Abstract
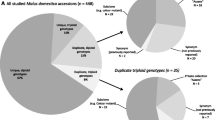
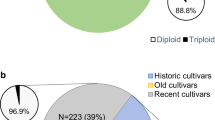
Apple cultivars and breeding lines that represent much of the diversity currently present in major European breeding programmes and are genetically related by their pedigree were examined for the trueness of their identity and parentage by consistency in marker scores using a genome-covering set of 80 microsatellite (SSR) markers and an ‘identity-by-descent’ approach. One hundred and twenty-five individuals were validated for the trueness-to-type of both their parents and 49 were validated for one of their parents, their second being unknown (23 individuals) or not available in this study (26 individuals). In addition, 15 individuals for which we lacked one of or both the direct parents were validated by consistency with tested parents of earlier generations. Furthermore, the identity of 28 founder cultivars was validated, their marker scores being consistent with descending cultivars and breeding lines. Four of the eight triploids identified were clearly shown to have arisen from unreduced egg cells. The assumed pedigree of 15 further individuals was found to be incorrect; fully consistent pedigrees were suggested for three of the cultivars. The pedigrees of a further eight individuals were confirmed through inference from the molecular data.
Similar content being viewed by others
References
Antofie A, Lateur M, Oger R, Patocchi A, Durel CE, van de Weg WE (2007) A new versatile database created for geneticists and breeders to link molecular and phenotypic data in perennial crops: the AppleBreed DataBase. Bioinformatics 23:882–891
Bink MCAM, Uimari P, Sillanpää MJ, Janss LLG, Jansen RC (2002) Multiple QTL mapping in related plant populations via a pedigree-analysis approach. Theor Appl Genet 104:751–762
Bink MCAM, Boer MP, ter Braak CJF, Jansen J, Voorrips RE, van de Weg WE (2008) Bayesian analysis of complex traits in pedigreed plant populations. Euphytica 161:85–96
Brooks RM, Olmo HP (1972) Register of new fruit and nut varieties, 2nd edn. Berkeley, Los Angeles, University of California Press, p 101
Cabe PR, Baumgaten A, Onan K, Luby JJ, Bedford DS (2005) Using microsatellite analysis to verify breeding records: a study of ‘Honeycrisp’ and other cold-hardy apple cultivars. HortScience 40:15–17
Dermen H (1955) A 2-4-2 chimera of McIntosh apple. J Wash Acad Sci 45:324–327
Gianfranceschi L, Soglio V (2004) The European Project HiDRAS: innovative multidisciplinary approaches to breeding high quality disease resistant apples. Acta Hortic 663:327–330
Gianfranceschi L, Koller B, Seglias N, Kellerhals M, Gessler C (1996) Molecular selection in apple for resistance to scab caused by Venturia inaequalis. Theor Appl Genet 93:199–204
King G, Brown L, Ryder C, Periam N (1998) Microsatellite markers for accession identification, pedigree analysis and assessment of allelic diversity in Malus genetic resources. ECPGR report of a working group on Malus/Pyrus—1st meeting (May 1997, Dublin). In: Maggioni L, Janes R, Hayes A, Swinburne T, Lipman E (eds) Biodiversity international publications, pp 104–113
Kouassi AB, Durel CE, Costa F, Tartarini S, van de Weg E, Evans K, Fernández-Fernández F, Govan C, Boudichevskaja A, Dunemann F, Antofie A, Lateur M, Stankiewicz-Kosyl M, Soska A, Tomala K, Lewandowski M, Rutkovski K, Zurawicz E, Guerra W, Laurens F (2009) Estimation of genetic parameters and prediction of breeding values for apple fruit-quality traits using pedigreed plant material in Europe. Tree Genet Genomes 5:659–672
Kruczyńsha DE, Rutkowski KP (2006) Quality and storage of Czech scab resistant apple cultivars. Phytopath Pol 39:53–61
McKenzie D (1979) The two types of Braeburn. Orchard N Z 1979:182
Morgan J, Richards A (2002) The new book of apples. Ebury Press, London
Patocchi A, Bigler B, Koller B, Kellerhals M, Gessler C (2004) Vr2: a new apple scab resistance gene. Theor Appl Genet 109:1087–1092
Patocchi A, Fernández-Fernández F, Evans K, Gobbin D, Rezzonico F, Boudichevskaia A, Dunemann F, Stankiewicz-Kosyl M, Mathis F, Durel CE, Gianfranceschi L, Costa F, Toller C, Cova V, Mott D, Komjanc M, Barbaro E, Kodde L, Rikkerink E, Gessler C, van de Weg WE (2009a) Development and test of 21 multiplex PCRs composed of SSRs spanning 80% of the apple genome. Tree Genet Genom 5:211–223
Patocchi A, Fernández-Fernández F, Evans K, Silfverburg-Dilworth E, Matasci CL, Gobbin D, Rezzonico F, Boudichevskaia A, Dunemann F, Stankiewicz-Kosyl M, Mathis F, Durel CE, Soglio V, Gianfranceschi L, Costa F, Toller C, Cova V, Mott D, Komjanc M, Barbaro E, Voorrips RE, Rikkerink E, Yamamoto T, Cevik V, Gessler C, van de Weg WE (2009b) Development of a set of apple SSRs markers spanning the apple genome, genotyping of HiDRAS plant material and validation of genotypic data. Acta Hortic 814:603–608
Pereira-Lorenzo S, Ramos-Cabrer AM, Díaz-Hernández MB (2007) Evaluation of genetic identity and variation of local apple cultivars (Malus × domestica Borkh.) from Spain using microsatellite markers. Genet Resour Crop Evol 54:405–420
Promo-Fruit (2009) http://www.promo-fruit.ch/de/rubinette.php
Reim S, Flachowsky H, Hanke M-V, Peil A (2009) Verifying the parents of the Pillnitzer apple cultivars. Acta Hortic 814:319–323
Stark P (1974) The ‘Ozark Gold’ apple. J Am Pomol Soc 28:20–21
Van de Weg WE, Voorips RE, Finkers R, Kodde LP, Jansen J, Bink MCAM (2004) Pedigree genotyping a new pedigree-based approach of QTL identification and allele mining. Acta Hort 663:45–50
Van Treuren R, Kemp H, Ernsting G, Jongejans B, Houtman H, Visser L (2010) Microsatellite genotyping of apple (Malus × domestica Borkh.) genetic resources in the Netherlands: application in collection management and variety identification. Genet Resour Crop Evol 57:853–865
Vinatzer BA, Patocchi A, Tartarini S, Gianfranceschi L, Sansavini S, Gessler C (2004) Isolation of two microsatellite markers from BAC clones of the Vf scab resistance region and molecular characterization of scab-resistant accessions in Malus germplasm. Plant Breeding 123:321–326
Voorrips RE (2007) Pedimap: software for visualization of genetic and phenotypic data in pedigrees. Plant Research International, Wageningen, the Netherlands http://www.plantbreeding.wur.nl/UK/software_pedimap.html
Acknowledgments
Charles Simon of the USDA-ARS of the Plant Genetic Resources Unit, Geneva is acknowledged for supplying leaf samples of PR14-126, PR14-152 and Starr, and Stefano Tartarini of the University of Bologna and Margaret Korbin of RIPF for supplying DNA of all other cultivars and breeding lines not cited in the tables. This publication was carried out with the financial support from the Commission of the European Communities (contract no. QLK5-CT-2002-01492), Directorate-General Research—Quality of Life and Management of Living Resources Programme. It does not necessarily reflect the Commission’s views and in no way anticipates its future policy in this area. Its content is the sole responsibility of the publishers.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Evans, K.M., Patocchi, A., Rezzonico, F. et al. Genotyping of pedigreed apple breeding material with a genome-covering set of SSRs: trueness-to-type of cultivars and their parentages. Mol Breeding 28, 535–547 (2011). https://doi.org/10.1007/s11032-010-9502-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11032-010-9502-5