Abstract
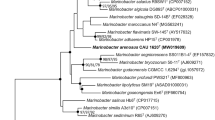
A novel Gram-stain-negative, aerobic, and rod-shaped bacterium with gliding motility, named strain ANRC-HE7T, was isolated from the seawater of Biological Bay adjacent to Fildes Peninsula, Antarctica. The optimal growth of this strain occurred at 28 °C, pH 7.5, and in the presence of 1.0% (w/v) NaCl. Strain ANRC-HE7T can produce amylase and harbors gene clusters involved in cellulose degradation. Phylogenetic analysis based on the 16S rRNA gene sequence showed that strain ANRC-HE7T formed a distinct lineage within the genus Maribacter and was closely related to Maribacter luteus RZ05T (98.4% sequence similarity), Maribacter polysiphoniae LMG 23671T (98.3%), and Maribacter arenosus CAU 1321T (97.3%). However, digital DNA–DNA hybridization and average nucleotide identity values between strain ANRC-HE7T and closely related strains were 17.4–49.1% and 70.9–92.7%, much lower than the cutoff values of 70% and 95%, respectively. On the other hand, strain ANRC-HE7T shared characteristics with most type strains within the genus. Its respiratory quinone was MK-6. The major fatty acids were iso-C15:0, summed feature 3 (C16:1 ω7c and/or C16:1 ω6c), and anteiso-C15:0. The major polar lipids were phosphatidylethanolamine, two unidentified aminolipids, four unidentified phospholipids, and five unidentified glycolipids. The DNA G + C content of strain ANRC-HE7T was 40.1%. Based on the results of the biochemical, phylogenetic, and chemotaxonomic analyses, strain ANRC-HE7T is suggested to represent a novel species of the genus Maribacter, for which the name Maribacter aquimaris sp. nov. is proposed. The type strain is ANRC-HE7T (= MCCC 1K03787T = KCTC 72532T).



Similar content being viewed by others
Data availability
The GenBank accession numbers of the 16S rRNA gene and the draft genome sequences of strain ANRC-HE7T are MN658399 and JABTCF000000000, respectively. All other data generated or analysed during this study are included in this published article and its supplementary information files.
References
Auch AF, Klenk HP, Göker M (2010) Standard operating procedure for calculating genome-to-genome distances based on high-scoring segment pairs. Stand Genom Sci 2:142–148. https://doi.org/10.4056/sigs.541628
Bankevich A, Nurk S, Antipov D, Gurevich AA, Dvorkin M, Kulikov AS, Lesin VM, Nikolenko SI, Pham S, Prjibelski AD, Pyshkin AV, Sirotkin AV, Vyahhi N, Tesler G, Alekseyev MA, Pevzner PA (2012) SPAdes: a new genome assembly algorithm and its applications to single-cell sequencing. J Comput Biol 19(5):455–477. https://doi.org/10.1089/cmb.2012.0021
Barbeyron T, Carpentier F, L’haridon S, Schuler M, Michel G, Amann R (2008) Description of Maribacter forsetii sp. nov., a marine Flavobacteriaceae isolated from North Sea water, and emended description of the genus Maribacter. Int J Syst Evol Microbiol 58:790–797. https://doi.org/10.1099/ijs.0.65469-0
Blin K, Shaw S, Steinke K, Villebro R, Ziemert N, Lee SY, Medema MH, Weber T (2019) antiSMASH 5.0: updates to the secondary metabolite genome mining pipeline. Nucl Acids Res 47:81–87. https://doi.org/10.1093/nar/gkz310
Buchfink B, Xie C, Huson DH (2014) Fast and sensitive protein alignment using DIAMOND. Nat Methods 12:59–60. https://doi.org/10.1038/nmeth.3176
Dong X, Cai M (2001) Determinative manual for routine bacteriology (English translation). Scientific Press, Beijing
Embley TM, Wait R (1994) Structural lipids of eubacteria. In: Goodfellow M, O’Donnell AG (eds) Chemical methods in prokaryotic systematics. Wiley, Chichester, pp 121–161
Fang C, Wu YH, Xamxidin M, Wang CS, Xu XW (2017) Maribacter cobaltidurans sp. nov., a heavy-metal-tolerant bacterium isolated from deep-sea sediment. Int J Syst Evol Microbiol 67:5261–5267. https://doi.org/10.1099/ijsem.0.002458
Felföldi T, Somogyi B, Marialigeti K, Vörös L (2009) Characterization of photoautotrophic picoplankton assemblages in turbid, alkalinelakes of the Carpathian Basin (Central Europe). J Limnol 68:385–395. https://doi.org/10.4081/jlimnol.2009.385
Felsenstein J (1981) Evolutionary trees from DNA sequences: a maximum likelihood approach. J Mol Evol 17:368–376. https://doi.org/10.1007/BF01734359
Felsenstein J (1985) Confidence limits on phylogenies: an approach using the bootstrap. Evolution 39(4):783–791. https://doi.org/10.1111/j.1558-5646.1985.tb00420.x
Hu J, Yang QQ, Ren Y, Zhang WW, Zheng G, Sun C, Pan J, Zhu XF, Zhang XQ, Wu M (2015) Maribacter thermophilus sp. nov., isolated from an algal bloom in an intertidal zone, and emended description of the genus Maribacter. Int J Syst Evol Microbiol 65:36–41. https://doi.org/10.1099/ijs.0.064774-0
Jackson SA, Kennedy J, Morrissey JP, O’Gara F, Dobson AD (2015) Maribacter spongiicola sp. nov. and Maribacter vaceletii sp. nov., isolated from marine sponges, and emended description of the genus Maribacter. Int J Syst Evol Microbiol 65:2097–2103. https://doi.org/10.1099/ijs.0.000224
Jin M, Kim M, Kim JY, Song HS, Cha IT, Roh SW, Lee SJ (2017) Maribacter pelagius sp. nov., isolated from seawater. Int J Syst Evol Microbiol 67:3834–3839. https://doi.org/10.1099/ijsem.0.002203
Kang H, Cha I, Kim H, Joh K (2018) Maribacter maritimus sp. nov., isolated from seawater. Int J Syst Evol Microbiol 68:2431–2436. https://doi.org/10.1099/ijsem.0.002843
Kämpfer P, Kroppenstedt RM (1996) Numerical analysis of fatty acid patterns of coryneform bacteria and related taxa. Can J Microbiol 42:989–1005. https://doi.org/10.1139/m96-128
Khan SA, Jeong SE, Baek JH, Jeon CO et al (2020) Maribacter algicola sp. nov., isolated from a marine red alga, Porphyridium marinum, and transfer of Maripseudobacter aurantiacus Chen et al. 2017 to the genus Maribacter as Maribacter aurantiacus comb. nov. Int J Syst Evol Microbiol 70:797–804. https://doi.org/10.1099/ijsem.0.003828
Kim KH, Jin HM, Jeong HI, Jeon CO (2016) Maribacter lutimaris sp. nov., isolated from marine sediment. Int J Syst Evol Microbiol 66:1773–1778. https://doi.org/10.1099/ijsem.0.000942
Kumar S, Stecher G, Tamura K (2016) MEGA7: molecular evolutionary genetics analysis version 7.0 for bigger datasets. Mol Biol Evol 33:1870–1874. https://doi.org/10.1093/molbev/msw054
Lo N, Jin HM, Jeon CO (2013) Maribacter aestuarii sp. nov., isolated from tidal flat sediment, and an emended description of the genus Maribacter. Int J Syst Evol Microbiol 63:3409–3414. https://doi.org/10.1099/ijs.0.050054-0
Liu A, Zhang YJ, Liu DK, Li XZ (2020) Maribacter luteus sp. nov., a marine bacterium isolated from intertidal sand of the Yellow Sea. Int J Syst Evol Microbiol 70:3497–3503. https://doi.org/10.1099/ijsem.0.004206
Meier-Kolthoff JP, Göker M, Spröer C, Klenk HP (2013) When should a DDH experiment be mandatory in microbial taxonomy? Arch Microbiol 195:413–418. https://doi.org/10.1007/s00203-013-0888-4
Na SI, Kim YO, Yoon SH, Ha SM, Baek I, Chun J (2018) UBCG: Up-to-date bacterial core gene set and pipeline for phylogenomic tree reconstruction. J Microbiol 56(4):280–285. https://doi.org/10.1007/s12275-018-8014-6
Nedashkovskaya OI, Kim SB, Han SK, Lysenko AM, Rohde M, Rhee MS, Frolova GM, Falsen E, Mikhailov VV, Bae KS (2004) Maribacter gen. nov., a new member of the family Flavobacteriaceae, isolated from marine habitats, containing the species Maribacter Maribacter sp. nov., Maribacter aquivivus sp. nov., Maribacter orientalis sp. nov. and Maribacter ulvicola sp. nov. Int J Syst Evol Microbiol 54:1017–1023. https://doi.org/10.1099/ijs.0.02849-0
Nedashkovskaya OI, Vancanneyt M, De Vos P, Kim SB, Lee MS, Mikhailov VV (2007) Maribacter polysiphoniae sp. nov., isolated from a red alga. Int J Syst Evol Microbiol 57:2840–2843. https://doi.org/10.1099/ijs.0.65181-0
Rzhetsky A, Nei M (1992) Statistical properties of the ordinary leastsquares, generalized least-squares, and minimum-evolution methods of phylogenetic inference. J Mol Evol 35:367–375. https://doi.org/10.1007/BF00161174
Saitou N, Nei M (1987) The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol Biol Evol 4:406–425. https://doi.org/10.1093/oxfordjournals.molbev.a040454
Sasser M (1990) Identification of bacteria by gas chromatography of cellular fatty acids. MIDI Technical Note 101. Newark, DE: MIDI
Thongphrom C, Kim JH, Kim W (2016) Maribacter arenosus sp. nov., isolated from marine sediment. Int J Syst Evol Microbiol 66:4826–4831. https://doi.org/10.1099/ijsem.0.001436
Tindall B (1990a) A comparative study of the lipid composition of Halobacterium saccharovorum from various sources. Syst Appl Microbiol 13:128–130. https://doi.org/10.1016/s0723-2020(11)80158-x
Tindall B (1990b) Lipid composition of Halobacterium lacusprofundi. FEMS Microbiol Lett 66:199–202. https://doi.org/10.1111/j.1574-6968.1990.tb03996.x
Wayne LG, Brenner DJ, Colwell RR, Grimont PAD, Kandler O, Krichevsky MI, Moore LH, Moore WEC, Murray RGE, Stackebrandt E, Starr MP, Truper HG (1987) Report of the ad hoc committee on reconciliation of approaches to bacterial systematics. Int J Syst Evol Microbiol 37:463–464. https://doi.org/10.1016/s0176-6724(88)80120-2
Xu L, Dong Z, Fang L, Luo Y, Wei Z, Guo H, Zhang G, Gu YQ, Coleman-Derr D, Xi Q, Wang Y (2019) OrthoVenn2: a web server for whole-genome comparison and annotation of orthologous clusters across multiple species. Nucl Acids Res 47:W52–W58. https://doi.org/10.1093/nar/gkz333
Yoon JH, Kang SJ, Lee SY, Lee CH, Oh TK (2005) Maribacter dokdonensis sp. nov., isolated from sea water off a Korean island, Dokdo. Int J Syst Evol Microbiol 55:2051–2055. https://doi.org/10.1099/ijs.0.63777-0
Yoon SH, Ha SM, Lim J, Kwon S, Chun J (2017a) A large-scale evaluation of algorithms to calculate average nucleotide identity. Antonie Van Leeuwenhoek 110:1281–1286. https://doi.org/10.1007/s10482-017-0844-4
Yoon SH, Ha SM, Kwon S et al (2017b) Introducing EzBioCloud: a taxonomically united database of 16S rRNA gene sequences and whole-genome assemblies. Int J Syst Evol Microbiol 67:1613–1617. https://doi.org/10.1099/ijsem.0.001755
Acknowledgements
Funding was provided by Special Foundation for National Science and Technology Basic Research Program of China (No. 2019FY100706), the Science Fund for Distinguished Young Scholars of Anhui Province (No. 2008085J12), and the Science and Technology Major Project of Anhui Province (No. 202103a06020006).
Funding
The authors have not disclosed any funding.
Author information
Authors and Affiliations
Contributions
YFZ, YWZ, LLM, WF, and MMH performed the experiments and wrote the manuscript, YZX and ZMF designed the study and revised the manuscript. All the authors read and approved the final version of the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical statement
This article does not contain any studies with animals performed by any of the authors.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Zhang, Y., Zhai, Y., Mu, L. et al. Maribacter aquimaris sp. nov., isolated from seawater adjacent to Fildes Peninsula, Antarctica. Antonie van Leeuwenhoek 116, 753–761 (2023). https://doi.org/10.1007/s10482-023-01844-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10482-023-01844-x