Abstract
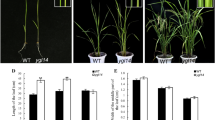
The yellow-green leaf color mutant (Ygm) is a spontaneous mutant derived from the common wheat (Triticum aestivum L.) cultivar Xinong1718. Genetic analysis has shown that a novel single incompletely dominant gene (Y1718) is responsible for the yellow leaf color phenotype. The progeny of Ygm exhibit three distinct leaf color phenotypes, i.e., yellow (Y), yellow-green (Yg), and normal green (G). Y plants have yellow-green leaves in the seedling stage, which become yellow or a strong gold-yellow in the booting stage, with dwarfism and thin tillers until the flowering stage, and underdeveloped thylakoid membranes without well-structured grana in the chloroplasts. Yg plants always have a yellow-green phenotype with a number of well-structured grana that are loosely connected with stroma lamellae in the chloroplasts, where their main agronomic traits are the same as Xinong1718 and G plants, but the seed yield is low. Compared with Xinong1718 and G plants, Y and Yg plants had much lower chlorophyll (Chl) a, Chl b, and carotenoid contents in the booting stage. Molecular analysis using an F2 population and F2:3 lines derived from a cross of Yg and Shannong1 indicated that the Y1718 gene is located on chromosome 2BS, where it is flanked by the simple sequence repeat marker Xwmc25 and expressed sequence tag-sequence tagged sites marker BE498358 at genetic distances of 1.7 and 4.0 cM, respectively. Our results facilitate the fine mapping and gene cloning of Y1718 to explore chlorophyll synthesis, metabolism, and development in wheat.




Similar content being viewed by others
References
Allen GC, Flores-Vergara MA, Krasynanski S, Kumar S, Thompson WF (2006) A modified protocol for rapid DNA isolation from plant tissues using cetyltrimethylammonium bromide. Nat Protoc 1(5):2320–2325. doi:10.1038/nprot.2006.384
An X, Liu C, Zhu C, Sun D, Feng Y, Zhang L (2015) Evaluation and analysis of the main photosynthetic properties and agronomic characteristics of three xantha wheat NILs. J Triticeae Crop 35(11):1476–1482. doi:10.7606/j.issn.1009-1041.2015.11.02
Ansari MJ, AL-ghamdi A, Kumar R, Usmani S, Al-attal Y, Nuru A, Mohamed AA, Singh K, Dhaliwal HS (2013) Characterization and gene mapping of a chlorophyll-deficient mutant clm1 of Triticum monococcum L. Biol Plant 57(3):442–448. doi:10.1007/s10.535-013-0307-3
Arnon DI (1949) Copper enzymes in isolated chloroplasts, polyphenoloxidase in Beta vulgaris. Plant Physiol 24(1):1–15. doi:10.1104/pp.24.1.1
Beale SI (2005) Green genes gleaned. Trends Plant Sci 10(7):309–312. doi:10.1016/j.tplants.2005.05.005
Cao L, Wang H, Sun DJ, Feng Y, Li XJ, Min DH (2010) Chloroplast thylakoid protein composition and characteristics of chlorophyll biosynthesis in a novel aurea mutant of wheat. J Triticeae Crops 30(4):638–643. doi:10.7606/j.issn.1009-1041.2010.04.10
Chen H, Li C, Liu L, Zhao J, Cheng X, Jiang G, Zhai W (2016) The Fd-GOGAT1 mutant gene lc7 confers resistance to Xanthomonas oryzae pv. Oryzae in rice. Sci Rep 6:26411. doi:10.1038/srep26411
Chu CG, Faris JD, Friesen TL, Xu SS (2006) Molecular mapping of hydrid necrosis genes Ne1 and Ne2 in hexaploid wheat using microsatellite markers. Theor Appl Genet 112(7):1374–1381. doi:10.1007/s00122-006-0239-9
Deng XJ, Zhang HQ, Wang Y, He F, Liu JL, Xiao X, Shu ZF, Li W, Wang GH, Wang GL (2014) Mapped clone and functional analysis of leaf-color gene ygl7 in a rice hydrid (Oryza sativa L. ssp. indica). PLoS ONE 9(6):e99564. doi:10.1371/journal.pone.0099564
Fitzmaurice WP, Nguyen LV, Wernsman EA, Thompson WF, Conkling MA (1999) Transposon tagging of the Sulfur gene of tobacco using engineered maize Ac/Dc elements. Genetics 153(4):1919–1928
Freeman TP, Duysen ME, Williams ND (1987) Effects of gene dosage on light harvesting chlorophyll accumulation, chloroplastment, and photosynthesis in wheat (Triticum aestivum). Can J Bot 65(10):2118–2123. doi:10.1139/b87-291
Gupta PK, Balyan HS, Edwards KJ, Isaac P, Korzun V, Röder M, Gautier MF, Joudrier P, Schlatter AR, Dubcovsky J, De La Pena RC, Khairallah M, Penner G, Hayden MJ, Sharp P, Keller B, Wang RCC, Hardouin JP, Jack P, Leroy P (2002) Genetic mapping of 66 new microsatellite (SSR) loci in bread wheat. Theor Appl Genet 105(2):413–422. doi:10.1007/s00122-002-0865-9
Hansson A, Willows RD, Roberts TH, Hansson M (2002) Three semidominant barley mutants with single amino acid substitutions in the smallest magnesium chelatase subunit form defective AAA+ hexamers. Proc Natl Acad Sci USA 99(21):13944–13949. doi:10.1073/pnas.212504499
Hua W, Liu Z, Zhu J, Xie C, Yang T, Zhou Y, Duan X, Sun Q, Liu Z (2009) Identification and genetic mapping of Pm42, a new recessive wheat powdery mildew resistance gene derived from wild emmer (Triticum turgidum var. dicoccoides). Theor Appl Genet 119(2):223–230. doi:10.1007/s00122-009-1031-4
Jung KH, Hur J, Ryu Ch, Choi Y, Chung YY, Miyao A, Hirochika H, An G (2003) Characterization of a rice chlorophyll-deficient mutant using the T-DNA gene-trap system. Plant Cell Physiol 44(5):463–472. doi:10.1093/pcp/pcg064
Kato Y, Sun X, Zhang L, Sakamoto W (2012) Cooperative D1 degradation in the photosystem II repair mediated by chloroplastic proteases in Arabidopsis. Plant Physiol 159(4):1428–1439. doi:10.1104/pp.112.199042
Klindworth DL, Williams ND, Duysen ME (1995) Genetic analysis of chlorina mutants of durum wheat. Crop Sci 35(2):431–436. doi:10.2135/cropsci1995.0011183X003500020026x
Kosambi DD (1943) The estimation of map distances from recombination values. Ann Eugen 12(1):172–175. doi:10.1111/j.1469-1809.1943.tb02321.x
Kosuge K, Watanabe N, Kuboyama T (2011) Comparative genetic mapping of homoeologous genes for the chlorina phenotype in the genus Triticum. Euphytica 179(2):257–263. doi:10.1007/s10681-010-0302-0
Koussevitzky S, Nott A, Mockler TC, Hong F, Sachetto-Martins G, Surpin M, Lim J, Mittler R, Chory J (2007) Signals from chloroplasts converge to regulate nuclear gene expression. Science 316(5825):715–719. doi:10.1126/science.1140516
Kusumi K, Sakata C, Nakamura T, Kawasaki S, Yoshimura A, Iba K (2011) A plastid protein NUS1 is essential for build-up of the genetic system for early development under cold stress conditions. Plant J 68(6):1039–1050. doi:10.1111/j.1365-313X.2011.04755x
Li N, Jia J, Xia C, Liu X, Kong X (2013) Characterization and mapping of novel chlorophyll deficient mutant genes in durum wheat. Breed Sci 63(2):169–175. doi:10.1270/jsbbs.63.169
Li Q, Zhu FY, Gao X, Sun Y, Li S, Tao Y, Lo C, Liu H (2014) Young leaf chlorosis 2 encodes the stroma-localized hemeoxygenase 2 which is required for normal tetrapyrrole biosynthesis in rice. Planta 240(4):701–712. doi:10.1007/s00425-014-2116-0
Li W, Tang S, Zhang S, Shan J, Tang C, Chen Q, Jia G, Han Y, Zhi H, Diao X (2016) Gene mapping and functional analysis of the novel leaf color gene SiYGL1 in foxtail millet [Setaria italica (L.) P. Beauv]. Physiol Plant 157(1):24–37. doi:10.1111/ppl.12405
Liu Z, Zhu J, Cui Y, Liang Y, Wu H, Song W, Liu Q, Yang T, Sun Q, Liu Z (2012) Identification and comparative mapping of a powdery mildew resistance gene derived from wild emmer (Triticum turgidum var. dicoccoides) on chromosome 2BS. Theor Appl Genet 124(6):1041–1049. doi:10.1007/s00122-011-1767-5
Liu J, Wang JY, Yao XY, Zhang Y, Li JQ, Wang XX, Xu ZJ, Chen WF (2015) Characterization and fine mapping of thermo-sensitive chlorophyll deficit mutant1 in rice (Oryza sativa L.). Breed Sci 65(2):161–169. doi:10.1270/jsbbs.65.161
Nagata N, Tanaka R, Satoh S, Tanaka A (2005) Identification of a viny1 reductase gene for chlorophyll synthesis in Arabidopsis thaliana and implications for the evolution of Prochlorococcus species. Plant Cell 17(1):233–240. doi:10.1105/tpc.104.027276
Neatby KW (1963) A chlorophyll mutation in wheat. J Hered 24:159–162
Papenbrock J, Gräfe S, Kruse E, Hänel F, Grimm B (1997) Mg-chelatase of tobacco: identification of a ChlD cDNA sequence encoding a third subunit, analysis of the interaction of the three subunits with the yeast two-hybrid system, and reconstitution of the enzyme activity by co-expression of recombinant CHLD, CHLH and CHLI. Plant J 12(5):981–990. doi:10.1046/j.1365-313X.1997.12050981.x
Qin D, Dong J, Xu F, Guo G, Ge S, Xu Q, Xu Y, Li M (2015) Characterization and fine mapping of a novel barley stage green-revertible albino gene (HvSGRA) by bulked segregant analysis based on SSR assay and specific length amplified fragment sequencing. BMC Genom 16(1):838. doi:10.1186/s12864-015-2015-1
Röder MS, Korzun V, Wendehake K, Plaschke J, Tixier MH, Leroy P, Ganal MW (1998) A microsatellite map of wheat. Genetics 149(4):2007–2023. doi:10.1016/B0-12-227620-5/00113-0
Sawers RJ, Viney J, Farmer PR, Bussey RR, Olsefski G, Anufrikova K, Hunter CN, Brutnell TP (2006) The maize Oil yellow1 (Oy1) gene encodes the I subunit of magnesim chelatase. Plant Mol Biol 60(1):95–106. doi:10.1007/s11103-005-2882-0
Sear ER (1956) Neatby’s virescent. Wheat Inf Serv 3:5
Sear ER (1957) Effect of chromosomes XII and XVI on the action of Neatby’s virescent. Wheat Inf Serv 6:1
Sears LMS, Sears ER (1968) The mutants chlorina-1 and Hermsen’s virescent. In: Proceedings of the 3rd International Wheat Genetics Symposium, Australian Academy of Sciences, Canberra, Australia, pp. 299–304
Shen XK, Ma LX, Zhong SF, Liu N, Zhang M, Chen WQ, Zhou YL, Li HJ, Chang ZJ, Li X, Bai GH, Zhang HY, Tan FQ, Ren ZL, Luo PG (2015) Identification and genetic mapping of the putative Thinopyrum intermedium-derived dominant powdery mildew resistance genePmL962 on wheat chromosome arm 2BS. Theor Appl Genet 128(3):517–528. doi:10.1007/s00122-014-2449-x
Soldatova O, Apchelimov A, Radukina N, Ezhova T, Shestakov S, Ziemann V, Hedtke B, Grimm B (2005) An Arabidopsis mutant that is resistant to the protoporphyrinogen oxidase inhibitor acifluorfen shows regulatory changes in tetrapyrrole biosynthesis. Mol Genet Genom 273(4):311–318. doi:10.1007/s00438-005-1129-6
Somers DJ, Isaac P, Edwards K (2004) A high-density microsatellite consensus map for bread wheat (Triticum aestivum L.). Theor Appl Genet 109(6):1105–1114. doi:10.1007/s00122-004-1740-7
Song QJ, Shi JR, Singh S, Fickus EW, Costa JM, Lewis L, Gill BS, Ward R, Cregan PB (2005) Development and mapping of microsatellite (SSR) markers in wheat. Theor Appl Genet 110(3):550–560. doi:10.1007/s00122-004-1871-x
Sugimoto H, Kusumi K, Noguchi K, Yano M, Yoshimura A, Iba K (2007) The rice nuclear gene, VIRESCENT 2, is essential for chloroplast development and encodes a novel type of guanylate kinase targeted to plastids and mitochondria. Plant J 52(3):512–527. doi:10.1111/j.1365-313X.2007.03251.x
Tanaka A, Tanaka R (2006) Chlorophyll metabolism. Curr Opin Plant Biol 9(3):248–255. doi:10.1016/j.pbi.2006.03.011
Watanabe N, Koval SF (2003) Mapping of chlorina mutant genes on the long arm of homoeologous group 7 chromosomes in common wheat with partial deletion lines. Euphytica 129(3):259–265. doi:10.1023/A:1022276724354
Williams ND, Joppa L, Duysen ME, Freenman TP (1985) Inheritance of three chlorophyll-deficient mutants of common wheat. Crop Sci 25(6):1023–1025. doi:10.2135/cropsci1985.0011183X002500060030x
Yang DL, Jing RL, Chang XP, Li W (2007) Quantitative trait loci mapping for chlorophyll fluorescence and associated traits in wheat (Triticum aestivum). J Intergr Plant Biol 49(5):646–654. doi:10.1111/j.1744-7909.2007.00443.x
Zhang H, Li J, Yoo JH, Yoo SC, Cho SH, Koh HJ, Seo HS, Paek NC (2006) Rice Chlorina-1 and Chlorona-9 encode ChlD and ChlI subunits of Mg-chelatase, a key enzyme for chlorophyll synthesis and chloroplast development. Plant Mol Biol 62(3):325–337. doi:10.1007/s11103-006-9024-z
Zhao H, Yu L, Huai ZX, Wang XH, Ding GD, Chen SS, Li P, Xu FS (2014) Mapping and candidate gene identification defining Bnchd1-1, a locus involved in chlorophyll biosynthesis in Brassica napus. Acta Physiol Plant 36(4):859–870. doi:10.1007/s11738-013-1464-x
Zheng K, Zhao J, Lin D, Chen J, Xu J, Zhou H, Teng S, Dong Y (2016) The rice TCM5 gene encoding a novel Deg protease protein is essential for chloroplast development under high temperatures. Rice 9(1):13. doi:10.1186/s12284-016-0086-5
Zhou K, Ren Y, Lv J, Wang Y, Liu F, Zhou F, Zhao S, Chen S, Peng C, Zhang X, Guo X, Cheng Z, Wang J, Wu F, Jiang L, Wan J (2013) Young leaf chlorosis 1, a chloroplast-localized gene required for chlorophyll and lutein accumulation during early leaf development in rice. Planta 237(1):279–292. doi:10.1007/s00425-012-1756-1
Zhu XG, Long SP, Ort DR (2010) Improving photosynthetic efficiency for greater yield. Annu Rev Plant Biol 61(1):235–261. doi:10.1146/annurev-arplant-042809-112206
Acknowledgements
The study was financially supported by National Sci-Tech Support Project of China (2013BAD01B02), the Zhongying Tang Breeding Foundation of Northwest A&F University and the Basic Scientific Research Fund of Northwest A&F University. We are grateful to Dr. Duncan E. Jackson for improving this manuscript.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors have no conflict of interest to declare.
Rights and permissions
About this article
Cite this article
Zhang, L., Liu, C., An, X. et al. Identification and genetic mapping of a novel incompletely dominant yellow leaf color gene, Y1718, on chromosome 2BS in wheat. Euphytica 213, 141 (2017). https://doi.org/10.1007/s10681-017-1894-4
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s10681-017-1894-4