Abstract
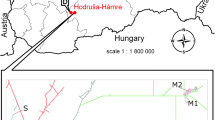
To unveil the potential effect of metal presence to antibiotic tolerance proliferation, four sites of surface landfills containing tailings from metal processing in Slovakia (Hnúšťa, Hodruša, Košice) and Poland (Tarnowskie Góry) were investigated. Tolerance and multitolerance to selected metals (Cu, Ni, Pb, Fe, Zn, Cd) and antibiotics (ampicillin, tetracycline, chloramphenicol, and kanamycin) and interrelationships between them were evaluated. A low bacterial diversity (Shannon–Wiener index from 0.83 to 2.263) was detected in all sampling sites. Gram-positive bacteria, mostly belonging to the phylum Actinobacteria, dominated in three of the four sampling sites. The recorded percentages of tolerant bacterial isolates varied considerably for antibiotics and metals from 0 to 57% and 0.8 to 47%, respectively, among the sampling sites. Tolerances to chloramphenicol (45–57%) and kanamycin (32–45%) were found in three sites. Multitolerance to several metals and antibiotics in the range of 24 to 48% was recorded for three sites. A significant positive correlation (p < 0.05) for the co-occurrence of tolerance to each studied metal and at least one of the antibiotics was observed. Exposure time to the metal (landfill duration) was an important factor for the development of metal- as well as antibiotic-tolerant isolates. The results show that metal-contaminated sites represent a significant threat for human health not only for their toxic effects but also for their pressure to antibiotic tolerance spread in the environment.




Similar content being viewed by others
Data availability
The manuscript has no associated data.
References
Aleem, A., Isar, J., & Malik, A. (2003). Impact of long-term application of industrial wastewater on the emergence of resistance traits in Azotobacter chroococcum isolated from rhizosphere soil. Bioresources and Technology, 86, 7–13. https://doi.org/10.1016/S0960-8524(02)00134-7
Ash, R. J., Mauck, B. A. N. D., & Morgan, M. (2020). Antibiotic resistance of Gram-negative bacteria in rivers, United States. Emerging Infectious Diseases, 8(7), 713. https://doi.org/10.3201/eid0807.010264
Bălăşoiu, M., Bălăşoiu, A. T., Mănescu, R., et al. (2014). Pseudomonas aeruginosa resistance phenotypes and phenotypic highlighting methods. Current Health Sciences Journal, 40(2), 85–92. https://doi.org/10.12865/CHSJ.40.02.01
Chen, J., LI, J., Zhang, H., et al. (2019). Bacterial heavy-metal and antibiotic resistance genes in a copper tailing dam area in Northern China. Frontiers in Microbiology, 10, 1916. https://doi.org/10.3389/fmicb.2019.01916
De Souza, M. J., Nair, S., Bharathi, P. L., et al. (2006). Metal and antibiotic-resistance in psychrotrophic bacteria from Antarctic marine waters. Ecotoxicology, 15(4), 379–384. https://doi.org/10.1007/s10646-006-0068-2
Di Cesare, A., Eckert. E., & Corno, G. (2016). Co-selection of antibiotic and heavy metal resistance in freshwater bacteria. Journal of Limnology, 59-66. https://doi.org/10.4081/jlimnol.2016.1198
Dickinson, A. W., Power, A., Hansen, M. G., et al. (2019). Heavy metal pollution and co-selection for antibiotic resistance: A microbial palaeontology approach. Environment International, 132, 105117. https://doi.org/10.1016/j.envint.2019.105117
Ding, J., Zhu, D., Wang, Y., et al. (2021). Exposure to heavy metal and antibiotic enriches antibiotic resistant genes on the tire particles in soil. Science of the Total Environment, 792, 148417. https://doi.org/10.1016/j.scitotenv.2021.148417
Gadd, G. M. (2010). Metals, minerals and microbes: Geomicrobiology and bioremediation. Microbiology, 156(3), 609–643. https://doi.org/10.1099/mic.0.037143-0
Glibota, N., Grande, M., Galvez, A., et al. (2020). Genetic determinants for metal tolerance and antimicrobial resistance detected in bacteria isolated from soils of olive tree farms.
Hill, T. C. J., Walsh, K. A., Harris, J. A., et al. (2003). Using ecological diversity measures with bacterial communities. FEMS Microbiology Ecology, 43(1), 1–11. https://doi.org/10.1111/j.1574-6941.2003.tb01040.x
Ji, X., Shen, Q., Liu, F., et al. (2012). Antibiotic resistance gene abundances associated with antibiotics and heavy metals in animal manures and agricultural soils adjacent to feedlots in Shanghai. China Journal of Hazardous Materials, 235, 178–185. https://doi.org/10.1016/j.jhazmat.2012.07.040
Jia, L., Wu, W., Zhou, Q., et al. (2022). New insights on the synergetic removal of nutrients and sulfonamides in solid carbon/manganese ore supported denitrification system: Water quality, microbial community and antibiotic resistance genes. Chemical Engineering Journal, p.136992. https://doi.org/10.1016/j.cej.2022.136992
Kasemodel, M. C., Sakamoto, I. K., Varesche, M. B. A., et al. (2019). Potentially toxic metal contamination and microbial community analysis in an abandoned Pb and Zn mining waste deposit. Science of the Total Environment, 675, 367–379. https://doi.org/10.1016/j.scitotenv.2019.04.223
Kwon, J. H., & Powderly, W. G. (2021). The post-antibiotic era is here. Science, 373(6554), 471–471. https://doi.org/10.1126/science.abl5997
Lin, X., Sun, Z., Zhao, L., et al. (2019). The toxicity thresholds of metal (loid) s to soil-dwelling springtail Folsomia candida—A review. Ecotoxicology and Environmental Safety, 180, 632–645. https://doi.org/10.1016/j.ecoenv.2019.04.068
Mandal, M., Das, S. N., & Mandal, S. (2020). Principal component analysis exploring the association between antibiotic resistance and heavy metal tolerance of plasmid-bearing sewage wastewater bacteria of clinical relevance. Access Microbiology, 2(3). https://doi.org/10.1099/acmi.0.000095
Mazhar, S. H., Li, X., Rashid, A., SU, et al. (2021). Co-selection of antibiotic resistance genes, and mobile genetic elements in the presence of heavy metals in poultry farm environments. Science of the Total Environment, 755, 142702. https://doi.org/10.1016/j.scitotenv.2020.142702
Munees. A., & Malik. A. (2014). Prevalence of heavy metal and antibiotic resistance in bacterial isolates from metal polluted soils. Microbiology Journal, 4, 12–21. https://doi.org/10.3923/mj.2014.12.21
Nanda, M., Kumar, V., & Sharma, D. K. (2019). Multimetal tolerance mechanisms in bacteria: The resistance strategies acquired by bacteria that can be exploited to ‘clean-up’ heavy metal contaminants from water. Aquatic Toxicology, 212, 1–10. https://doi.org/10.1016/j.aquatox.2019.04.011
Neethu, C. S., Mujeeb RahimaN, K. M., Saramma, A. V., et al. (2015). Heavy-metal resistance in Gram-negative bacteria isolated from Kongsfjord, Arctic. Canadian Journal of Microbiology, 61(6), 429–435. https://doi.org/10.1139/cjm-2014-0803
Nguyen, C. C., Hugie, C. N., Kile, M. L., et al. (2019). Association between heavy metals and antibiotic-resistant human pathogens in environmental reservoirs: A review. Frontiers of Environmental Science & Engineering, 13(3), 1–17. https://doi.org/10.2147/IDR.S175967
Nosáľová, L., Maliničová, L., Kisková, J., et al. (2021). Cultivable microbiota associated with gold ore from the Rozália gold mine, Hodruša-Hámre, Slovakia. Geomicrobiology Journal, 38(5), 415–425. https://doi.org/10.1080/01490451.2021.1871685
Nosáľová, L. (2019). Štúdium baktérií z biogeochemického cyklu zlata: diploma thesis. Košice: UPJŠ, 2019.
Peng, S., Dolfing, J., Feng, Y., et al. (2018). Enrichment of the antibiotic resistance gene tet (L) in an alkaline soil fertilized with plant derived organic manure. Frontiers in Microbiology, 9, 1140. https://doi.org/10.3389/fmicb.2018.01140
Rajbanshi, A. (2008). Study on heavy metal resistant bacteria in Guheswori sewage treatment plant. Our Nature, 6(1), 52–57. https://doi.org/10.3126/on.v6i1.1655
Rebelo, A., Mourão, J., Freitas, A.R., et al. (2021). Diversity of metal and antibiotic resistance genes in Enterococcus spp. from the last century reflects multiple pollution and genetic exchange among phyla from overlapping ecosystems. Science of The Total Environment, 787, 147548. https://doi.org/10.1016/j.scitotenv.2021.147548
Salam, L. B. (2020). Unravelling the antibiotic and heavy metal resistome of a chronically polluted soil. 3 Biotech, 10, pp.1–23. https://doi.org/10.1007/s13205-020-02219-z
Sinegani, A. A. S., & Younessi, N. (2017). Antibiotic resistance of bacteria isolated from heavy metal-polluted soils with different land uses. Journal of Global Antimicrobial Resistance, 10, 247–255. https://doi.org/10.1016/j.jgar.2017.05.012
Silvester, R., Madhavan, A., Kokkat, A., et al. (2022). Global surveillance of antimicrobial resistance and hypervirulence in Klebsiella pneumoniae from LMICs: An in-silico approach. Science of the Total Environment, 802, 149859. https://doi.org/10.1016/j.scitotenv.2021.149859
Srivastava, P., & Kowshik, M. (2013). Mechanisms of metal resistance and homeostasis in Haloarchaea. Archaea. ID 732864. https://doi.org/10.1155/2013/732864
Timková, I., Lachká, M., Kisková, J., et al. (2020). High frequency of antibiotic tolerance in deep subsurface heterotrophic cultivable bacteria from the Rozália Gold Mine, Slovakia. Environmental Science and Pollution Research, 27(35), 44036–44044. https://doi.org/10.1007/s11356-020-10347-5
Timoney, J. F., Port, J., Giles, J., et al. (1978). Heavy-metal and antibiotic resistance in the bacterial flora of sediments of New York Bight. Applied and Environmental Microbiology, 36(3), 465–472. https://doi.org/10.1128/aem.36.3.465-472.1978
Walter, T., Klim, J., Jurkowski, et al. (2020). Plasmidome of an environmental Acinetobacter lwoffii another difference witch originating from a former gold and arsenic mine. Plasmid, 110, 102505. https://doi.org/10.1016/j.plasmid.2020.102505
Wu, L., Chen, J., Xiao, et al. (2018). Barcoded pyrosequencing reveals a shift in the bacterial community in the rhizosphere and rhizoplane of Rehmannia glutinosa under consecutive monoculture. International Journal of Molecular Sciences, 19(3), 850. https://doi.org/10.3390/ijms19030850
Wu, X., Cobbina, S. J., Mao, G., et al. (2016). A review of toxicity and mechanisms of individual and mixtures of heavy metals in the environment. Environmental Science and Pollution Research, 23(9), 8244–8259. https://doi.org/10.1007/s11356-016-6333-x
Zagui, G. S., Moreira, N. C., Santos, et al. (2020). High occurrence of heavy metal tolerance genes in bacteria isolated from wastewater: A new concern?. Environmental Research, p. 110352. https://doi.org/10.1016/j.envres.2020.110352
Zhang, Y., Gu, A. Z., Cen, T., et al. (2018a) Sub-inhibitory concentrations of heavy metals facilitate the horizontal transfer of plasmid-mediated antibiotic resistance genes in water environment. Environmental Pollution, 237, 74-82. https://doi.org/10.1016/j.envpol.2018.01.032
Zhang, Y., Zhang, H., Zhang, Z., et al. (2018b). PH effect on heavy metal release from a polluted sediment. Journal of Chemistry, 2018. https://doi.org/10.1155/2018/7597640
Zhao, W., Wang, B., & Yu, G. (2018). Antibiotic resistance genes in China: Occurrence, risk, and correlation among different parameters. Environmental Science and Pollution Research, 25(22), 21467–21482.https://doi.org/10.1007/s11356-018-2507-z
Zhou, Q., Wang, M., Zhong, X., et al. (2019). Dissemination of resistance genes in duck/fish polyculture ponds in Guangdong Province: Correlations between Cu and Zn and antibiotic resistance genes. Environmental Science and Pollution Research, 26(8), 8182–8193. https://doi.org/10.1007/s11356-018-04065-2
Zhu, Y. G., Johnson, T. A., Su, J. Q., et al. (2013). Diverse and abundant antibiotic resistance genes in Chinese swine farms. Proceedings of the National Academy of Sciences, 110(9), 3435–3440. https://doi.org/10.1073/pnas.1222743110
Funding
The work was supported by financial aid from Slovak Grant Agency (project No. VEGA 1/0018/22 and VEGA 2/0142/19) and from Polish National Agency for Academic Exchange (PPN/BIL/2018/1/00026/U/00001).
Author information
Authors and Affiliations
Contributions
Author M.L. wrote the paper. Authors J.S-K and A.L. conceived and designed the analysis. Authors M.L., K.S., and V.P. performed the MIC analysis. Author M.L. performed statistical analyses. Authors L.N. and I.T. performed the MALDI-TOF analysis. Authors J.W. and I.J. performed the chemical analysis of the ore. M.L., J.S-K., and A.L. edited the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
Jana Sedlákova-Kadukova and Alena Luptakova have received research support from Slovak Grant Agency. Joanna Willner received research support from Polish National Agency for Academic Exchange. The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Highlights
• Metals enhance the spread of antibiotics tolerance in metal-polluted environments
• Old mine landfills are more likely to spread antibiotic resistance than active ones
• Correlation between antibiotic and metal tolerance was proved
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Lachka, M., Soltisova, K., Nosalova, L. et al. Metal-containing landfills as a source of antibiotic tolerance. Environ Monit Assess 195, 262 (2023). https://doi.org/10.1007/s10661-022-10873-4
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s10661-022-10873-4