Singer SD, Zhang Y, Yin X, Zheng Y, Fan C, Wang X (2013d) Genome-wide identification and analysis of the aldehyde dehydrogenase (ALDH) gene superfamily in apple (Malus × domestica Borkh.). Plant Physiol Biochem 71:268–282
Li X, Yin X, Wang H, Li J, Guo C, Gao H, Zheng Y, Fan C, Wang X (2015) Genome-wide identification and analysis of the apple (Malus × domestica Borkh.) TIFY gene family. Tree Genet Genomes 11:1–13
Li G, Ma J, Tan M, Mao J, An N, Sha G, Zhang D, Zhao C, Han M (2016) Transcriptome analysis reveals the effects of sugar metabolism and auxin and cytokinin signaling pathways on root growth and development of grafted apple. BMC Genom 17:1
Liebhard R, Gianfranceschi L, Koller B, Ryder C, Tarchini R, Van de Weg E, Gessler C (2002) Development and characterisation of 140 new microsatellites in apple (Malus x domestica Borkh.). Mol Breed 10:217–241
Liebhard R, Kellerhals M, Pfammatter W, Jertmini M, Gessler C (2003a) Mapping quantitative physiological traits in apple (Malus × domestica Borkh.). Plant Mol Biol 52:511–526
Liebhard R, Koller B, Gianfranceschi L, Gessler C (2003b) Creating a saturated reference map for the apple (Malus × domestica Borkh.) genome. Theor Appl Genet 106:1497–1508
Los DA, Murata N (2004) Membrane fluidity and its roles in the perception of environmental signals. Biochimica et Biophysica Acta (BBA)-Biomembranes 1666:142–157
Lozano L, Iglesias I, Micheletti D, Troggio M, Kumar S, Volz RK, Allan AC, Chagné D, Gardiner SE (2014) Feasibility of genome-wide association analysis using a small single nucleotide polymorphism panel in an apple breeding population segregating for fruit skin color. J Am Soc Hortic Sci 139:619–626
Luo X-C, Sun M-H, Xu R-R, Shu H-R, Wang J-W, Zhang S-Z (2014) Genomewide identification and expression analysis of the ARF gene family in apple. J Genet 93:785–797
Lv J, Rao J, Johnson F, Shin S, Zhu Y (2015) Genome-wide identification of jasmonate biosynthetic genes and characterization of their expression profiles during apple (Malus × domestica) fruit maturation. Plant Growth Regul 75:355–364
Mahajan S, Pandey GK, Tuteja N (2008) Calcium-and salt-stress signaling in plants: shedding light on SOS pathway. Arch Biochem Biophys 471:146–158
Maliepaard C, Alston F, Van Arkel G, Brown L, Chevreau E, Dunemann F, Evans K, Gardiner S, Guilford P, Van Heusden A (1998) Aligning male and female linkage maps of apple (Malus pumila Mill.) using multi-allelic markers. Theor Appl Genet 97:60–73
Meng D, Li Y, Bai Y, Li M, Cheng L (2016) Genome-wide identification and characterization of WRKY transcriptional factor family in apple and analysis of their responses to waterlogging and drought stress. Plant Physiol Biochem 103:71–83
Migicovsky Z, Gardner KM, Money D, Sawler J, Bloom JS, Moffett P, Chao CT, Schwaninger H, Fazio G, Zhong G-Y (2016) Genome to phenome mapping in apple using historical data. Plant Genome. doi: 10.3835/plantgenome2015
Moradi F, Ismail AM (2007) Responses of photosynthesis, chlorophyll fluorescence and ROS-scavenging systems to salt stress during seedling and reproductive stages in rice. Ann Bot (Lond) 99:1161–1173
N’diaye A, Van de Weg W, Kodde L, Koller B, Dunemann F, Thiermann M, Tartarini S, Gennari F, Durel C (2008) Construction of an integrated consensus map of the apple genome based on four mapping populations. Tree Genet Genomes 4:727–743
Ning L, Sun M-h, Jiang Z-s, Shu H-r, Zhang S-z (2016) Genome-wide analysis of the synonymous codon usage patterns in apple. J Integr Agric 15:983–991
Nishitani C, Hirai N, Komori S, Wada M, Okada K, Osakabe K, Osakabe Y (2016) Efficient genome editing in apple using a CRISPR/Cas9 system. Scientific reports 6
Noormohammadi Z, Fazeli S, Sheidai M, Farahani F (2015) Molecular and genome size analyses of somaclonal variation in apple rootstocks Malling 7 and Malling 9. Acta Biol Szegediensis 59: 139–149
Östlund G, Schmitt T, Forslund K, Köstler T, Messina DN, Roopra S, Frings O, Sonnhammer EL (2010) InParanoid 7: new algorithms and tools for eukaryotic orthology analysis. Nucleic Acids Res 38:D196–D203
Pashaiasl M, Ebrahimi M, Ebrahimie E (2016) Identification of the key regulating genes of diminished ovarian reserve (DOR) by network and gene ontology analysis. Mol Biol Rep 43:923–937
Proost S, Van Bel M, Vaneechoutte D, Van de Peer Y, Inze D, Mueller-Roeber B, Vandepoele K (2015) PLAZA 3.0: an access point for plant comparative genomics. Nucleic Acids Res 43:D974–D981
Rudell DR, Mattheis JP, Curry EA (2008) Prestorage ultraviolet–white light irradiation alters apple peel metabolome. J Agric Food Chem 56:1138–1147
Rudell D, Mattheis J, Hertog M (2009) Apple storage management using postharvest metabolomics. Australasian Postharvest Conference
Sarowar S, Wang D, Zhao Y, Guerra R, Zheng D, Korban S (2010) Transcriptome analysis of apple blossom after challenging with fire blight pathogen Erwinia amylovora wild type and mutant strains. In XII International Workshop on Fire Blight 896: 245–251
Shannon PA, Markiel O, Ozier NS, Baliga JT, Wang D, Ramage N, Amin B, Schwikowski, Ideker T (2003) Cytoscape: a software environment for integrated models of biomolecular interaction networks. Genome Res 13:2498–2504
Shin S, Zheng P, Fazio G, Mazzola M, Main D, Zhu Y (2016) Transcriptome changes specifically associated with apple (Malus domestica) root defense response during Pythium ultimum infection. Physiol Mol Plant Pathol 94:16–26
Sikorskaite S, Haimi P, Rajamaki M, Valkonen J, Gelvonauskiene D, Staniene G, Stanys V, Baniulis D (2012) Proteome analysis of cell nuclei enriched subcellular fraction of apple (Malus × domestica Borkh.). FEBS J 279:69–69
Silfverberg-Dilworth E, Matasci C, Van de Weg W, Van Kaauwen M, Walser M, Kodde L, Soglio V, Gianfranceschi L, Durel C, Costa F (2006) Microsatellite markers spanning the apple (Malus x domestica Borkh.) genome. Tree Genet Genomes 2:202–224
Su H, Zhang S, Yuan X, Chen C, Wang X-F, Hao Y-J (2013) Genome-wide analysis and identification of stress-responsive genes of the NAM-ATAF1, 2-CUC2 transcription factor family in apple. Plant Physiol Biochem 71:11–21
Sun J, Zhang G, Li S, Ivanov AR, Fenyo D, Lisacek F, Murthy SK, Karger BL, Brusic V (2014) Pathway analysis and transcriptomics improve protein identification by shotgun proteomics from samples comprising small number of cells—a benchmarking study. BMC Genom 15:S1
Tadiello A (2015) Genome wide transcriptome investigation of fruit development and ripening in apple. In plant and animal genome XXIII Conference. Plant and Animal Genome
Tian Y, Dong Q, Ji Z, Chi F, Cong P, Zhou Z (2015) Genome-wide identification and analysis of the MADS-box gene family in apple. Gene 555:277–290
Troggio M, Zharkikh A, Pindo M, Baldi P, Costa F, Chagné D, Micheletti D, Coppola G, Cestaro A, Lasserre P (2010) 1 PAG-XVIII (P470) a sequence-anchored integrated genetic linkage map for apple, malus X
Troggio M, Chagne D, Cestaro A, Kumar S, Crowhurst RN (2012) Apple, from genome to breeding. Tree Genet Genomes 8:509–529
Vimolmangkang S, Zheng D, Han Y, Khan MA, Soria-Guerra RE, Korban SS (2014) Transcriptome analysis of the exocarp of apple fruit identifies light-induced genes involved in red color pigmentation. Gene 534:78–87
Wang N, Yue Z, Liang D, Ma F (2014) Genome-wide identification of members in the YTH domain-containing RNA-binding protein family in apple and expression analysis of their responsiveness to senescence and abiotic stresses. Gene 538:292–305
Willekens H, Chamnongpol S, Davey M, Schraudner M, Langebartels C, Van Montagu M, Inzé D, Van Camp W (1997) Catalase is a sink for H2O2 and is indispensable for stress defence in C3 plants. EMBO J 16:4806–4816
Xing L, Zhang D, Song X, Weng K, Shen Y, Li Y, Zhao C, Ma J, An N, Han M (2016) Genome-wide sequence variation identification and floral-associated trait comparisons based on the re-sequencing of the ‘nagafu No. 2’and ‘Qinguan’varieties of apple (Malus domestica Borkh.). Front Plant Sci 7:908
Xu J, Xing S, Cui H, Chen X, Wang X (2016) Genome-wide identification and characterization of the apple (Malus domestica) HECT ubiquitin-protein ligase family and expression analysis of their responsiveness to abiotic stresses. Mol Genet Genom 291:635–646
Yuan H, Zhao K, Lei H, Shen X, Liu Y, Liao X, Li T (2013) Genome-wide analysis of the GH3 family in apple (Malus × domestica). BMC Genom 14:1
Zhang Q, Ma B, Li H, Chang Y, Han Y, Li J, Wei G, Zhao S, Khan MA, Zhou Y (2012) Identification, characterization, and utilization of genome-wide simple sequence repeats to identify a QTL for acidity in apple. BMC Genom 13:1
Zhang S, Chen GH, Liu Y, Chen H, Yang G, Yuan X, Jiang Z, Shu H (2013) Apple gene function and gene family database: an integrated bioinformatics database for apple research. Plant Growth Regul 70:199–206
Zhang J, Karmakar S, Yu M, Mitter N, Zou J, Yu C (2014) Protein therapy: synthesis of silica vesicles with controlled entrance size for high loading, sustained release, and cellular delivery of therapeutical proteins (small 24/2014). Small 10:4986
Zhang C-x, Tian Y, Cong P-h (2015) Proteome analysis of pathogen-responsive proteins from apple leaves induced by the alternaria blotch Alternaria alternata. PLoS ONE 10:e0122233
Zhu J-K (2003) Regulation of ion homeostasis under salt stress. Curr Opin Plant Biol 6:441–445
Acknowledgements
This work was supported by the National Natural Science Foundation in China (Grant No. 31401822) and International Science & Technology Cooperation Program of China (2015DFA31190).
Author information
Authors and Affiliations
Corresponding authors
Additional information
Lin Liu and Xiao-cu Luo have contributed equally to the work.
Rights and permissions
About this article
Cite this article
Liu, L., Luo, Xc., Ge, Hj. et al. Apple, from omics to systemic function. Plant Growth Regul 83, 1–11 (2017). https://doi.org/10.1007/s10725-017-0276-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10725-017-0276-1