Abstract
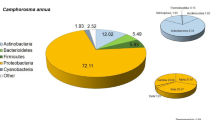
Hypersaline ecosystems host a particular microbiota, which can be specifically recruited by halophytes. In order to broaden our knowledge of hypersaline ecosystems, an in natura study was conducted on the microbiota associated with the halophyte Halocnemum strobilaceum from alkaline-saline arid soil in Algeria. We collected and identified a total of 414 strains isolated from root tissues (RT), root-adhering soil (RAS), non-adhering rhizospheric soil (NARS) and bulk soil (BS) using different NaCl concentrations. Our data showed that halophilic and halotolerant bacterial isolates in BS and the rhizosphere belonged to 32 genera distributed in Proteobacteria (49%), Firmicutes (36%), Actinobacteria (14%) and Bacteroidetes (1%). Bacterial population size and species diversity were greatly increased in the rhizosphere (factor 100). The reservoir of diversity in BS was dominated by the genera Bacillus and Halomonas. Bacillus/Halomonas ratio decreased with the proximity to the roots from 2.2 in BS to 0.3 at the root surface. Salt screening of the strains showed that species belonging to nine genera were able to grow up to 5.1 M NaCl. Thus, we found that H. strobilaceum exerted a strong effect on the diversity of the recruited microbiota with an affinity strongly attributed to the genus Halomonas.







Similar content being viewed by others
Data availability
Sequences of the strains used in this study were deposited at the National Center for Biotechnology Information Sequence Read Archive under the accession number PRJNA765921.
Abbreviations
- RT:
-
Root tissues
- RAS:
-
Root adhering soil
- NARS:
-
Non-adhering rhizospheric soil
- BS:
-
Bulk soil
References
Al-Mailem DM, Sorkhoh NA, Marafie M et al (2010) Oil phytoremediation potential of hypersaline coasts of the Arabian Gulf using rhizosphere technology. Bioresour Technol 101:5786–5792. https://doi.org/10.1016/j.biortech.2010.02.082
Alsharif W, Saad MM, Hirt H (2020) Desert microbes for boosting sustainable agriculture in extreme environments. Front Microbiol 11:1666. https://doi.org/10.3389/fmicb.2020.01666
Bachar A, Al-Ashhab A, Soares MIM et al (2010) Soil microbial abundance and diversity along a low precipitation gradient. Microb Ecol 60:453–461. https://doi.org/10.1007/s00248-010-9727-1
Bachar A, Soares MIM, Gillor O (2012) The effect of resource islands on abundance and diversity of bacteria in arid soils. Microb Ecol 63:694–700. https://doi.org/10.1007/s00248-011-9957-x
Bais HP, Weir TL, Perry LG et al (2006) The role of root exudates in rhizosphere interactions with plants and other organisms. Annu Rev Plant Biol 57:233–266. https://doi.org/10.1146/annurev.arplant.57.032905.105159
Benardini JN, Sawyer J, Venkateswaran K, Nicholson WL (2003) Spore UV and acceleration resistance of endolithic Bacillus pumilus and Bacillus subtilis isolates obtained from Sonoran Desert basalt: implications for Lithopanspermia. Astrobiology 3:709–717. https://doi.org/10.1089/153110703322736033 (PMID: 14987476)
Bhagat N, Raghav M, Dubey S, Bedi N (2021) Bacterial exopolysaccharides: insight into their role in plant abiotic stress tolerance. J Microbiol Biotechnol 31:1045–1059. https://doi.org/10.4014/jmb.2105.05009
Bibi F, Ullah I, Alvi SA et al (2017) Isolation, diversity, and biotechnological potential of rhizo- and endophytic bacteria associated with mangrove plants from Saudi Arabia. Genet Mol Res 16:1–12. https://doi.org/10.4238/gmr16029657
Bibi F, Strobel GA, Naseer MI et al (2018) Halophytes-associated endophytic and rhizospheric bacteria: diversity, antagonism and metabolite production. Biocontrol Sci Technol 28:192–213. https://doi.org/10.1080/09583157.2018.1434868
Brader G, Compant S, Mitter B et al (2014) Metabolic potential of endophytic bacteria. Curr Opin Biotechnol 27:30–37. https://doi.org/10.1016/j.copbio.2013.09.012
Bulgarelli D, Garrido-oter R, Mchardy AC, Schulze-lefert P (2015) Structure and function of the bacterial root microbiota in wild and domesticated barley. Cell Host Microbe 17:392–403. https://doi.org/10.1016/j.chom.2015.01.011
Chandra P, Dhuli P, Verma P et al (2020) Culturable microbial diversity in the rhizosphere of different biotypes under variable salinity. Trop Ecol 61:291–300. https://doi.org/10.1007/s42965-020-00089-3
Chaudhary DR, Saxena J, Lorenz N et al (2012) Microbial profiles of rhizosphere and bulk soil microbial communities of biofuel crops switchgrass (Panicum virgatum L.) and jatropha (Jatropha curcas L.). Appl Environ Soil Sci 2012:906864. https://doi.org/10.1155/2012/906864
Chun J, Oren A, Ventosa A et al (2018) Proposed minimal standards for the use of genome data for the taxonomy of prokaryotes. Int J Syst Evol Microbiol 68:461–466. https://doi.org/10.1099/ijsem.0.002516
Edwards J, Johnson C, Santos-medellín C et al (2015) Structure, variation, and assembly of the root-associated microbiomes of rice. Proc Natl Acad Sci USA 112:E911–E920. https://doi.org/10.1073/pnas.1414592112
El Khalloufi F, Oufdou K, Bertrand M et al (2016) Microbiote shift in the Medicago sativa rhizosphere in response to cyanotoxins extract exposure. Sci Total Environ 539:135–142. https://doi.org/10.1016/j.scitotenv.2015.08.127
Etesami H, Beattie GA (2018) Mining halophytes for plant growth-promoting halotolerant bacteria to enhance the salinity tolerance of non-halophytic crops. Front Microbiol 9:148. https://doi.org/10.3389/fmicb.2018.00148
Feng J, Wang J, Fan P et al (2015) High-throughput deep sequencing reveals that microRNAs play important roles in salt tolerance of euhalophyte Salicornia europaea. BMC Plant Biol 15:1–17. https://doi.org/10.1186/s12870-015-0451-3
Flowers TJ, Colmer TD (2008) Salinity tolerance in halophytes. New Phytol 179:945–963. https://doi.org/10.1111/j.1469-8137.2008.02531.x
Flowers TJ, Colmer TD (2015) Plant salt tolerance: adaptations in halophytes. Ann Bot 115:327–331. https://doi.org/10.1093/aob/mcu267
Flowers TJ, Hajibagheri MA, Clipson NJW (1986) Halophytes. Q Rev Biol 61:313–337. https://doi.org/10.1086/415032
Gao L, Ma J, Liu Y et al (2021) Diversity and biocontrol potential of cultivable endophytic bacteria associated with halophytes from the west Aral Sea Basin. Microorganisms 9:1448. https://doi.org/10.3390/microorganisms9071448
Gee GW, Bauder JW (1986) Particle-size analysis. In: Klute A (ed) Methods of soil analysis: part 1 physical and mineralogical methods, agronomy monograph no. 9 (2nd edn). American Society of Agronomy/Soil Science Society of America, Madison, USA, pp 383–411. https://doi.org/10.2136/sssabookser5.1.2ed.c15
Haichar FEZ, Marol C, Berge O et al (2008) Plant host habitat and root exudates shape soil bacterial community structure. ISME J 2:1221–1230. https://doi.org/10.1038/ismej.2008.80
Hardoim PR, van Overbeek LS, Berg G et al (2015) The hidden world within plants: ecological and evolutionary considerations for defining functioning of microbial endophytes. Microbiol Mol Biol Rev 79:293–320. https://doi.org/10.1128/mmbr.00050-14
IUSS Working Group WRB (2015) World reference base for soil resources 2014, update 2015 international soil classification system for naming soils and creating legends for soil maps. Worls Soil Resources Reports No. 106 FAO, Rome, E-ISBN 978-92-5-108370-3
Jafari B, Hanifezadeh M, Parvin MSJ (2012) Molecular study of bacteria associated with Salicornia symbiotic bacteria as a candidate for Hormozgan salty zone culturing by Persian Gulf water irrigation. African J Microbiol Res 6:4687–4695. https://doi.org/10.5897/ajmr11.1132
Jha B, Gontia I, Hartmann A (2012) The roots of the halophyte Salicornia brachiata are a source of new halotolerant diazotrophic bacteria with plant growth-promoting potential. Plant Soil 356:265–277. https://doi.org/10.1007/s11104-011-0877-9
Kasim WA, Gaafar RM, Abou-Ali RM et al (2016) Effect of biofilm forming plant growth promoting rhizobacteria on salinity tolerance in barley. Ann Agric Sci 61:217–227. https://doi.org/10.1016/j.aoas.2016.07.003
Kearl J, McNary C, Lowman JS et al (2019) Salt-tolerant halophyte rhizosphere bacteria stimulate growth of alfalfa in salty soil. Front Microbiol 10:1849. https://doi.org/10.3389/fmicb.2019.01849
Le Houérou HN (1993) Salt-tolerant plants for the arid regions of the Mediterranean isoclimatic zone. Towards Ration Use High Salin Toler Plants I:403–422. https://doi.org/10.1007/978-94-011-1858-3_42
Li Y, Kong Y, Teng D et al (2018) Rhizobacterial communities of five co-occurring desert halophytes. PeerJ 6:1–27. https://doi.org/10.7717/peerj.5508
Llamas I, del Moral A, Martínez-Checa F et al (2006) Halomonas maura is a physiologically versatile bacterium of both ecological and biotechnological interest. Antonie Van Leeuwenhoek 89:395–403. https://doi.org/10.1007/s10482-005-9043-9
Mapelli F, Marasco R, Rolli E et al (2013) Potential for plant growth promotion of rhizobacteria associated with Salicornia growing in Tunisian hypersaline soils. Biomed Res Int 2013:248078. https://doi.org/10.1155/2013/248078
Marasco R, Rolli E, Ettoumi B et al (2012) A drought resistance-promoting microbiome is selected by root system under desert farming. PLoS ONE 7:e48479. https://doi.org/10.1371/journal.pone.0048479
Marasco R, Mapelli F, Rolli E et al (2016) Salicornia strobilacea (synonym of Halocnemum strobilaceum) grown under different tidal regimes selects rhizosphere bacteria capable of promoting plant growth. Front Microbiol 7:1286. https://doi.org/10.3389/fmicb.2016.01286
McCaig AE, Glover LA, Prosser JI (1999) Molecular analysis of bacterial community structure and diversity in unimproved and improved upland grass pastures. Appl Environ Microbiol 65:1721–1730. https://doi.org/10.1128/aem.65.4.1721-1730.1999
Menasria T, Monteoliva-Sánchez M, Benammar L et al (2019) Culturable halophilic bacteria inhabiting Algerian saline ecosystems: a source of promising features and potentialities. World J Microbiol Biotechnol 35:132. https://doi.org/10.1007/s11274-019-2705-y
Merino C, Nannipieri P, Matus F (2015) Soil carbon controlled by plant, microorganism and mineralogy interactions. J Soil Sci Plant Nutr 15:321–332. https://doi.org/10.4067/s0718-95162015005000030
Mishra A, Tanna B (2017) Halophytes: potential resources for salt stress tolerance genes and promoters. Front Plant Sci 8:829. https://doi.org/10.3389/fpls.2017.00829
Mukhtar S, Mirza BS, Mehnaz S et al (2018) Impact of soil salinity on the microbial structure of halophyte rhizosphere microbiome. World J Microbiol Biotechnol 34:1–17. https://doi.org/10.1007/s11274-018-2509-5
Mukhtar S, Mehnaz S, Malik KA (2021) Comparative study of the rhizosphere and root endosphere microbiomes of Cholistan Desert plants. Front Microbiol 12:1–14. https://doi.org/10.3389/fmicb.2021.618742
Oliveira V, Gomes NCM, Cleary DFR et al (2014) Halophyte plant colonization as a driver of the composition of bacterial communities in salt marshes chronically exposed to oil hydrocarbons. FEMS Microbiol Ecol 90:647–662. https://doi.org/10.1111/1574-6941.12425
Ozenda P (1982) Les végétaux dans la biosphère. Doin, Paris
Pawlowski J, Holzmann M (2002) Molecular phylogeny of Foraminifera – a review. Eur J Protistol 10:1–10
Qurashi AW, Sabri AN (2012) Bacterial exopolysaccharide and biofilm formation stimulate chickpea growth and soil aggregation under salt stress. Brazilian J Microbiol 43:1183–1191. https://doi.org/10.1590/S1517-83822012000300046
Rathore AP, Chaudhary DR, Jha B (2017) Seasonal patterns of microbial community structure and enzyme activities in coastal saline soils of perennial halophytes. L Degrad Dev 28:1779–1790. https://doi.org/10.1002/ldr.2710
Ravindran KC, Venkatesan K, Balakrishnan V et al (2007) Restoration of saline land by halophytes for Indian soils. Soil Biol Biochem 39:2661–2664. https://doi.org/10.1016/j.soilbio.2007.02.005
Rengasamy P (2010) Soil processes affecting crop production in salt-affected soils. Funct Plant Biol 37:613–620. https://doi.org/10.1071/FP09249
Ruppel S, Franken P, Witzel K (2013) Properties of the halophyte microbiome and their implications for plant salt tolerance. Funct Plant Biol 40:940–951. https://doi.org/10.1071/FP12355
Sáenz-Mata J, Palacio-Rodríguez R, Sánchez-Galván H, Balagurusamy N (2016) Plant growth promoting rhizobacteria associated to halophytes: Potential applications in agriculture. In: Khan MA, Boër B, Özturk M et al (eds) Sabkha ecosystems, vol 5. The Americas, Springer N, pp 11–27
Sampedro I, Pérez-Mendoza D, Toral L et al (2020) Effects of halophyte root exudates and their components on chemotaxis, biofilm formation and colonization of the halophilic bacterium Halomonas Anticariensis FP35T. Microorganisms 8:575. https://doi.org/10.3390/microorganisms8040575
Saul-Tcherkas V, Steinberger Y (2011) Soil microbial diversity in the vicinity of a Negev Desert shrub-Reaumuria negevensis. Microb Ecol 61:64–81. https://doi.org/10.1007/s00248-010-9763-x
Sgroy V, Cassán F, Masciarelli O et al (2009) Isolation and characterization of endophytic plant growth-promoting (PGPB) or stress homeostasis-regulating (PSHB) bacteria associated to the halophyte Prosopis strombulifera. Appl Microbiol Biotechnol 85:371–381. https://doi.org/10.1007/s00253-009-2116-3
Shabala S (2013) Learning from halophytes: physiological basis and strategies to improve abiotic stress tolerance in crops. Ann Bot 112:1209–1221. https://doi.org/10.1093/aob/mct205
Shabala S, Bose J, Hedrich R (2014) Salt bladders: do they matter? Trends Plant Sci 19:687–691. https://doi.org/10.1016/j.tplants.2014.09.001 (Epub 2014 Oct 28 PMID: 25361704)
Shurigin V, Egamberdieva D, Li L et al (2020) Endophytic bacteria associated with halophyte Seidlitzia rosmarinus Ehrenb. ex Boiss. from saline soil of Uzbekistan and their plant beneficial traits. J Arid Land 12:730–740. https://doi.org/10.1007/s40333-020-0019-4
Siddikee MA, Chauhan PS, Anandham R et al (2010) Isolation, characterization, and use for plant growth promotion under salt stress, of ACC deaminase-producing halotolerant bacteria derived from coastal soil. J Microbiol Biotechnol 20:1577–1584. https://doi.org/10.4014/jmb.1007.07011
Stackebrandt E, Goebel BM (1994) Taxonomic note: A place for DNA- DNA reassociation and 16S rRNA sequence analysis in the present species definition in bacteriology. Int J Syst Bacteriol 44:846–849. https://doi.org/10.1099/00207713-44-4-846
Szymańska S, Płociniczak T, Piotrowska-Seget Z, Hrynkiewicz K (2016) Endophytic and rhizosphere bacteria associated with the roots of the halophyte Salicornia europaea L. - community structure and metabolic potential. Microbiol Res 192:37–51. https://doi.org/10.1016/j.micres.2016.05.012
Tian XY, Zhang CS (2017) Illumina-based analysis of endophytic and rhizosphere bacterial diversity of the coastal halophyte Messerschmidia sibirica. Front Microbiol 8:2288. https://doi.org/10.3389/fmicb.2017.02288
Tiwari S, Singh P, Tiwari R et al (2011) Salt-tolerant rhizobacteria-mediated induced tolerance in wheat (Triticum aestivum) and chemical diversity in rhizosphere enhance plant growth. Biol Fertil Soils 47:907–916. https://doi.org/10.1007/s00374-011-0598-5
Upadhyay SK, Singh JS, Singh DP (2011) Exopolysaccharide-producing plant growth-promoting rhizobacteria under salinity condition. Pedosphere 21:214–222. https://doi.org/10.1016/S1002-0160(11)60120-3
Vlisidis AC (1966) The determination of sulfate and sulfide sulfur in rocks or minerals. Geol Surv Bull 1214-D:D1–D5
Walkley A, Black IA (1934) An examination of the degtjareff method for determining soil organic matter, and a proposed modification of the chromic acid titration method. Soil Sci 37:29–38. https://doi.org/10.1097/00010694-193401000-00003
Weisburg WG, Barns SM, Pelletier DA, Lane DJ (1991) 16S ribosomal DNA amplification for phylogenetic study. J Bacteriol 173:697–703. https://doi.org/10.1128/jb.173.2.697-703.1991
Yamamoto K, Matsutani M, Shiwa Y et al (2020) Comparative analysis of bacterial diversity and community structure in the rhizosphere and root endosphere of two halophytes, Salicornia europaea and Glaux maritima, collected from two brackish lakes in Japan. Microbes Environ 35:ME20072. https://doi.org/10.1264/jsme2.ME20072
Yuan Z, Druzhinina IS, Labbé J et al (2016) Specialized microbiome of a halophyte and its role in helping non-host plants to withstand salinity. Sci Rep 6:32467. https://doi.org/10.1038/srep32467
Acknowledgements
The authors thank Institut Technique de Développement de l’Agronomie Saharienne (ITDAS) for performing the pedological analysis.
Funding
This work was supported by the French-Algerian Hubert Curien Partnership Program (PHC) TASSILI (14MDU907).
Author information
Authors and Affiliations
Contributions
SB, NB, TH and YK: designed the research. SB: carried out sample processing. WA, TH and YK: provided the experimental materials. SB: performed the laboratory work with contribution from NB. MB and PO: conducted bioinformatic data analysis. SB: wrote the first draft. TH and YK: supervised this study. All authors read, reviewed and edited previous versions of the manuscript. All authors approved the submission.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no conflict of interest.
Additional information
Communicated by A. Oren.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Behairi, S., Baha, N., Barakat, M. et al. Bacterial diversity and community structure in the rhizosphere of the halophyte Halocnemum strobilaceum in an Algerian arid saline soil. Extremophiles 26, 18 (2022). https://doi.org/10.1007/s00792-022-01268-x
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00792-022-01268-x