Abstract
Main conclusion
SMAX/SMXL family genes were successfully identified and characterized in the chickpea and lentil and gene expression data revealed several genes associated with the modulation of plant branching and powerful targets for use in transgenesis and genome editing.
Abstract
Strigolactones (SL) play essential roles in plant growth, rooting, development, and branching, and are associated with plant resilience to abiotic and biotic stress conditions. Likewise, karrikins (KAR) are “plant smoke-derived molecules” that act in a hormonal signaling pathway similar to SL playing an important role in seed germination and hairy root elongation. The SMAX/SMXL family genes are part of these two signaling pathways, in addition to some of these members acting in a still little known SL- and KAR-independent signaling pathway. To date, the identification and functional characterization of the SMAX/SMXL family genes has not been performed in the chickpea and lentil. In this study, nine SMAX/SMXL genes were systematically identified and characterized in the chickpea and lentil, and their expression profiles were explored under different unstressless or different stress conditions. After a comprehensive in silico characterization of the genes, promoters, proteins, and protein-protein interaction network, the expression profile for each gene was determined using a meta-analysis from the RNAseq datasets and complemented with real-time PCR analysis. The expression profiles of the SMAX/SMXL family genes were very dynamic in different chickpea and lentil organs, with some genes assuming a tissue-specific expression pattern. In addition, these genes were significantly modulated by different stress conditions, indicating that SMAX/SMXL genes, although working in three distinct signaling pathways, can act to modulate plant resilience. Most CaSMAX/SMXL and partner genes such as CaTiE1 and CaLAP1, have a positive correlation with the plant branching level, while most LcSMAX/SMXL genes were less correlated with the plant branching level. The SMXL6, SMXL7, SMXL8, TiE1, LAP1, BES1, and BRC1 genes were highlighted as powerful targets for use in transgenesis and genome editing aiming to develop chickpea and lentil cultivars with improved architecture. Therefore, this study presented a detailed characterization of the SMAX/SMXL genes in the chickpea and lentil, and provided new insights for further studies focused on each SMAX/SMXL gene.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Strigolactones (SL) are phytohormones that play essential roles in plant growth, rooting, development, and branching, and act to improve plant resilience (Yang et al. 2020; Li et al. 2022; Zhang et al. 2022). For the SL biosynthesis, all-trans-β-carotene are metabolized by the DWARF27 (D27) enzyme to produce 9′-cis-β-carotenoid, and the next steps of this pathway have the involvement of carotenoid cleavage dioxygenase (CCD) enzymes for synthesis of 9′-cis-β-10′-carotenal, carlactone, and 5-deoxylstrigol (Lopez-Obando et al. 2015; Wang et al. 2015; Bennett et al. 2016; Wallner et al. 2017; Sun et al. 2022). In the SL-dependent signaling pathway, the DWARF14 (D14) protein acts as an SL receptor, which binds and changes its molecular structure (Yao et al. 2016). D14 and SL complexes bind to D53/Suppressor of MAX2 1-Like (SMXL; formally named in Arabidopsis thaliana as AtSMXL6, AtSMXL7, and AtSMXL8) proteins in the nucleus, then recruit SCFMAX2 (SCF MORE AXILLARY GROWTH 2) protein to form D14-SL-SCFMAX2-D53/SMXLs complex (Bennett et al. 2016). SL promotes D53/SMXL ubiquitination, while the 26S proteasome specifically recognizes D53/SMXL proteins and directs to degradation, unlocking SL-dependent signal transduction and releasing BRANCHED 1 (BRC1) transcription factor (Zhou et al. 2013; Wang et al. 2015; Bennett et al. 2016). In this way, since D53/SMXL proteins are not degraded, SL-dependent signal transduction is inhibited while branching and tillering are promoted (Zhou et al. 2013; Zhang et al. 2022). Thus, D53/SMXL proteins are the final target proteins of this signaling pathway. Therefore, SL and D53/SMXL play a critical role in BRC1-mediated regulation of shoot branching and plant elongation (Zhao et al. 2015).
Likewise, karrikins (KAR) are butenolide molecules derived from plant smoke that act in a signaling pathway similar to the SL pathway (Bennett et al. 2016; Yang et al. 2019). In the KAR-dependent signaling pathway, KARRIKIN-INSUSCEPTIBLE2 (KAI2) protein acts as a KAR receptor (Villaécija-Aguilar et al. 2019). Subsequently, the KAR-KAI2 complex interacts with SCFMAX2 and SUPPRESSOR OF MAX2 1 (AtSMAX1/SMXL1), thus triggering ubiquitylation and targeting the AtSMAX1/SMXL1 and AtSMXL2 proteins for degradation (Carbonnel et al. 2020; Wang et al. 2020). Consequently, KAR-responsive genes, such as ACC synthase 7, which catalyzes ethylene biosynthesis, are transcriptionally up-regulated (Carbonnel et al. 2020). So, the KAR-dependent signaling pathway regulates seed germination and hairy root elongation (Wallner et al. 2017; Villaécija-Aguilar et al. 2019; Carbonnel et al. 2020). In addition, AtSMXL2 protein is also supposed to act in both SL- and KAR-dependent signaling pathways (Wang et al. 2020). Meanwhile, AtSMXL3, AtSMXL4, and AtSMXL5 proteins act in a yet unknown SL- and KAR-independent signaling pathway regulating phloem formation (Wallner et al. 2017). Given these previous studies, the biological importance of the SL- and KAR-dependent or -independent signaling pathways for plant development and resilience was determined in model plants. However, to date, little information about SMAX/SMXL family genes and their expression profile was generated from chickpea and lentil.
Chickpea (Cicer arietinum L.) and lentil (Lens culinaris Medik) are crops of outstanding importance for human food worldwide (Landi et al. 2021). The chickpea is a self-pollinated diploid, dicotyledonous, with a 738 Mb genome size organized in eight chromosomes (2n = 16) and up to 28,200 annotated genes (Varshney et al. 2013). In contrast, lentil is a self-pollinated diploid, dicotyledonous, with a 3.69 Gb genome size organized in seven chromosomes (2n = 14) and 58,243 annotated genes (Ramsay et al. 2023). Several germplasm banks with a high number of accessions, lines, and cultivars with high phenotypic variability are available for these two legumes. In particular, several cultivars of chickpea and lentil have a high number of branches and the absence of a typical dominant stem (Singh et al. 2019b; Silva-Perez et al. 2022). These intrinsic features related to plant architecture make it difficult to manage the chickpea and lentil crops in the field. For example, making mechanized harvesting more difficult and increasing lodging and susceptibility to pathogens (Tripathi et al. 2022). So, understanding the molecular mechanisms that orchestrate plant branching is essential for the development of chickpea and lentil cultivars with an improved architecture (Koul et al. 2022).
In this study, were systematically identified and characterized nine SMAX/SMXL family genes in the chickpea and lentil. The orthologous genes in the chickpea and lentil were identified using as reference AtSMAX/SMXL genes from A. thaliana. Subsequently, evolutionary relationships, features of sequences, the basic structure of genes, chromosomal localization of genes, cis-regulatory elements in promoter sequences, conserved motifs and domains in protein sequences, protein–protein interaction network, and three-dimensional (3D) structures of the SMAX/SMXL proteins were successfully performed and their biological implications were discussed. Expression profiles of the SMAX/SMXL genes in different organs of chickpea and lentil unstressed plants and under abiotic and biotic stress conditions were performed using a meta-analysis approach from RNAseq datasets. Finally, expression profiles of all identified SMAX/SMXL genes in the chickpea and lentil contrasting cultivars, such as little branched and highly branched cultivars, were determined in axillary and apical buds using real-time PCR assays. These collective data describe the sequence features and expression profile of each SMAX/SMXL gene and reveal the key players involved in the branching of chickpea and lentil. Therefore, our results provide a solid basis for further functional studies in the chickpea and lentil focused on each SMAX/SMXL gene. Furthermore, these data provide powerful genes for use in both transgenesis and genome editing to improve the architecture of these leguminous crops.
Materials and methods
Chickpea and lentil sequences and features
Chickpea genome sequences and features were retrieved of the CGIAR v1 assembly and Cicer arietinum v1 annotation dataset (Varshney et al. 2013) from the Phytozome v13 database (Goodstein et al. 2012). Meanwhile, lentil genome sequences and features were retrieved of the Lcu.2RBY assembly and Lens culinaris CDC Redberry genome v2 annotation dataset (Ramsay et al. 2023) from the Pulse Crop database (Humann et al. 2019). In addition, additional features were retrieved of the USask assembly and Lens culinaris v1 dataset from the Phytozome database. The A. thaliana genome sequences were retrieved from the TAIR10 dataset (Cheng et al. 2017).
Sequences analysis
Protein subcellular localization was predicted using LOCALIZER software (Sperschneider et al. 2017). Conserved domains in gene and protein sequences were identified using the PFAM database (El-Gebali et al. 2019), CD database (Marchler-Bauer et al. 2015), and InterPro Scan (Blum et al. 2021). Sequences were aligned using MUSCLE software (Edgar 2004) and curated by the Gblocks model, while evolutionary analyses were performed with Phylogeny.fr web service using maximum likelihood estimation (MLE) method with aLRT SH-like branch support and GTR (for nucleotide sequences) and WAG (for amino acid sequences) substitution models (Dereeper et al. 2008). SMAX/SMXL gene structures were displayed by the Gene Structure Display web server (Hu et al. 2014), while an unrooted evolutionary tree was inferred by the Neighbor-Joining (NJ) method (Saitou and Nei 1987) with 5000 bootstrap replicates using MEGA11 software (Tamura et al. 2021). The chromosomal localization of the SMAX/SMXL genes on chickpea and lentil genomes was generated using the MapGene2Chrom program (Jiangtao et al. 2015). Sequences of 2000 nucleotides upstream of the start codon were retrieved from the Phytozome database and submitted in the PlantCARE program to predict the cis-regulatory elements (Lescot et al. 2002). The conserved motifs in protein sequences were identified with the MEME Suite web server (Bailey et al. 2015). Protein–protein interaction network among SMAX/SMXL with partner proteins was predicted by the STRING database using the Cicer arietinum NCBI:txid3827 dataset as a reference (Szklarczyk et al. 2020).
Protein 3D structures
SMAX/SMXL protein sequences were processed by the FASTA program from the EMBL-EBI webpage (https://www.ebi.ac.uk/Tools/sss/fasta/) for search proteins that most resemble (best score) with our query sequences. This tool performed a local heuristic search by sequence similarity from a protein or nucleotide databases for a query protein of the same type (Madeira et al. 2019, 2022). EMBL-EBI webpage produced the folded protein structure in HTML color-coded by the predicted local distance difference test (plDDT) and generated a PAE plot. The results refer to the “best model fit”, which represents 3D structures according to the plDDT, using a scale that goes from 0 to 100%. This parameter indicates an estimate of how the predicted structure agrees with an experimentally determined structure (Tunyasuvunakool et al. 2021). For pair-to-pair comparison, 3D structures were processed by the MODELLER v10.4 program (Webb and Sali 2016). The previous selection was based on root mean square deviation (RMSD) set to a maximum limit of 10 Angstroms (Å).
Meta-analysis from RNAseq datasets
For tissue-specific expression in the chickpea, RNAseq datasets used in the meta-analysis were generated as reported by Jain et al. (2022) from Cicer arietinum cultivar ICC 4958 growth under room and field conditions. Thirty-two tissue samples representing different organs and developmental stages in at least three independent biological replicates were collected and analyzed. The expression level of each gene was FPKM normalized. In addition, for lentil tissue-specific expression, RNAseq datasets used in the meta-analysis were generated from Lens culinaris cultivar Cassab grown in a glasshouse at 22 °C with a 16 h photoperiod as described by Sudheesh et al. (2016). Different tissue samples were harvested from four-week-old plants using three biological replicates. Tissue-specific RNAseq datasets were normalized by log-transformed counts using the 75th percentile method. On the other hand, the RNAseq datasets used for meta-analysis from chickpea plants under abiotic and biotic stress were: (i) root tissue under drought stress experiment I and shoot tissue under drought stress (Mashaki et al. 2018); (ii) root tissue under salinity stress experiment I and root tissue under drought stress experiment II (Garg et al. 2016); (iii) before-flowering root tissue under heat stress, before-flowering leaf tissue under heat stress, after-flowering root tissue under heat stress, and after-flowering leaf tissue under heat stress (Kudapa et al. 2023); (iv) flower tissue under salinity stress (Kaashyap et al. 2022); (v) root tissue under salinity stress experiment II and shoot tissue under salinity stress (Kumar et al. 2021b); and (vi) root tissue under drought stress experiment III (Kumar et al. 2019). In contrast, RNAseq datasets used from lentil were: (i) seedling tissue infected by ascochyta blight from resistant cultivar and seedling tissue infected by ascochyta blight disease from susceptible cultivar (Khorramdelazad et al. 2018); (ii) seedling tissue under heat stress from tolerant cultivar and seedling tissue under heat stress from susceptible cultivar (Singh et al. 2019a); (iii) seedling tissue under heat stress (Sohrabi et al. 2022); (iv) seedling tissue infected by dry root rot disease (Mishra et al. 2021); (v) root tissue under alkalinity stress from tolerant cultivar and root tissue under alkalinity stress from susceptible cultivar (Singh et al. 2022a); (vi) root and leaf tissues under drought stress (Morgil et al. 2019); (vii) leaf tissue under drought stress from tolerant cultivar, leaf tissue under drought stress from susceptible cultivar, leaf tissue under heat stress experiment I from tolerant cultivar, leaf tissue under heat stress experiment I from susceptible cultivar, leaf tissue under salinity stress from tolerant cultivar, leaf tissue under salinity stress from susceptible cultivar, leaf tissue under alkalinity stress from tolerant cultivar, and leaf tissue under alkalinity stress from susceptible cultivar (Singh et al. 2022b); (viii) root tissue under salinity stress from tolerant cultivar, shoot tissue under salinity stress from tolerant cultivar, root tissue under salinity stress from susceptible cultivar, and shoot tissue under salinity stress from susceptible cultivar (Singh et al. 2021); and (ix) leaf tissue under heat stress experiment II from tolerant cultivar and leaf tissue under heat stress experiment II from susceptible cultivar (Kumar et al. 2021a). Heatmaps were generated by the SRplot web server (https://www.bioinformatics.com.cn/srplot) using log2 fold change values (stress treatment/control treatment).
Contrasting cultivars and plant materials
The chickpea cultivars Blanco lechoso and FLIP07-318C, and lentil cultivars Castellana and Campisi were previously selected among several other cultivars under greenhouse conditions as being the most contrasting in terms of plant branching (data not shown). The phenotypic analysis to define these contrasting cultivars was carried out determining the number of branches per plant at certain times after seed germination. In order to characterize the branching of these four selected cultivars, at least 15 plants (three replicates with five plants each) of each cultivar were evaluated, and branches were counted at 25- (stage I) and 40-day-old (stage I) plants. For this, seeds were superficially sterilized with 1.5% sodium hypochlorite solution, washed abundantly with distilled water, soaked for 5 min in distilled water, and germinated in Petri dishes containing humid filter paper for three days at room temperature. Germinated seeds with a 1–2 cm radicle were transferred to pots containing commercial substrate and kept in a greenhouse at room temperature.
RNA and gene expression
Axillary and apical buds were collected from 20-day-old plants. Total RNA was isolated with GenUP™ Total RNA kit (Biotechrabbit, Volmerstraße, Berlin, Germany) and RNA integrity was checked in agarose electrophoresis. RNA samples were treated with RNase-free RQ1 DNase I (Promega) and used for cDNA synthesis using oligo-(dT)20 primer and SuperScript III RT mix (Life Technologies, Carlsbad, CA, USA). The cDNA samples were diluted 1:10 (v:v) and real-time PCR assays were performed in QuantStudio 7 Flex Real-Time PCR system (Applied Biosystems, Waltham, MA, USA). The PCR mix consisted of 3 µL cDNA, 0.1 µM gene-specific primers (Suppl. Table S1), and SYBR Green PCR Master mix (Applied Biosystems). Relative expression was calculated with the 2^-∆Ct formula using CaCAC (Reddy et al. 2016) and LcTUB (Sinha et al. 2019) as reference genes for normalization (Suppl. Table S1). The CaG6PD, CaTIP41, LcRPL2, and LcRBC1 reference genes were also tested with a reduced number of samples, but CaCAC and LcTUB were considered more stable in our samples. Three biological replicates for each treatment and ten plants for each biological replicate were used. All cDNA samples were carried out in technical triplicate reactions. Target-specific amplification was confirmed by the occurrence of a single peak in the melting curve. Expression data were statistically evaluated using the SASM-Agri software (Canteri et al. 2001) while heatmaps were generated by the SRplot web server.
Results
Basic features of the SMAX/SMXL family members identified in the chickpea and lentil
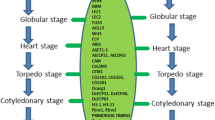
To identify the candidates SMAX/SMXL family genes in the chickpea and lentil genomes were used as reference the coding (CDS) and amino acid sequences of the orthologous AtSMAX/SMXL genes previously identified in A. thaliana. The biological importance of the AtSMAX/SMXL proteins in the SL- and KAR-dependent or SL- and KAR-independent signaling pathways for A. thaliana was previously proposed by Soundappan et al. (2015), Carbonnel et al. (2020), Villaécija-Aguilar et al. (2019), and Wallner et al. (2017), and used in this study as information support for chickpea and lentil (Fig. 1a). The in silico analyses allowed identify nine genes in the chickpea (CaSMAX1/SMXL1 to CaSMXL9) and other nine genes in the lentil (LcSMAX1/SMXL1 to LcSMXL9) genomes (Tables 1 and 2). Phylogenetic relationships from nucleotide and amino acid sequences were used to define the orthologous and their corresponding genes in the chickpea and lentil (Fig. 1b and c). In particular, CaSMAX/SMXL proteins showed predicted subcellular localization at chloroplast or nucleus, predominantly with PF02861, PF07724, IPR023150, IPR027417, IPR004176, IPR003959, and KOG1051 as main domains of the PFAM, InterPro, and KOG, respectively. The CaSMAX/SMXL coding sequences ranged between 1866 to 3252 nucleotides in length, and deduced protein sequences ranged from 622 to 1084 amino acids in length. In addition, molecular weight (Mw) ranged between 70.4 to 120.4 kDa, and isoelectric point (pI) ranged from 5.8 to 8.2 (Table 1). In contrast, LcSMAX/SMXL proteins showed predicted subcellular localization in the chloroplast, mitochondria, and nucleus, predominantly with PF02861, PF07724, IPR023150, IPR027417, IPR004176, IPR003959, and KOG1051 as main domains of the PFAM, InterPro, and KOG, respectively. The LcSMAX/SMXL coding sequences ranged from 1716 to 3246 nucleotides in length, and deduced protein sequences ranged from 572 to 1082 amino acids in length. In addition, molecular weight ranged from 64 to 120.5 kDa, while isoelectric point ranged from 5.7 to 7.5 (Table 2). The detailed features concerning the SMAX/SMXL chickpea and lentil genes, such as gene identifiers, orthologous gene in Arabidopsis, subcellular localization, conserved domains in protein sequences, chromosomal localizations of these genes, coding and amino acid sequences length, protein molecular weight, and isoelectric point are systematically listed in Tables 1 and 2. Therefore, these data showed that the SMAX/SMXL family genes of chickpea and lentil have several conserved features among them, indicating that they can act redundantly in some functions, while other features are specific to some members, also suggesting role specificity for these members.
Identification and characterization of the SMAX/SMXL family genes in the chickpea and lentil using previously characterized A. thaliana genes as reference. a Overview of involvement of the AtSMXL2, AtSMXL6, AtSMXL7, and AtSMXL8 proteins in the strigolactones (SL) signaling pathway of Arabidopsis. Role of the AtSMAX1/SMXL1 and AtSMXL2 proteins in the karrikin (KAR) signaling pathway of Arabidopsis. Role of the AtSMXL3, AtSMXL4, and AtSMXL5 proteins in the SL- and KAR-independent signaling pathway of Arabidopsis, a module not yet well characterized. AtTiE1, AtLAP1, AtBES1, and AtBRC1 proteins are also involved in the regulation of Arabidopsis branching. These three signaling pathways mediated by SMAX/SMXL proteins were previously proposed by Soundappan et al. (2015), Carbonnel et al. (2020), Villaécija-Aguilar et al. (2019), and Wallner et al. (2017). b and c Identification of the SMAX/SMXL genes in chickpea and lentil genomes using as reference SMAX/SMXL orthologous genes of Arabidopsis. Unrooted evolutionary trees were generated from nucleotide and amino acid sequences using the MLE method. d Basic structure of the SMAX/SMXL genes. The unrooted evolutionary tree was inferred using the NJ method
Phylogenetic relationships among the SMAX/SMXL family genes identified in different species
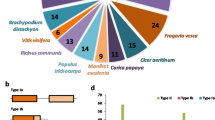
The evolutionary relationship among SMAX/SMXL family genes of chickpea and lentil in relation to orthologous genes in A. thaliana and Malus domestica were determined from unrooted phylogenetic trees constructed using 18 candidates SMAX/SMXL gene sequences identified in the chickpea and lentil, eight SMXL gene sequences identified in Arabidopsis, and ten SMXL gene sequences identified in M. domestica (Fig. 1a and b; Suppl. Fig. S1). Evolutionary relationships allowed identify the orthologous genes between chickpea, lentil, and A. thaliana, as well as paralogous genes within the same legume species (Fig. 1a and b). Three major groups were identified with at least 92% bootstrap support, with SMXL6, SMXL7, and SMXL8 genes of chickpea and lentil separately grouped in group I clustered with AtSMXL6 to AtSMXL8. Meanwhile, the SMAX1/SMXL1 genes were grouped in group II clustered with AtSMAX1 and AtSMXL2. Finally, the SMXL2, SMXL3, SMXL4, SMXL5, and SMXL9 genes of chickpea and lentil were separately grouped in group III clustered with AtSMXL3 to AtSMXL5 (Fig. 1a and b). This same organization in three main groups was maintained when including in the phylogenetic analysis the SMAX/SMXL sequences of M. domestica (Suppl. Fig. S1). Therefore, the phylogenetic relationship analysis showed that the SMAX/SMXL family genes of different plant species clustered forming three groups closely corresponding to their biological role in the three signaling pathways (SL-, KAR-dependent, and SL- and KAR-independent).
Structure and chromosomal localization of the SMAX/SMXL genes
The structural organization of the SMAX/SMXL family genes based on 5′-UTR, introns, exons, and 3′-UTR sequences was successfully determined (Fig. 1d). The number of introns/exons was very similar between these genes, ranging from three to four exons in each gene sequence for both chickpea and lentil. In addition, the length of the exon sequences was also similar between them, while the length of intron sequences was more variable, especially the SMXL3 and SMXL9 genes which included two short introns compared to SMXL6 and SMXL7 which showed two larger introns. The chromosomal localization of the SMAX/SMXL genes was also successfully determined in the chickpea (Fig. 2a) and lentil (Fig. 2b). The mapping data indicated that CaSMAX/SMXL genes were located on five out of eight chromosomes, one on chromosome 1 (CaSMXL4), one on chromosome 2 (CaSMXL7), three on chromosome 4 (CaSMXL2, CaSMXL3, and CaSMXL9), two on chromosome 5 (CaSMXL6 and CaSMXL8), and two on chromosome 7 (CaSMAX1/SMXL1 and CaSMXL5) (Fig. 2a). For lentil genes, the mapping data indicated that LcSMAX/SMXL genes were located on five out of seven chromosomes, three on chromosome 1 (LcSMXL2, LcSMXL3, and LcSMXL9), one on chromosome 2 (LcSMXL4), two on chromosome 3 (LcSMXL6 and LcSMXL8), one on chromosome 5 (LcSMXL7), and two on chromosome 7 (LcSMAX1/SMXL1 and LcSMXL5) (Fig. 2b). Based on this evidence, a similar distribution of these genes was observed between chickpea and lentil. In addition, these data may hypothesize two segmental duplication events (chromosome four and seven of chickpea and chromosome one and seven of lentil) for both plant species studied here, which eventually allowed an expansion of this SMAX/SMXL family.
Localizations of the SMAX/SMXL genes on chromosomes of a chickpea and b lentil. Positions are based on megabases (Mb). c Cis-regulatory elements in promoter sequences (2000 nucleotides upstream of the start codon) of the SMAX/SMXL genes. Number of each cis-regulatory element is shown in the heatmap. ABA abscisic acid, MeJA methyl jasmonate, MYB MYB transcription factor binding domain, MYC MYC transcription factor binding domain, SA salicylic acid
Cis-regulatory elements in SMAX/SMXL promoter sequences
Fourteen main cis-regulatory elements present in each promoter sequence for each of the 18 SMAX/SMXL genes were successfully identified (Fig. 2c). The four most prevalent cis-regulatory elements in these promoter sequences were light-responsive that ranged from three (LcSMXL9) to 17 (LcSMXL6, LcSMXL7, and LcSMXL8), MYB-related ranged from two (LcSMXL6) to 12 (LcSMAX1/SMXL1), ethylene-responsive ranged from one (LcSMXL7) to nine (CaSMAX1/SMXL1), and MYC-related ranged from zero (LcSMXL8) to eight (CaSMXL6). In particular, for promoter sequences of SMXL9 genes, three cis-regulatory elements associated with meristem-specific expression were identified, implying that these genes may develop an important role in the early stage of plant tissue development. Similarly, promoter sequences of CaSMAX1/SMXL1 and LcSMXL2 genes showed twelve and eight ABA-responsive cis-regulatory elements, respectively. In addition, several SMAX/SMXL genes showed cis-regulatory elements responsive to gibberellin, auxin, salicylic acid, and methyl jasmonate indicating that they can be modulated by other hormones (Fig. 2c). Furthermore, although these genes showed close evolutionary conservation (Fig. 1b and c), a considerable difference in the type and number of cis-regulatory elements were observed (Fig. 2c), indicating a possible variable transcriptional modulation among each gene of this family and also between chickpea and lentil. Therefore, these data suggested that SMAX/SMXL family genes may have their expression modulated by different hormones or influenced by abiotic and biotic stress conditions, and activated primarily in a tissue or plant stage-specific manner. Thus, these data suggested that SMAX/SMXL family genes of chickpea and lentil can play an important role in the modulation of plant growth and development and can be modulated by different abiotic and biotic stresses.
Conserved motifs and protein–protein interaction network of the SMAX/SMXL proteins
SMAX/SMXL protein sequences of chickpea and lentil were in silico evaluated to identify the top ten conserved motifs (Fig. 3a). The number and organization of these motifs were conserved among each gene in chickpea and lentil, and with considerable similarity to the orthologous in A. thaliana. In particular, some motifs or their position in the protein sequence were specific for certain SMAX/SMXL proteins, indicating that some of these proteins can assume particular characteristics to perform their biological function. Subsequently, a meta-analysis of the protein–protein interaction network was conducted with the STRING database for both chickpea and lentil (Fig. 3b). These data clearly showed that all SMAX/SMXL family members are highly interconnected in a major group composed of 14 proteins. Particularly in this group, D14L/KAI2, F-box MAX2, and heat shock protein 70 (HSP70, also named hypoxia up-regulated 1 protein, HYOU1) were shown to be central hub proteins together with SMAX/SMXL proteins of both chickpea and lentil. The fourth element of this major group, F-box SKIP25-like, showed to be directly related to the SMAX1/SMXL1 proteins and interconnected with D14L/KAI2 and F-box MAX2, potentially both acting in the core of the KAR-dependent signaling pathway. Therefore, these data showed that SMAX/SMXL proteins of chickpea and lentil can play quite conserved functions and, although they act in three different signaling pathways, they form protein–protein interaction networks with the same major hub proteins. The HSP70 was shown to be the main hub protein that interconnects with all SMAX/SMXL proteins.
Conserved motifs and protein–protein interaction network among SMAX/SMXL with partner proteins. a Top 10 conserved motifs in SMAX/SMXL protein sequences of chickpea and lentil. The unrooted evolutionary tree was generated from amino acid sequences using the MLE method. b Protein–protein interaction network predicted by the STRING database using Cicer arietinum dataset as reference. HSP70: Ca_07617 and Lcu.2RBY.3g044220.1; D14L/KAI2: Ca_09326, Ca_02196, Lcu.2RBY.5g006180.1, and Lcu.2RBY.L022510.1; F-box MAX2: Ca_19880 and Lcu.2RBY.4g047720.1; F-box SKIP25-like: Ca_10634 and Lcu.2RBY.5g013390.1. Known interactions are shown in light blue line: from curated databases, and pink line: experimentally determined. Predicted interactions are shown in dark green line: gene neighborhood, red line: gene fusions, and dark yellow: gene co-occurrence. Other protein–protein associations are shown in light green line: text-mining, black line: co-expression, and light blue line: protein homology
3D structure of the SMAX/SMXL proteins
The 3D structures of the SMAX/SMXL proteins from chickpea and lentil were modeled and compared to each other to verify potential structural similarity. Consistently, all 18 SMAX/SMXL proteins characterized in this study had their 3D structure successfully determined (Fig. 4a–r). In addition, structural comparisons between SMAX/SMXL proteins from chickpea and lentil were performed and when similarity was greater than 66% they were considered for further analysis. The pair-to-pair structure comparison revealed 37 comparisons with a large overlapping region with > 66% homology of residues when using RMSD values maximum of 10 Å (Suppl. Table S2). These data showed that some SMAX/SMXL proteins from chickpea and lentil have considerable structural similarities and differences from each other, both between plant species or among proteins of the same plant species. Therefore, the SMAX/SMXL family has highly conserved proteins within the same species, as well as conserved proteins between different plant species, suggesting high conservation of the SL- and KAR-dependent or -independent signaling pathways in the chickpea and lentil.
3D structure of the CaSMAX/SMXL and LcSMAX/SMXL proteins. Structure of a CaSMAX1/SMXL1, b CaSMXL2, c CaSMXL3, d CaSMXL4, e CaSMXL5, f CaSMXL6, g CaSMXL7, h CaSMXL8, i CaSMXL9, j LcSMAX/SMXL1, k LcSMXL2, l LcSMXL3, m LcSMXL4, n LcSMXL5, o LcSMXL6, p LcSMXL7, q LcSMXL8, and r LcSMXL9. Regions of different degrees of confidence are expressed with different colors according to predicted local distance difference test (plDDT) value, going from dark blue (very high confidence degree plDDT > 90%), cyan (high confident degree plDDT < 90%), yellow (relative confidence degree plDDT < 70%), and red (low confidence degree plDDT < 50%)
Dynamic expression of the SMAX/SMXL genes in different organs and under stress conditions
The expression profile of each of these SMAX/SMXL genes in different organs and in chickpea and lentil plants under different abiotic to biotic stress conditions was performed using a meta-analysis approach from RNAseq datasets. In untressed chickpea plants, it was observed that all SMAX/SMXL genes had their lowest expression level in the hairy root, endosperm, nodule, 30 days after pollination-seed, and androecium while the higher expression level observed was in the gynoecium (CaSMXL7 and CaSMXL4), pedicel and pod shell (CaSMXL8 and CaSMXL9, respectively), shoot apical meristem (CaSMXL3 and CaSMXL5), and in different flowers stages (CaSMAX1/SMXL1 and CaSMXL6) (Fig. 5a). In contrast, the highest expression level in lentil tissues observed was in immature seed and pod (LcSMXL3 and LcSMXL9), root (LcSMXL2, LcSMXL4, LcSMXL5, LcSMXL6, and LcSMXL8), stem (LcSMXL7), and flower and immature pod (LcSMAX1/SMXL1) (Fig. 5b).
Expression profiles of the SMAX/SMXL genes in different organs of chickpea and lentil, and plants under different abiotic and biotic stress conditions determined from a meta-analysis using RNAseq datasets. SMAX/SMXL gene expression levels in different organs of a chickpea and b lentil. Gene expression in chickpea plants under c drought, d salinity, e heat stress conditions. Gene expression in lentil plants under f drought, g salinity, and h heat stress conditions. Expression values in different organs of chickpea correspond to FPKM-normalized counts for each gene, while expression values in different organs of lentil correspond to log-transformed counts using the 75th percentile method. Expression values in plants under stress conditions correspond to Log2(fold-change) contrasting “treatment versus control”. The scale bar indicates the expression profile
Meanwhile, the meta-analysis also addressed the expression profiles of the CaSMAX/SMXL genes in chickpea plants under drought, salinity, and heat stress conditions (Fig. 5c–e). Overall, all CaSMAX/SMXL genes had their expression modulated by these different stress conditions. In particular, under drought conditions, all genes had their highest expression level in shoots, while CaSMXL6 and CaSMXL9 genes were also significantly more up-regulated in roots (Fig. 5c). In comparison, under salinity stress conditions, the CaSMAX/SMXL genes were up-regulated in the flower, root, and shoot, with some genes preferentially more up-regulated in certain tissues (Fig. 5d). Meanwhile, when under heat stress conditions, all CaSMAX/SMXL genes had their expression up-regulated in before-flowering and after-flowering leaves, except for CaSMAX1/SMXL1 and CaSMXL3 genes (Fig. 5e). Likewise, the meta-analysis also addressed the LcSMAX/SMXL gene expression level in lentil plants under drought, salinity, heat, alkalinity, and biotic stress conditions (Fig. 5f–h; Suppl. Fig. S2a and b). In addition, this analysis also included LcSMAX/SMXL gene expression in contrasting lentil cultivars (tolerant, resistant, and susceptible). Similar to that observed in the chickpea, the LcSMAX/SMXL genes were significantly up-regulated in lentil plants under different abiotic and biotic stress conditions and showed a positive correlation with plant tolerance level to both stress conditions. In particular, under drought, salinity, and alkalinity stress conditions, the greater up-regulation of the LcSMAX/SMXL genes observed was in roots of tolerant cultivars compared to the leaves of these same cultivars. Interestingly, LcSMAX/SMXL genes are also significantly up-regulated by biotic stress and mostly correlated positively with plant resistance level (Suppl. Fig. S2b). Therefore, these collective data showed that both SMAX/SMXL genes of chickpea and lentil were dynamically modulated by different abiotic and biotic stress conditions, with some of these genes taking a more tissue-specific expression, and with a positive correlation with plant tolerance or resistance level.
Expression of the SMAX/SMXL genes in contrasting cultivars in terms of branching
First, a screening was carried out with several cultivars of chickpea and lentil to identify the two most contrasting cultivars: little branched and highly branched. After phenotypic analysis based on the number of branches per plant, were selected the chickpea cultivars Blanco lechoso and FLIP07-318C, and lentil cultivars Castellana and Campisi as being little branched and highly branched, respectively (Table 3; Fig. 6a and b). During this phenotypic analysis carried out in the greenhouse, axillary and apical buds were sampled for further molecular analysis. Subsequently, the expression profiles of the SMAX/SMXL genes were performed by real-time PCR. In addition, BRC1 (transcription factor involved in plant branching for acting in the SL-dependent signaling pathway), TiE1 (TCP interactor containing EAR motif protein 1), LAP1 (LIKE-APETALA1), and BES1 (BRI1-EMS-SUPPRESSOR 1) genes, which encode proteins that are negative regulators of BRC1 protein, were also evaluated. For the chickpea, the highest expression level of these genes was observed in axillary buds of the highly branched cultivar (FLIP07-318C), except for CaBES1 and CaSMXL5 genes that showed higher expression in apical buds (Fig. 6c). These preliminary data showed that the expression profile of these genes in chickpea cultivars has a positive correlation with plant branching level. In the lentil, the expression profiles of these genes showed a lower correlation with plant branching level (Fig. 6d). In particular, the LcSMXL3, LcSMXL4, LcSMXL8, LcSMXL9, and LcLAP1 genes showed higher expression levels in apical buds of the highly branched cultivar (Campisi), while the other genes were more expressed in axillary buds of the little branched cultivar (Castellana). These data showed that each gene or group of SMAX/SMXL genes, despite having several conserved features, has a slightly different expression profile between chickpea and lentil cultivars. This expression profile was also dynamic at the tissue level (axillary and apical buds), collaborating with the dynamics of the three signaling pathways in which these genes are involved.
Contrasting cultivars and expression profiles of the SMAX/SMXL genes. Branching profile of the contrasting cultivars of a chickpea and b lentil. Heatmap representation displaying the expression patterns of the SMAX/SMXL genes in the c chickpea cultivars Blanco lechoso (little branched) and FLIP07-318C (highly branched), and d lentil cultivars Castellana (little branched) and Campisi (highly branched). Gene expression levels were explored by real-time PCR. The scale bar indicates the relative expression level
Overall, the real-time PCR data showed that CaSMAX1/SMXL1 (Fig. 7a), CaSMXL3 (Fig. 7c), CaSMXL4 (Fig. 7d), CaSMXL5 (Fig. 7e), CaSMXL6 (Fig. 7f), CaSMXL7 (Fig. 7g), CaSMXL8 (Fig. 7h), CaSMXL9 (Fig. 7i), CaTiE1 (Fig. 7k), and CaLAP1 (Fig. 7l) genes were more expressed with statistical significance in the highly branched cultivar, therefore, with a positive correlation with plant branching level. Meanwhile, CaSMXL2 and CaBES1 genes showed no significant difference in expression level between different tissues and contrasting cultivars (Fig. 7b and m). In contrast, the CaBRC1 gene showed lower expression with statistical significance in apical buds of the highly branched cultivar, a negative correlation with plant branching level (Fig. 7j). Meanwhile, the real-time PCR data from lentil showed that LcSMAX1/SMXL1 (Fig. 7n), LcSMXL2 (Fig. 7o), LcSMXL4 (Fig. 7q), LcSMXL5 (Fig. 7r), LcSMXL6 (Fig. 7s), LcSMXL8 (Fig. 7u), LcLAP1 (Fig. 7a1), and LcBES1 (Fig. 7a2) genes were more expressed in a tissue-specific manner, with no clear correlation with plant branching level. In contrast, the expression profiles of LcSMXL7 (Fig. 7t), LcBRC1 (Fig. 7y), and LcTiE1 (Fig. 7z) genes showed a negative correlation with plant branching level, while LcSMXL3 (Fig. 7p) and LcSMXL9 (Fig. 7x) genes showed a positive correlation with plant branching level. Therefore, these expression data revealed that most CaSMAX/SMXL and partner genes are positively correlated with branching level, except the CaBRC1 gene with a negative correlation. In comparison, the gene expression profiles in the lentil were mostly tissue-dependent, but LcSMXL3 and LcSMXL9 genes were positively correlated with the branching level. Furthermore, these collective data, in addition to providing new regulatory information for each gene, provide powerful targets such as SMXL6, SMXL7, SMXL8, TiE1, LAP1, BES1, and BRC1 genes for eventual use in transgenesis and genome editing for the development of chickpea and lentil cultivars with improved architecture.
Expression profile of the SMAX/SMXL genes measured by real-time PCR in axillary and apical buds of chickpea cultivars Blanco lechoso and FLIP07-318C and lentil cultivars Castellana and Campisi. Expression profiles of chickpea a CaSMAX1/SMXL1, b CaSMXL2, c CaSMXL3, d CaSMXL4, e CaSMXL5, f CaSMXL6, g CaSMXL7, h CaSMXL8, i CaSMXL9, j CaBRC1, k CaTiE1, l CaLAP1, and m CaBES1, and lentil n LcSMAX1/SMXL1, o LcSMXL1, p LcSMXL2, q LcSMXL3, r LcSMXL4, s LcSMXL5, t LcSMXL6, u LcSMXL7, v LcSMXL8, x LcSMXL9, y CaBRC1, z CaTiE1, a1 CaLAP1, and a2 CaBES1 genes. Gene expression values were calculated with the 2^-∆Ct formula and normalized with CaCAC and LcTUB as endogenous reference genes (Suppl. Table S1). Error bars represent confidence intervals corresponding to three biological replicates consisting of 10 plants each replicate (n = 10). Different letters on the bars indicate significant statistical differences according to Tukey’s test at a 95% significance level
Discussion
The increasing climate changes, eminent geopolitical conflicts, growth of the global population, and high demand for healthy food are major factors that are challenging agriculture around the world (Arif et al. 2021). Chickpea and lentil are important crops for the food security of several European and Asian countries (Landi et al. 2021; Karalija et al. 2022). The plant breeding of these legumes to improve agronomic traits associated with abiotic and biotic stress tolerance, seed yield, nutritional features, and plant architecture are important requirements to produce more at a lower cost (Basso et al. 2020, 2023; Asati et al. 2022). Particularly related to the architecture of chickpea and lentil plants, significant efforts are still needed to develop superior cultivars better adapted to mechanized planting and harvesting systems (Yang et al. 2021). Fortunately, for both chickpea and lentil there are currently available a huge amount of accessions, genotypes, and cultivars in germplasm banks worldwide that can be explored to develop these more adapted cultivars (Piergiovanni 2022; Basso et al. 2023). Although knowledge of the genetic basis associated with different agronomical traits has been explored in recent years, little is known about the molecular mechanism involved in plant branching and architecture of these two legume crops. The SL, together with other hormones, is one of the main regulators of the plant branching mechanism (Yang et al. 2020; Li et al. 2022; Zhang et al. 2022). In particular, the biological role of SL and KAR hormones in plant development and resilience to abiotic stresses has not yet been explored in detail in the chickpea and lentil. Therefore, improving knowledge about the SL and KAR signaling pathways can help to understand and develop biotechnological strategies related to plant architecture (Yang et al. 2019). In especial, SMAX/SMXL family genes are important players that act in the SL- and KAR-dependent and -independent signaling pathways (Wallner et al. 2017; Carbonnel et al. 2020; Wang et al. 2020; Li et al. 2022). Overall, SMAX/SMXL family genes are organized into three major functional groups, which correspond to the involvement of these members in the three signaling pathways described above. These three pathways act mainly on (i) shoot branching and elongation, (ii) seed germination and root elongation, and (iii) phloem formation (Soundappan et al. 2015; Wallner et al. 2017; Villaécija-Aguilar et al. 2019). These functional roles of the SMAX/SMXL proteins have been consistently studied in Arabidopsis. However, to date, SMAX/SMXL family genes have not yet been studied and explored in the chickpea and lentil.
In this study, were identified and characterized nine SMAX/SMXL family genes in the chickpea and lentil, and further analyses were performed focusing on the involvement of these genes in plant branching. The eight SMAX/SMXL family genes of Arabidopsis were used as a reference to successfully find the orthologous genes in chickpea and lentil genomes. In silico analyses from gene and protein sequences revealed that these members are highly conserved in the chickpea and lentil but also with some particular features. In addition, were observed the presence of the Clp-N and P-loop NTPase domains in all SMAX/SMXL proteins of chickpea and lentil, which are related to the nuclear localization and ubiquitination of these proteins, respectively (Liang et al. 2016; Khosla et al. 2020). These features indicated that the SMAX/SMXL genes of chickpea and lentil may range from redundant to very specific functions, similar to those observed with SMAX/SMXL family members of Arabidopsis (Carbonnel et al. 2020; Wang et al. 2020). Phylogenetic relationship data showed the organization of the SMAX/SMXL genes of chickpea and lentil into three major groups closely related to their putative biological function, as well as also observed with SMAX/SMXL genes of Arabidopsis (Soundappan et al. 2015). Similarly, other recent studies have also identified and characterized SMAX/SMXL family genes in different plant species, such as M. domestica (Li et al. 2018), soybean (Zhang et al. 2022), cotton (Jia et al. 2022), and Populus trichocarpa (Sun et al. 2023). In particular, in these studies were identified a variable number of SMAX/SMXL genes but not very different from those observed in the chickpea and lentil, except for the soybean with 31 members (Zhang et al. 2022). This higher number of SMAX/SMXL members in the soybean can be explained by the highly duplicated genome (Schmutz et al. 2010). Also, another important feature observed was the high conservation of these members in different plant species, thus suggesting the high importance of this protein family in green plants. Also, these data support the biological importance of SL- and KAR-dependent or -independent signaling pathways mediated by the SMAX/SMXL genes for seed germination, plant development, branching, and resilience. Our data also showed that segmental duplication can have occurred in the chickpea and lentil for some of these genes since some of them were highly associated with the same chromosome and relatively closely located. In the soybean, there was no tandem duplication event of the SMAX/SMXL genes, but 27 segmental duplication events related to 31 SMAX/SMXL genes were detected (Zhang et al. 2022). In M. domestica, duplication events of the SMAX/SMXL genes were also suggested as responsible for the expansion of this family (Li et al. 2018).
The promoter sequences of the SMAX/SMXL genes showed a considerable number of cis-regulatory elements associated mainly with responses to light, hormones, and defense response, while some were associated with tissue-specific expression in the chickpea and lentil. These data suggested that the expression profile of the SMAX/SMXL genes can be dynamically influenced in the chickpea and lentil by the plant development stage, in a tissue-specific manner, and under abiotic and biotic stress conditions. In the soybean, despite the number of GmSMAX/SMXL genes being approximately three times greater, the number and widespread distribution of these cis-regulatory elements were similar to those observed in the chickpea and lentil (Zhang et al. 2022). In the P. trichocarpa, light-responsive and environmental stress-related cis-regulatory elements were also the most abundant in their promoter sequences (Sun et al. 2023). Subsequently, the protein–protein interaction network of the SMAX/SMXL proteins of chickpea and lentil was evidenced, composed of a major group of 14 proteins, which includes five partner proteins. In particular, an HSP70/HYOU1 protein was identified as a major hub to be interconnected with all SMAX/SMXL proteins of chickpea and lentil. The HSP70/HYOU1 proved to be highly conserved in several plant species, which contain a cl17037 domain (nucleotide-binding domain of the sugar kinase/HSP70/actin superfamily). The Arabidopsis HSP70/HYOU1 protein is considered a chaperone complex protein of the endoplasmic reticulum involved in the cellular response to hypoxia and negative regulation of hypoxia-induced intrinsic apoptotic signaling pathway (Behnke et al. 2015). However, the functional relationship between HSP70/HYOU1 proteins and the SL- and KAR-dependent signaling pathway has not yet been elucidated. In addition, SMAX1/SMXL1 proteins besides showing their interaction network with D14L/KAI2, HSP70/HYOU1, and F-box MAX2, also showed an interaction network with a F-box SKIP25-like, and a KAR-related protein involved in protein ubiquitination (Nelson et al. 2010; Sepulveda et al. 2022). Therefore, these data confirmed the high relationship and conservation of functions between the SMAX/SMXL proteins of chickpea and lentil. In agreement, the 3D structure data of the SMAX/SMXL proteins revealed that these members have similar structures in the chickpea and lentil, but the small differences in the structure and composition can be important to play the different roles in SL and KAR signaling pathways. This similar pattern of high structural conservation among SMAX/SMXL proteins was also observed in P. trichocarpa (Sun et al. 2023).
In order to explore the expression pattern of these genes in different tissues of unstressed and stressed chickpea and lentil plants, was performed meta-analysis from RNAseq datasets. The meta-analysis data showed that SMAX/SMXL family genes are highly expressed at very dynamic levels in all plant tissues, indicating that they may play a remarkable role in plant growth and development. A similar tissue-specific expression pattern was observed with the SMAX/SMXL family genes of M. domestica and P. trichocarpa (Li et al. 2018; Sun et al. 2023). Likewise, the expression profiles of these SMAX/SMXL genes were also dynamic and significantly modulated in chickpea and lentil plants under different abiotic and biotic stress conditions. Similar results of SMAX/SMXL gene expression were observed in M. domestica under polyethylene glycol, abscisic acid, salinity, and cold stress treatments (Li et al. 2018). In the cotton, most SMAX/SMXL family genes were significantly up-regulated or down-regulated by at least one cold, heat, drought, or salinity stress condition (Jia et al. 2022). In addition, the expression profile of SMAX/SMXL family genes was also evaluated by real-time PCR in contrasting cultivars of chickpea and lentil in terms of branching. Especially regarding SMAX1/SMXL1, SMXL6, SMXL7, and SMXL8 genes of chickpea, which are directly involved in the SL signaling pathway and act in the plant branching, showed a positive correlation with the plant branching level. Meanwhile, these same genes of lentil had an expression not clearly correlated with plant branching level. In contrast, the BRC1 genes showed a negative correlation with plant branching levels of both chickpea and lentil cultivars. Previous studies showed that a higher expression level of BRC1 genes is associated with reduced plant branching (Aguilar-Martínez et al. 2007). The expression profiles of the TiE1, BES1, and LAP1 genes also revealed, at least in the chickpea, a positive correlation with plant branching level. In particular, these three proteins act as inhibitors of BRC1 protein accumulation, directly impacting plant branching (Diao et al. 2019; Hu et al. 2020; Maurya et al. 2020a). In this way, the SMXL6, SMXL7, SMXL8, TiE1, LAP1, and BES1 genes are powerful targets for use in genome editing aiming a gene knockout and, consequently, developing chickpea and lentil cultivars with an improved architecture. In this context, the triple knockdown of the AtSMXL6, AtSMXL7, and AtSMXL8 genes in Arabidopsis resulted in improved drought tolerance and reduced plant branching (Yang et al. 2020). Similarly, the knockdown of the AtTiE1 gene in Arabidopsis also resulted in reduced plant branching (Diao et al. 2019). In the same sense, due to the fact that LAP1 and BES1 proteins inhibit BRC1, the knockout of these two genes can also result in a phenotype of reduced plant branching (Hu et al. 2020; Maurya et al. 2020a). Similarly, BRC1 genes of chickpea and lentil are powerful targets for use in transgenesis aiming a gene overexpression and, consequently, developing chickpea and lentil cultivars with an improved architecture. The knockdown of the AtBRC1 gene in Arabidopsis resulted in plants with highly branched, suggesting that overexpression of BRC1 genes in the chickpea and lentil can result in reduced plant branching (Aguilar-Martínez et al. 2007; Maurya et al. 2020b). Therefore, these collective data provided new evidence to be exploited for the regulation of SMAX/SMXL and partner genes by transgenesis or genome editing, as well as by traditional plant breeding, to achieve genetic improvements associated with plant architecture (Basso et al. 2019).
Conclusion
In this study, a comprehensive identification and characterization of the SMAX/SMXL genes from chickpea and lentil at the level of gene, protein, promotor sequence, and gene expression was systematically provided. Gene expression data revealed the expression profiles of all these genes in different organs of plants unstressed and plants under different stress conditions (abiotic or biotic), as well as in contrasting cultivars in terms of branching. These results showed that SMAX/SMXL genes are dynamically modulated both in the chickpea and lentil, with a positive correlation with the branching level of chickpea cultivars and a tissue-specific expression manner for the lentil. These collective data also highlighted the involvement of SMAX/SMXL genes in SL- and KAR-dependent and -independent signaling pathways. In addition, revealed some genes which can be interesting targets for the development of biotechnological tools based on transgenesis or genome editing to reduce plant branching and improve plant architecture. Furthermore, this study will help to understand better the biological role of SMAX/SMXL genes in branching and resilience to stresses of chickpea and lentil.
Data availability statement
All data generated were available in the manuscript as figures, tables, and supplementary tables. All gene sequences and their respective ID numbers were described in the materials and methods, and in which public and open-access databases were retrieved. All datasets used in the meta-analysis were retrieved from previously published articles and were properly cited in this work. It is also worth noting that these datasets used already had the differential expression information analyzed and normalized, so we only work with the final data.
Abbreviations
- SMAX1:
-
SUPPRESSOR OF MAX2 1
- SMXL:
-
Suppressor of MAX2 1-Like
- TiE1:
-
TCP interactor containing EAR motif protein 1
- LAP1:
-
LIKE-APETALA1
- BES1:
-
BRI1-EMS-SUPPRESSOR 1
References
Aguilar-Martínez JA, Poza-Carrión C, Cubas P (2007) Arabidopsis BRANCHED1 acts as an integrator of branching signals within axillary buds. Plant Cell 19:458–472. https://doi.org/10.1105/tpc.106.048934
Arif A, Parveen N, Waheed MQ, Atif RM, Waqar I, Shah TM (2021) A comparative study for assessing the drought-tolerance of chickpea under varying natural growth environments. Front Plant Sci 11:607869. https://doi.org/10.3389/fpls.2020.607869
Asati R, Tripathi MK, Tiwari S, Yadav RK, Tripathi N (2022) Molecular breeding and drought tolerance in chickpea. Life 12:1846. https://doi.org/10.3390/life12111846
Bailey TL, Johnson J, Grant CE, Noble WS (2015) The MEME suite. Nucleic Acids Res 43(W1):W39-49. https://doi.org/10.1093/nar/gkp335
Basso MF, Ferreira PCG, Kobayashi AK, Harmon FG, Nepomuceno AL, Molinari HBC, Grossi-de-Sa MF (2019) MicroRNAs and new biotechnological tools for its modulation and improving stress tolerance in plants. Plant Biotechnol J 17:1482–1500. https://doi.org/10.1111/pbi.13116
Basso MF, Arraes FBM, Grossi-de-Sa M, Moreira VJV, Alves-Ferreira M, Grossi-de-Sa MF (2020) Insights into genetic and molecular elements for transgenic crop development. Front Plant Sci 15(11):509. https://doi.org/10.3389/fpls.2020.00509
Basso M, Contaldi F, Lo Celso F, Karalija E, Paz-Carrasco L, Barone G, Ferrante A, Martinelli F (2023) Expression profile of the NCED/CCD genes in chickpea and lentil during abiotic stress reveals a positive correlation with increased plant tolerance. Plant Sci 336:111817. https://doi.org/10.1016/j.plantsci.2023.111817
Behnke J, Feige MJ, Hendershot LM (2015) BiP and its nucleotide exchange factors Grp170 and Sil1: mechanisms of action and biological functions. J Mol Biol 427(7):1589–1608. https://doi.org/10.1016/j.jmb.2015.02.011
Bennett T, Liang Y, Seale M, Ward S, Müller D, Leyser O (2016) Strigolactone regulates shoot development through a core signalling pathway. Biol Open 5:1806–1820. https://doi.org/10.1242/bio.021402
Blum M, Chang HY, Chuguransky S, Grego T, Kandasaamy S, Mitchell A, Nuka G, Paysan-Lafosse T, Qureshi M, Raj S, Richardson L, Salazar GA, Williams L, Bork P, Bridge A, Gough J, Haft DH, Letunic I, Marchler-Bauer A, Mi H, Natale DA, Necci M, Orengo CA, Pandurangan AP, Rivoire C, Sigrist CJA, Sillitoe I, Thanki N, Thomas PD, Tosatto SCE, Wu CH, Bateman A, Finn RD (2021) The InterPro protein families and domains database: 20 years on. Nucleic Acids Res 49(D1):D344–D354. https://doi.org/10.1093/nar/gkaa977
Canteri MG, Althaus RA, Filho JV, Giglioti ÉA, Godoy CV (2001) SASM-Agri: Sistema para análise e separação de médias em experimentos agrícolas pelos métodos Scott-Knott, Tukey e Duncan. Revista Brasileira De Agrocomputação 1(2):18–24
Carbonnel S, Das D, Varshney K, Kolodziej MC, Villaécija-Aguilar JA, Gutjahr C (2020) The karrikin signaling regulator SMAX1 controls Lotus japonicus root and root hair development by suppressing ethylene biosynthesis. PNAS 117:21757–21765. https://doi.org/10.1073/pnas.200611111
Cheng C-Y, Krishnakumar V, Chan AP, Thibaud-Nissen F, Schobel S, Town CD (2017) Araport11: a complete reannotation of the Arabidopsis thaliana reference genome. Plant J 89:789–804. https://doi.org/10.1111/tpj.13415
Dereeper A, Guignon V, Blanc G, Audic S, Buffet S, Chevenet F, Dufayard JF, Guindon S, Lefort V, Lescot M, Claverie JM, Gascuel O (2008) Phylogeny. fr: robust phylogenetic analysis for the non-specialist. Nucleic Acids Res 36(Web Server issue):W465–W469. https://doi.org/10.1093/nar/gkn180
Diao Y, Zhan J, Zhao Y, Liu L, Liu P, Wei X, Ding Y, Sajjad M, Hu W, Wang P, Ge X (2019) GhTIE1 regulates branching through modulating the transcriptional activity of TCPs in cotton and Arabidopsis. Front Plant Sci 10:1348. https://doi.org/10.3389/fpls.2019.01348
Edgar RC (2004) MUSCLE: multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res 32(5):1792–1797. https://doi.org/10.1093/nar/gkh340
El-Gebali S, Mistry J, Bateman A, Eddy SR, Luciani A, Potter SC, Qureshi M, Richardson LJ, Salazar GA, Smart A, Sonnhammer ELL, Hirsh L, Paladin L, Piovesan D, Tosatto SCE, Finn RD (2019) The Pfam protein families database in 2019. Nucleic Acids Res 47(D1):D427–D432. https://doi.org/10.1093/nar/gky995
Garg R, Shankar R, Thakkar B, Kudapa H, Krishnamurthy L, Mantri N, Varshney RK, Bhatia S, Jain M (2016) Transcriptome analyses reveal genotype- and developmental stage-specific molecular responses to drought and salinity stresses in chickpea. Sci Rep 6:19228. https://doi.org/10.1038/srep19228
Goodstein DM, Shu S, Howson R, Neupane R, Hayes RD, Fazo J, Mitros T, Dirks W, Hellsten U, Putnam N, Rokhsar DS (2012) Phytozome: a comparative platform for green plant genomics. Nucleic Acids Res 40:D1178–D1186. https://doi.org/10.1093/nar/gkr944
Hu B, Jin J, Guo A-Y, Zhang H, Luo J, Gao G (2014) GSDS 2.0: an upgraded gene feature visualization server. Bioinformatics 31:1296–1297. https://doi.org/10.1093/bioinformatics/btu817
Hu J, Ji Y, Hu X, Sun S, Wang X (2020) BES1 functions as the co-regulator of D53-like SMXLs to inhibit BRC1 expression in strigolactone-regulated shoot branching in Arabidopsis. Plant Commun 1:100014. https://doi.org/10.1016/j.xplc.2019.100014
Humann J, Jung S, Cheng C-H, Lee T, Zheng P, Frank M, McGaughey D, Scott K, Buble K, Yu J, Hough H, Sanad M, Coyne C, McGee R, Main D (2019) Cool season food legume genome database: a resource for pea, lentil, faba bean and chickpea genetics, genomics and breeding. Proceedings of the International Plant and Animal Genome Conference, San Diego, CA, USA
Jain M, Bansal J, Rajkumar MS, Garg R (2022) An integrated transcriptome mapping the regulatory network of coding and long non-coding RNAs provides a genomics resource in chickpea. Commun Biol 5:1106. https://doi.org/10.1038/s42003-022-04083-4
Jia T, Zhu L, Xiao G, Li H (2022) Genome-wide identification and expression analysis of the SMXL gene family in cotton. Scientia Sinica Vitae 52:1868–1882. https://doi.org/10.1360/SSV-2021-0023
Jiangtao C, Yingzhen K, Qian W, Yuhe S, Daping G, Jing L, Guanshan L (2015) MapGene2Chrom, a tool to draw gene physical map based on Perl and SVG languages. Yi chuan = Hereditas 37:91–97. https://doi.org/10.16288/j.yczz.2015.01.013
Kaashyap M, Ford R, Mann A, Varshney RK, Siddique KHM, Mantri N (2022) Comparative flower transcriptome network analysis reveals DEGs involved in chickpea reproductive success during salinity. Plants 11:434. https://doi.org/10.3390/plants11030434
Karalija E, Vergata C, Basso MF, Negussu M, Zaccai M, Grossi-de-Sa MF, Martinelli F (2022) Chickpeas’ tolerance of drought and heat: Current knowledge and next steps. Agronomy 12:2248. https://doi.org/10.3390/agronomy12102248
Khorramdelazad M, Bar I, Whatmore P, Smetham G, Bhaaskaria V, Yang Y, Bai SH, Mantri N, Zhou Y, Ford R (2018) Transcriptome profiling of lentil (Lens culinaris) through the first 24 hours of Ascochyta lentis infection reveals key defence response genes. BMC Genom 19:108. https://doi.org/10.1186/s12864-018-4488-1
Khosla A, Morffy N, Li Q, Faure L, Chang SH, Yao J, Zheng J, Cai ML, Stanga J, Flematti GR, Waters MT, Nelson DC (2020) Structure-function analysis of SMAX1 reveals domains that mediate its karrikin-induced proteolysis and interaction with the receptor KAI2. Plant Cell 32(8):2639–2659. https://doi.org/10.1105/tpc.19.00752
Koul B, Sharma K, Sehgal V, Yadav D, Mishra M, Bharadwaj C (2022) Chickpea (Cicer arietinum L.) biology and biotechnology: from domestication to biofortification and biopharming. Plants 11:2926. https://doi.org/10.3390/plants11212926
Kudapa H, Barmukh R, Garg V, Chitikineni A, Samineni S, Agarwal G, Varshney RK (2023) Comprehensive transcriptome profiling uncovers molecular mechanisms and potential candidate genes associated with heat stress response in chickpea. Int J Mol Sci 24:1369. https://doi.org/10.3390/ijms24021369
Kumar M, Chauhan AS, Kumar M, Yusuf MA, Sanyal I, Chauhan PS (2019) Transcriptome sequencing of chickpea (Cicer arietinum L.) genotypes for identification of drought-responsive genes under drought stress condition. Plant Mol Biol Report 37:186–203. https://doi.org/10.1007/s11105-019-01147-4
Kumar J, Gupta DS, Kesari R, Verma R, Murugesan S, Basu PS, Soren KR, Gupta S, Singh NP (2021a) Comprehensive RNAseq analysis for identification of genes expressed under heat stress in lentil. Physiol Plant 173:1785–1807. https://doi.org/10.1111/ppl.13419
Kumar N, Soren KR, Bharadwaj C, Priya PRS, Shrivastava AK, Pal M, Roorkiwal M, Kumar K, Patil BS, Soni A, Nimmy MS, Siddique KHM, Varshney RK (2021b) Genome-wide transcriptome analysis and physiological variation modulates gene regulatory networks acclimating salinity tolerance in chickpea. Environ Exp Bot 187:104478. https://doi.org/10.1016/j.envexpbot.2021.104478
Landi N, Piccolella S, Ragucci S, Faramarzi S, Clemente A, Papa S, Pacifico S, Di Maro A (2021) Valle agricola chickpeas: nutritional profile and metabolomics traits of a typical landrace legume from southern Italy. Foods. https://doi.org/10.3390/foods10030583
Lescot M, Déhais P, Thijs G, Marchal K, Moreau Y, Van de Peer Y, Rouzé P, Rombauts S (2002) PlantCARE, a database of plant cis-acting regulatory elements and a portal to tools for in silico analysis of promoter sequences. Nucleic Acids Res 30:325–327. https://doi.org/10.1093/nar/30.1.325
Li R, An J-P, You C-X, Wang X-F, Hao Y-J (2018) Genome-wide analysis and identification of the SMXL gene family in apple (Malus × domestica). Tree Genet Genom 14:61. https://doi.org/10.1007/s11295-018-1275-8
Li Q, Martín-Fontecha ES, Khosla A, White ARF, Chang S, Cubas P, Nelson DC (2022) The strigolactone receptor D14 targets SMAX1 for degradation in response to GR24 treatment and osmotic stress. Plant Commun 3:100303. https://doi.org/10.1016/j.xplc.2022.100303
Liang Y, Ward S, Li P, Bennett T, Leyser O (2016) SMAX1-LIKE7 signals from the nucleus to regulate shoot development in Arabidopsis via partially EAR motif-independent mechanisms. Plant Cell 28(7):1581–1601. https://doi.org/10.1105/tpc.16.00286
Lopez-Obando M, Ligerot Y, Bonhomme S, Boyer FD, Rameau C (2015) Strigolactone biosynthesis and signaling in plant development. Development (Cambridge, England) 142:3615–3619. https://doi.org/10.1242/dev.120006
Madeira F, Park YM, Lee J, Buso N, Gur T, Madhusoodanan N, Basutkar P, Tivey ARN, Potter SC, Finn RD, Lopez R (2019) The EMBL-EBI search and sequence analysis tools APIs in 2019. Nucleic Acids Res 47:W636–W641. https://doi.org/10.1093/nar/gkz268
Madeira F, Pearce M, Tivey ARN, Basutkar P, Lee J, Edbali O, Madhusoodanan N, Kolesnikov A, Lopez R (2022) Search and sequence analysis tools services from EMBL-EBI in 2022. Nucleic Acids Res 50:W276–W279. https://doi.org/10.1093/nar/gkac240
Marchler-Bauer A, Derbyshire MK, Gonzales NR, Lu S, Chitsaz F, Geer LY, Geer RC, He J, Gwadz M, Hurwitz DI, Lanczycki CJ, Lu F, Marchler GH, Song JS, Thanki N, Wang Z, Yamashita RA, Zhang D, Zheng C, Bryant SH (2015) CDD: NCBI’s conserved domain database. Nucleic Acids Res 43(1Database issue):D222–D226. https://doi.org/10.1093/nar/gku1221
Mashaki KM, Garg V, Ghomi AAN, Kudapa H, Chitikineni A, Nezhad KZ, Yamchi A, Soltanloo H, Varshney RK, Thudi M (2018) RNA-Seq analysis revealed genes associated with drought stress response in kabuli chickpea (Cicer arietinum L.). PLoS ONE 13:e0199774. https://doi.org/10.1371/journal.pone.0199774
Maurya JP, Miskolczi PC, Mishra S, Singh RK, Bhalerao RP (2020a) A genetic framework for regulation and seasonal adaptation of shoot architecture in hybrid aspen. PNAS 117:11523–11530. https://doi.org/10.1073/pnas.2004705117
Maurya JP, Singh RK, Miskolczi PC, Prasad AN, Jonsson K, Wu F, Bhalerao RP (2020b) Branching regulator BRC1 mediates photoperiodic control of seasonal growth in hybrid aspen. Curr Biol 30(1):122-126.e2. https://doi.org/10.1016/j.cub.2019.11.001
Mishra GP, Aski MS, Bosamia T, Chaurasia S, Mishra DC, Bhati J, Kumar A, Javeria S, Tripathi K, Kohli M, Kumar RR, Singh AK, Devi J, Kumar S, Dikshit HK (2021) Insights into the host-pathogen interaction pathways through RNA-seq analysis of Lens culinaris Medik in response to Rhizoctonia bataticola infection. Genes (Basel) 13(1):90. https://doi.org/10.3390/genes13010090
Morgil H, Tardu M, Cevahir G, Kavakli İH (2019) Comparative RNA-seq analysis of the drought-sensitive lentil (Lens culinaris) root and leaf under short- and long-term water deficits. Funct Integr Genom 19:715–727. https://doi.org/10.1007/s10142-019-00675-2
Nelson DC, Flematti GR, Riseborough JA, Ghisalberti EL, Dixon KW, Smith SM (2010) Karrikins enhance light responses during germination and seedling development in Arabidopsis thaliana. PNAS 107(15):7095–7100. https://doi.org/10.1073/pnas.0911635107
Piergiovanni AR (2022) Ex situ conservation of plant genetic resources: An overview of chickpea (Cicer arietinum L.) and lentil (Lens culinaris Medik) worldwide collections. Diversity 14:941. https://doi.org/10.3390/d14110941
Ramsay L, Koh CS, Kagale S, Gao D, Kaur S, Haile T, Gela TS, Chen L-A, Cao Z, Konkin DJ, Toegelová H, Doležel J, Rosen BD, Stonehouse R, Humann JL, Main D, Coyne CJ, McGee RJ, Cook DR, Penmetsa RV, Vandenberg A, Chan C, Banniza S, Edwards D, Bayer PE, Batley J, Udupa SM, Bett KE (2021) Genomic rearrangements have consequences for introgression breeding as revealed by genome assemblies of wild and cultivated lentil species. BioRxiv. https://doi.org/10.1101/2021.07.23.453237
Reddy DS, Bhatnagar-Mathur P, Reddy PS, Cindhuri KS, Ganesh AS, Sharma KK (2016) Identification and validation of reference genes and their impact on normalized gene expression studies across cultivated and wild cicer species. PLoS One 11:e0148451. https://doi.org/10.1371/journal.pone.0148451
Saitou N, Nei M (1987) The Neighbor-Joining method: A new method for reconstructing phylogenetic trees. Mol Biol Evol 4:406–425. https://doi.org/10.1093/oxfordjournals.molbev.a040454
Schmutz J, Cannon SB, Schlueter J, Ma J, Mitros T, Nelson W, Hyten DL, Song Q, Thelen JJ, Cheng J, Xu D, Hellsten U, May GD, Yu Y, Sakurai T, Umezawa T, Bhattacharyya MK, Sandhu D, Valliyodan B, Lindquist E, Peto M, Grant D, Shu S, Goodstein D, Barry K, Futrell-Griggs M, Abernathy B, Du J, Tian Z, Zhu L, Gill N, Joshi T, Libault M, Sethuraman A, Zhang XC, Shinozaki K, Nguyen HT, Wing RA, Cregan P, Specht J, Grimwood J, Rokhsar D, Stacey G, Shoemaker RC, Jackson SA (2010) Genome sequence of the palaeopolyploid soybean. Nature 463(7278):178–183. https://doi.org/10.1038/nature08670
Sepulveda C, Guzmán MA, Li Q, Villaécija-Aguilar JA, Martinez SE, Kamran M, Khosla A, Liu W, Gendron JM, Gutjahr C, Waters MT, Nelson DC (2022) Karrikin up-regulated F-box 1 (KUF1) imposes negative feedback regulation of karrikin and KAI2 ligand metabolism in Arabidopsis thaliana. PNAS 119(11):e2112820119. https://doi.org/10.1073/pnas.2112820119
Silva-Perez V, Shunmugam ASK, Rao S, Cossani CM, Tefera AT, Fitzgerald GJ, Armstrong R, Rosewarne GM (2022) Breeding has selected for architectural and photosynthetic traits in lentils. Front Plant Sci 13:925987. https://doi.org/10.3389/fpls.2022.925987
Singh D, Singh CK, Taunk J, Jadon V, Pal M, Gaikwad K (2019a) Genome wide transcriptome analysis reveals vital role of heat responsive genes in regulatory mechanisms of lentil (Lens culinaris Medikus). Sci Rep 9:12976. https://doi.org/10.1038/s41598-019-49496-0
Singh U, Gaur PM, Chaturvedi SK, Hazra KK, Singh G (2019b) Changing plant architecture and density can increase chickpea productivity and facilitate for mechanical harvesting. Int J Plant Product 13:193–202. https://doi.org/10.1007/s42106-019-00047-7
Singh D, Singh CK, Taunk J, Sharma S, Gaikwad K, Singh V, Sanwal SK, Singh D, Sharma PC, Pal M (2021) Transcriptome skimming of lentil (Lens culinaris Medikus) cultivars with contrast reaction to salt stress. Funct Integr Genom 21:139–156. https://doi.org/10.1007/s10142-020-00766-5
Singh D, Singh CK, Taunk J, Gaikwad K, Singh V, Sanwal SK, Karwa S, Singh D, Sharma PC, Yadav RK, Pal M (2022a) Linking genome wide RNA sequencing with physio-biochemical and cytological responses to catalogue key genes and metabolic pathways for alkalinity stress tolerance in lentil (Lens culinaris Medikus). BMC Plant Biol 22:99. https://doi.org/10.1186/s12870-022-03489-w
Singh D, Taunk J, Singh CK, Chaudhary P, Gaikwad K, Yadav RK, Singh D, Pal M (2022b) Comparative RNA sequencing for deciphering nodes of multiple abiotic stress tolerance in lentil (Lens culinaris Medikus). Plant Gene 31:100373. https://doi.org/10.1016/j.plgene.2022.100373
Sinha R, Sharma TR, Singh AK (2019) Validation of reference genes for qRT-PCR data normalisation in lentil (Lens culinaris) under leaf developmental stages and abiotic stresses. Physiol Mol Biol Plants 25:123–134. https://doi.org/10.1007/s12298-018-0609-1
Sohrabi SS, Ismaili A, Nazarian-Firouzabadi F, Fallahi H, Hosseini SZ (2022) Identification of key genes and molecular mechanisms associated with temperature stress in lentil. Gene 807:145952. https://doi.org/10.1016/j.gene.2021.145952
Soundappan I, Bennett T, Morffy N, Liang Y, Stanga JP, Abbas A, Leyser O, Nelson DC (2015) SMAX1-LIKE/D53 family members enable distinct MAX2-dependent responses to strigolactones and karrikins in Arabidopsis. Plant Cell 27:3143–3159. https://doi.org/10.1105/tpc.15.00562
Sperschneider J, Catanzariti A-M, DeBoer K, Petre B, Gardiner DM, Singh KB, Dodds PN, Taylor JM (2017) LOCALIZER: subcellular localization prediction of both plant and effector proteins in the plant cell. Sci Rep 7:44598. https://doi.org/10.1038/srep44598
Sudheesh S, Verma P, Forster JW, Cogan NOI, Kaur S (2016) Generation and characterisation of a reference transcriptome for lentil (Lens culinaris Medik). Int J Mol Sci 17:1887. https://doi.org/10.3390/ijms17111887
Sun H, Li W, Burritt DJ, Tian H, Zhang H, Liang X, Miao Y, Mostofa MG, Tran L-SP (2022) Strigolactones interact with other phytohormones to modulate plant root growth and development. Crop J 10:1517–1527. https://doi.org/10.1016/j.cj.2022.07.014
Sun M, Wang D, Liu C, Liu Y, Niu M, Wang J, Li J (2023) Genome-wide identification and analysis of the SUPPRESSOR of MAX2 1-LIKE gene family and its interaction with DWARF14 in poplar. BMC Plant Biol 23:105. https://doi.org/10.1186/s12870-023-04118-w
Szklarczyk D, Gable AL, Nastou KC, Lyon D, Kirsch R, Pyysalo S, Doncheva NT, Legeay M, Fang T, Bork P, Jensen LJ, von Mering C (2020) The STRING database in 2021: customizable protein-protein networks, and functional characterization of user-uploaded gene/measurement sets. Nucleic Acids Res 49:D605–D612. https://doi.org/10.1093/nar/gkaa1074
Tamura K, Stecher G, Kumar S (2021) MEGA11: Molecular evolutionary genetics analysis version 11. Mol Biol Evol 38:3022–3027. https://doi.org/10.1093/molbev/msab120
Tripathi K, Kumari J, Gore PG, Mishra DC, Singh AK, Mishra GP, Gayacharan C, Dikshit HK, Singh N, Semwal DP, Mehra R, Bhardwaj R, Bansal R, Rana JC, Kumar A, Gupta V, Singh K, Sarker A (2022) Agro-morphological characterization of lentil germplasm of Indian National Genebank and development of a core set for efficient utilization in lentil improvement programs. Front Plant Sci 12:751429. https://doi.org/10.3389/fpls.2021.751429
Tunyasuvunakool K, Adler J, Wu Z, Green T, Zielinski M, Žídek A, Bridgland A, Cowie A, Meyer C, Laydon A, Velankar S, Kleywegt GJ, Bateman A, Evans R, Pritzel A, Figurnov M, Ronneberger O, Bates R, Kohl SAA, Potapenko A, Ballard AJ, Romera-Paredes B, Nikolov S, Jain R, Clancy E, Reiman D, Petersen S, Senior AW, Kavukcuoglu K, Birney E, Kohli P, Jumper J, Hassabis D (2021) Highly accurate protein structure prediction for the human proteome. Nature 596:590–596. https://doi.org/10.1038/s41586-021-03828-1
Varshney RK, Song C, Saxena RK, Azam S, Yu S, Sharpe AG, Cannon S, Baek J, Rosen BD, Tar’an B, Millan T, Zhang X, Ramsay LD, Iwata A, Wang Y, Nelson W, Farmer AD, Gaur PM, Soderlund C, Penmetsa RV, Xu C, Bharti AK, He W, Winter P, Zhao S, Hane JK, Carrasquilla-Garcia N, Condie JA, Upadhyaya HD, Luo M-C, Thudi M, Gowda CLL, Singh NP, Lichtenzveig J, Gali KK, Rubio J, Nadarajan N, Dolezel J, Bansal KC, Xu X, Edwards D, Zhang G, Kahl G, Gil J, Singh KB, Datta SK, Jackson SA, Wang J, Cook DR (2013) Draft genome sequence of chickpea (Cicer arietinum) provides a resource for trait improvement. Nat Biotechnol 31:240–246. https://doi.org/10.1038/nbt.2491
Villaécija-Aguilar JA, Hamon-Josse M, Carbonnel S, Kretschmar A, Schmidt C, Dawid C, Bennett T, Gutjahr C (2019) SMAX1/SMXL2 regulate root and root hair development downstream of KAI2-mediated signalling in Arabidopsis. PLoS Genet 15:e1008327. https://doi.org/10.1371/journal.pgen.1008327
Wallner ES, López-Salmerón V, Belevich I, Poschet G, Jung I, Grünwald K, Sevilem I, Jokitalo E, Hell R, Helariutta Y, Agustí J, Lebovka I, Greb T (2017) Strigolactone- and karrikin-independent SMXL proteins are central regulators of phloem formation. Curr Biol 27:1241–1247. https://doi.org/10.1016/j.cub.2017.03.014
Wang L, Wang B, Jiang L, Liu X, Li X, Lu Z, Meng X, Wang Y, Smith SM, Li J (2015) Strigolactone signaling in arabidopsis regulates shoot development by targeting D53-like SMXL repressor proteins for ubiquitination and degradation. Plant Cell 27:3128–3142. https://doi.org/10.1105/tpc.15.00605
Wang L, Xu Q, Yu H, Ma H, Li X, Yang J, Chu J, Xie Q, Wang Y, Smith SM, Li J, Xiong G, Wang B (2020) Strigolactone and karrikin signaling pathways elicit ubiquitination and proteolysis of SMXL2 to regulate hypocotyl elongation in Arabidopsis. Plant Cell 32:2251–2270. https://doi.org/10.1105/tpc.20.00140
Webb B, Sali A (2016) Comparative protein structure modeling using MODELLER. Curr Protocols Bioinform 54:561–567. https://doi.org/10.1002/cpbi.3
Yang T, Lian Y, Wang C (2019) Comparing and contrasting the multiple roles of butenolide plant growth regulators: Strigolactones and karrikins in plant development and adaptation to abiotic stresses. Int J Mol Sci 20:6270. https://doi.org/10.3390/ijms20246270
Yang T, Lian Y, Kang J, Bian Z, Xuan L, Gao Z, Wang X, Deng J, Wang C (2020) The SUPPRESSOR of MAX2 1 (SMAX1)-like SMXL6, SMXL7 and SMXL8 act as negative regulators in response to drought stress in Arabidopsis. Plant Cell Physiol 61:1477–1492. https://doi.org/10.1093/pcp/pcaa066
Yang T, Liu K, Poppy L, Mulenga A, Gampe C (2021) Minimizing lentil harvest loss through improved agronomic practices in sustainable agro-systems. Sustainability 13:1896. https://doi.org/10.3390/su13041896
Yao R, Ming Z, Yan L, Li S, Wang F, Ma S, Yu C, Yang M, Chen L, Chen L, Li Y, Yan C, Miao D, Sun Z, Yan J, Sun Y, Wang L, Chu J, Fan S, He W, Deng H, Nan F, Li J, Rao Z, Lou Z, Xie D (2016) DWARF14 is a non-canonical hormone receptor for strigolactone. Nature 536:469–473. https://doi.org/10.1038/nature19073
Zhang H, Wang L, Gao Y, Guo Y, Zheng N, Xu X, Xu M, Wang W, Liu C, Liu W, Yang W (2022) Genome-wide identification of SMXL gene family in soybean and expression analysis of GmSMXLs under shade stress. Plants 11(18):2410. https://doi.org/10.3390/plants11182410
Zhao LH, Zhou XE, Yi W, Wu Z, Liu Y, Kang Y, Hou L, de Waal PW, Li S, Jiang Y, Scaffidi A, Flematti GR, Smith SM, Lam VQ, Griffin PR, Wang Y, Li J, Melcher K, Xu HE (2015) Destabilization of strigolactone receptor DWARF14 by binding of ligand and E3-ligase signaling effector DWARF3. Cell Res 25:1219–1236. https://doi.org/10.1038/cr.2015.122
Zhou F, Lin Q, Zhu L, Ren Y, Zhou K, Shabek N, Wu F, Mao H, Dong W, Gan L, Ma W, Gao H, Chen J, Yang C, Wang D, Tan J, Zhang X, Guo X, Wang J, Jiang L, Liu X, Chen W, Chu J, Yan C, Ueno K, Ito S, Asami T, Cheng Z, Wang J, Lei C, Zhai H, Wu C, Wang H, Zheng N, Wan J (2013) D14-SCF(D3)-dependent degradation of D53 regulates strigolactone signalling. Nature 504:406–410. https://doi.org/10.1038/nature12878
Funding
Open access funding provided by Università degli Studi di Firenze within the CRUI-CARE Agreement. MFB is grateful to CNPq for a postdoctoral research fellowship (process number: 106655/2023-0). The authors were supported by PRIMA-The Partnership for Research and Innovation in the Mediterranean Area. The PRIMA program is an Art.185 initiative supported and funded under Horizon 2020, the European Union’s Framework Program for Research and Innovation. Project No. 1432-LEGU-MED2-Legumes in biodiversity-based farming systems in Mediterranean basin project funded. In addition, FM was supported by the Israeli Ministry of Science and Technology and the Italian Ministry of Foreign Affairs and International Cooperation (Grant # 3-17924; project name: Resilient Hummus).
Author information
Authors and Affiliations
Contributions
MFB performed identification and in silico analysis of the SMAX/SMXL gene and promoter sequences, plant experiments, RNA, cDNA, and gene expression assays using real-time PCR. CMB helped with in silico analysis. FC performed the meta-analysis related to different stress conditions. FLC and GB analyzed the protein 3D structures. MFB prepared the data and wrote the final manuscript. FM, MFGS, and AF provided intellectual input in the final version of the manuscript, while FM provided financial resources. All authors approved the final version.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that this research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Consent for publication or ethics approval and consent to participate
Not applicable.
Additional information
Communicated by Dorothea Bartels.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Marcos Fernando Basso: This work was part of the TCC of the first author in the MBA in agribusiness USP/ESALQ 2022/23.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Basso, M.F., Contaldi, F., Lo Celso, F. et al. Identification and expression profile of the SMAX/SMXL family genes in chickpea and lentil provide important players of biotechnological interest involved in plant branching. Planta 259, 1 (2024). https://doi.org/10.1007/s00425-023-04277-y
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00425-023-04277-y