Abstract
Main conclusion
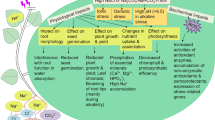
SbMYB15, R2R3-type MYB was induced by the different stresses, and conferred stress tolerance in transgenic tobacco by regulating the expression of stress-responsive genes.
MYBs are the master regulators of various metabolic processes and stress responses in plants. In this study, we functionally characterised a R2R3-type SbMYB15 transcription factor (TF) from the extreme halophyte Salicornia brachiata. The SbMYB15 acts as a transcriptional activator. Transcriptional analysis showed that SbMYB15 transcript was strongly upregulated in red shoots and was also induced by different stresses; however, its expression remained unchanged with ABA. Overexpression of SbMYB15 in tobacco significantly improved salinity and dehydration tolerance. The enhanced tolerance of the transgenic plants was defined by the changes in chlorophyll, malondialdehyde (MDA), proline, total soluble sugar and total amino acid contents. The transgenic plants exhibited a higher membrane stability and reduced electrolyte leakage, H2O2 and O −2 content compared to the wild type (WT). With ionic stress, transgenics showed a low Na+ and a high K+ content. In the transgenic plants, the expression of stress-responsive genes such as LEA5, ERD10D, PLC3, LTP1, HSF2, ADC, P5CS, SOD and CAT was enhanced in the presence of salinity, dehydration and heat. Exposure to gradual salinity and dehydration resulted in an increased stomatal conductance, water use efficiency, photosynthesis rate, photochemical quenching and reduced transpiration rate. Thus, SbMYB15 served as an important mediator of stress responses regulating different stress signalling pathways, leading to enhanced stress tolerance.








Similar content being viewed by others
Abbreviations
- ABA:
-
Abscisic acid
- Ci:
-
Intercellular CO2 concentration
- Ca:
-
Ambient CO2 concentration
- ETR:
-
Photosynthetic electron transport rate
- g:
-
Stomatal conductance
- MDA:
-
Malondialdehyde
- NPQ:
-
Non-photochemical fluorescence quenching
- PEG:
-
Polyethylene glycol
- SA:
-
Salicylic acid
- TF:
-
Transcription factor
- VA:
-
Transgenic plants transformed with pCAMBIA 1301
- WUE:
-
Water use efficiency
References
Abe H, Urao T, Ito T, Seki M, Shinozaki K, Yamaguchi-Shinozaki K (2003) Arabidopsis AtMYC2 (bHLH) and AtMYB2 (MYB) function as transcriptional activators in abscisic acid signaling. Plant Cell 15:63–78
Adams III WW, Muller O, Cohu CM, Demmig-Adams B (2014) Photosystem II efficiency and non-photochemical fluorescence quenching in the context of source-sink balance. In: Demmig-Adams B, Garab G, Adam III WW, Govindjee (eds) Non-photochemical quenching and energy dissipation in plants, algae and cyanobacteria. Springer, Dordrecht, pp 503–529. doi:10.1007/978-94-017-9032-1
Agarwal PK, Jha B (2010) Transcription factors in plants and ABA dependent and independent abiotic stress signalling. Biol Plant 54:201–212
Agarwal S, Sairam RK, Srivastava GC, Tyagi A, Meena RC (2005) Role of ABA, salicylic acid, calcium and hydrogen peroxide on antioxidant enzymes induction in wheat seedlings. Plant Sci 169:559–570
Agarwal PK, Shukla PS, Gupta K, Jha B (2013) Bioengineering for salinity tolerance in plants: state of the art. Mol Biotechnol 54:102–123
Alcázar R, Marco F, Cuevas JC, Patron M, Ferrando A, Carrasco P, Tiburcio AF, Altabella T (2006) Involvement of polyamines in plant response to abiotic stress. Biotechnol Lett 28:1867–1876
Ashraf M, Harris PJC (2013) Photosynthesis under stressful environments: an overview. Photosynthetica 51:163–190
Baker NR (2008) Chlorophyll fluorescence: a probe of photosynthesis in vivo. Annu Rev Plant Biol 59:89–113
Batra NG, Sharma V, Kumari N (2014) Drought-induced changes in chlorophyll fluorescence, photosynthetic pigments, and thylakoid membrane proteins of Vigna radiata. J Plant Interact 9:712–721
Biswal B, Joshi PN, Raval MK, Biswal UC (2011) Photosynthesis, a global sensor of environmental stress in green plants: stress signalling and adaptation. Curr Sci 101:47–56
Braun EL, Grotewold E (1999) Newly discovered plant c-myb-like genes rewrite the evolution of the plant myb gene family. Plant Physiol 121:21–24
Chen BJ, Wang Y, Hu YL, Wu Q, Lin ZP (2005) Cloning and characterization of a drought-inducible MYB gene from Boea crassifolia. Plant Sci 168:493–500
Chen YH, Yang XY, He K, Liu MH et al (2006) The MYB transcription factor superfamily of Arabidopsis: expression analysis and phylogenetic comparison with the rice MYB family. Plant Mol Biol 60:107–124
Chen N, Yang Q, Pan L, Chi X, Chen M, Hu D, Yang Z, Wang T, Wang M, Yu S (2014) Identification of 30 MYB transcription factor genes and analysis of their expression during abiotic stress in peanut (Arachis hypogaea L.). Gene 533:332–345
Chomczynski P, Chomczynski P, Sacchi N, Sacchi N (2006) The single-step method of RNA isolation by acid guanidinium thiocyanate–phenol–chloroform extraction: twenty-something years on. Nat Protoc 1:581–585
Close TJ (1997) Dehydrins: a commonalty in the response of plants to dehydration and low temperature. Physiol Plant 100:291–296
Cominelli E, Galbiati M, Vavasseur A, Conti L, Sala T, Vuylsteke M, Leonhardt N, Dellaporta SL, Tonelli C (2005) A guard-cell-specific MYB transcription factor regulates stomatal movements and plant drought tolerance. Curr Biol 15:1196–1200
Cook GD, Dixon JR, Leopold AC (1964) Transpiration: its effects on plant leaf temperature. Science 144:546–547
Crooks GE, Hon G, Chandonia JM, Brenner SE (2004) WebLogo: a sequence logo generator. Genome Res 14:1188–1190
Doyle JJ, Doyle JL (1987) A rapid DNA isolation procedure for small quantities of fresh leaf tissue. Phytochem Bull 19:11–15
Du H, Feng B-R, Yang S-S, Huang Y-B, Tang Y-X (2012) The R2R3-MYB transcription factor gene family in maize. PLoS One 7:e37463
Dubos C, Stracke R, Grotewold E, Weisshaar B, Martin C, Lepiniec L (2010) MYB transcription factors in Arabidopsis. Trends Plant Sci 15:573–581
Flexas J, Escalona JM, Evain S, Gulías J, Moya I, Osmond CB, Medrano H (2002) Steady-state chlorophyll fluorescence (Fs) measurements as a tool to follow variations of net CO2 assimilation and stomatal conductance during water-stress in C3 plants. Physiol Plant 114:231–240
Gao JJ, Zhang Z, Peng R-H, Xiong A-S, Xu J, Zhu B, Yao QH (2011) Forced expression of Mdmyb10, a myb transcription factor gene from apple, enhances tolerance to osmotic stress in transgenic Arabidopsis. Mol Biol Rep 38:205–211
Garg AK, Kim J, Owens TG, Ranwala AP, Do Choi Y, Kochian LV, Wu RJ (2002) Trehalose accumulation in rice plants confers high tolerance levels to different abiotic stresses. Proc Natl Acad Sci USA 99:15898–15903
Guo L, Yang H, Zhang X, Yang S (2013) Lipid transfer protein 3 as a target of MYB96 mediates freezing and drought stress in Arabidopsis. J Exp Bot 64:1755–1767
Gupta K, Agarwal PK, Reddy MK, Jha B (2010) SbDREB2A, an A-2 type DREB transcription factor from extreme halophyte Salicornia brachiata confers abiotic stress tolerance in Escherichia coli. Plant Cell Rep 29:1131–1137
Gupta K, Jha B, Agarwal PK (2014) A dehydration-responsive element binding (DREB) transcription factor from the succulent halophyte Salicornia brachiata enhances abiotic stress tolerance in transgenic tobacco. Mar Biotechnol 16:657–673
He Y, Li W, Lv J, Jia Y, Wang M, Xia G (2012) Ectopic expression of a wheat MYB transcription factor gene, TaMYB73, improves salinity stress tolerance in Arabidopsis thaliana. J Exp Bot 63:1511–1522
Hoagland DR, Arnon DI (1950) The water-culture method for growing plants without soil. Calif Agric Exp Stn Circ 347:1–32
Hodges DM, DeLong JM, Forney CF, Prange RK (1999) Improving the thiobarbituric acid-reactive-substances assay for estimating lipid peroxidation in plant tissues containing anthocyanin and other interfering compounds. Planta 207:604–611
Horsch RB, Fry JE, Hoffmann NL, Eichholtz D, Rogers SG, Farley RT (1985) A simple and general method for transferring genes into plants. Science 227:1229–1231
Jha B, Agarwal PK, Reddy PS, Lal S, Sopory SK, Reddy MK (2009) Identification of salt-induced genes from Salicornia brachiata, an extreme halophyte through expressed sequence tags analysis. Genes Genet Syst 84:111–120
Jia L, Clegg MT, Jiang T (2004) Evolutionary dynamics of the DNA-binding domains in putative R2R3-MYB genes identified from rice subspecies indica and japonica genomes. Plant Physiol 134:575–585
Jung C, Seo JS, Han SW, Koo YJ, Kim CH, Song SI, Nahm BH, Do Choi Y, Cheong JJ (2008) Overexpression of AtMYB44 enhances stomatal closure to confer abiotic stress tolerance in transgenic Arabidopsis. Plant Physiol 146:623–635
Kang G, Li G, Guo T (2014) Molecular mechanism of salicylic acid-induced abiotic stress tolerance in higher plants. Acta Physiol Plant 36:2287–2297
Karaba A, Dixit S, Greco R, Aharoni A, Trijatmiko KR, Marsch-Martinez N, Krishnan A, Nataraja KN, Udayakumar M, Pereira A (2007) Improvement of water use efficiency in rice by expression of HARDY, an Arabidopsis drought and salt tolerance gene. Proc Natl Acad Sci USA 104:15270–15275
Kranz HD, Denekamp M, Greco R et al (1998) Towards functional characterisation of the members of the R2R3-MYB gene family from Arabidopsis thaliana. Plant J 16:263–276
Lee HG, Seo PJ (2015) The MYB96-HHP module integrates cold and ABA signaling to activate the CBF-COR pathway in Arabidopsis. Plant J. doi:10.1111/tpj.12866
Li C, Ng CK-Y, Fan L-M (2014) MYB transcription factors, active players in abiotic stress signaling. Environ Exp Bot. doi:10.1016/j.envexpbot.2014.06.014
Liao Y, Zou H-F, Wang H-W, Zhang W-K, Ma B, Zhang J-S, Chen SY (2008) Soybean GmMYB76, GmMYB92, and GmMYB177 genes confer stress tolerance in transgenic Arabidopsis plants. Cell Res 18:1047–1060
Lichtenthaler HK (1987) Chlorophylls and carotenoids: pigments of photosynthetic biomembranes. Methods Enzymol 148:350–382
Lindemose S, O’Shea C, Jensen MK, Skriver K (2013) Structure, function and networks of transcription factors involved in abiotic stress responses. Int J Mol Sci 14:842–878
Liu F, Pang SJ (2010) Stress tolerance and antioxidant enzymatic activities in the metabolisms of the reactive oxygen species in two intertidal red algae Grateloupia turuturu and Palmaria palmata. J Exp Mar Biol Ecol 382:82–87
Liu J, Shi DC (2010) Photosynthesis, chlorophyll fluorescence, inorganic ion and organic acid accumulations of sunflower in responses to salt and salt–alkaline mixed stress. Photosynthetica 48:127–134
Liu X, Yang L, Zhou X, Zhou M, Lu Y, Ma L, Ma H, Zhang Z (2013) Transgenic wheat expressing Thinopyrum intermedium MYB transcription factor TiMYB2R-1 shows enhanced resistance to the take-all disease. J Exp Bot 64:2243–2253
Livak KJ, Schmittgen TD (2001) Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) method. Methods 25:402–408
Ma Q, Dai X, Xu Y et al (2009) Enhanced tolerance to chilling stress in OsMYB3R-2 transgenic rice is mediated by alteration in cell cycle and ectopic expression of stress genes. Plant Physiol 150:244–256
Martínez-Ferri E, Balaguer L, Valladares F, Chico JM, Manrique E (2000) Energy dissipation in drought-avoiding and drought-tolerant tree species at midday during the Mediterranean summer. Tree Physiol 20:131–138
Mattana M, Biazzi E, Consonni R, Locatelli F, Vannini C, Provera S, Coraggio I (2005) Overexpression of Osmyb4 enhances compatible solute accumulation and increases stress tolerance of Arabidopsis thaliana. Physiol Plant 125:212–223
Maxwell K, Johnson GN (2000) Chlorophyll fluorescence–a practical guide. J Exp Bot 51:659–668
Mengiste T, Chen X, Salmeron J, Dietrich R (2003) The BOTRYTIS SUSCEPTIBLE1 gene encodes an R2R3 MYB transcription factor protein that is required for biotic and abiotic stress responses in Arabidopsis. Plant Cell 15:2551–2565
Mukherjee SP, Choudhuri MA (1983) Implications of water stress-induced changes in the levels of endogenous ascorbic acid and hydrogen peroxide in Vigna seedlings. Physiol Plant 58:166–170
Murashige T, Skoog F (1962) A revised medium for rapid growth and bio assays with tobacco tissue cultures. Physiol Plant 15:473–497
Ogata K, Kanei-Ishii C, Sasaki M, Hatanaka H, Nagadoi A, Enari M, Nakamura H, Nishimura Y, Ishii S, Sarai A (1996) The cavity in the hydrophobic core of Myb DNA-binding domain is reserved for DNA recognition and trans-activation. Nat Struct Biol 3:178–187
Osmond CB, Grace SC (1995) Perspectives on photoinhibition and photorespiration in the field: quintessential inefficiencies of the light and dark reactions of photosynthesis? J Exp Bot 46:1351–1362
Parida AK, Jha B (2013) Physiological and biochemical responses reveal the drought tolerance efficacy of the halophyte Salicornia brachiata. J Plant Growth Regul 32:342–352
Pasquali G, Biricolti S, Locatelli F, Baldoni E, Mattana M (2008) Osmyb4 expression improves adaptive responses to drought and cold stress in transgenic apples. Plant Cell Rep 27:1677–1686
Qin Y, Wang M, Tian Y, He W, Han L, Xia G (2012) Over-expression of TaMYB33 encoding a novel wheat MYB transcription factor increases salt and drought tolerance in Arabidopsis. Mol Biol Rep 39:7183–7192
Raffaele S, Vailleau F, Léger A, Joubès J, Miersch O, Huard C, Blée E, Mongrand S, Domergue F, Roby D (2008) A MYB transcription factor regulates very-long-chain fatty acid biosynthesis for activation of the hypersensitive cell death response in Arabidopsis. Plant Cell 20:752–767
Riechmann JL (2000) Arabidopsis transcription factors: genome-wide comparative analysis among eukaryotes. Science 290:2105–2110
Rost B, Liu J (2003) The PredictProtein server. Nucleic Acids Res 31:3300–3304
Seo PJ, Park CM (2010) MYB96-mediated abscisic acid signals induce pathogen resistance response by promoting salicylic acid biosynthesis in Arabidopsis. New Phytol 186:471–483
Shabala S, Pottosin I (2014) Regulation of potassium transport in plants under hostile conditions: implications for abiotic and biotic stress tolerance. Physiol Plant 151:257–279
Shavrukov Y (2013) Salt stress or salt shock: which genes are we studying? J Exp Bot 64:119–127
Shepherd CT, Lauter ANM, Scott MP (2009) Determination of transgene copy number by real-time quantitative PCR. In: Scott MP (ed) Methods in molecular biology: transgenic maize. Humana Press, New York, pp 129–134
Shi J, Fu XZ, Peng T, Huang XS, Fan QJ, Liu JH (2010) Spermine pretreatment confers dehydration tolerance of citrus in vitro plants via modulation of antioxidative capacity and stomatal response. Tree Physiol 30:914–923
Shim JS, Jung C, Lee S, Min K, Lee YW, Choi Y, Lee JS, Song JT, Kim JK, Choi YD (2013) AtMYB44 regulates WRKY70 expression and modulates antagonistic interaction between salicylic acid and jasmonic acid signaling. Plant J 73:483–495
Shukla PS, Agarwal PK, Jha B (2012) Improved salinity tolerance of Arachis hypogaea (L.) by the interaction of halotolerant plant-growth-promoting rhizobacteria. J Plant Growth Regul 31:195–206
Stracke R, Werber M, Weisshaar B (2001) The R2R3-MYB gene family in Arabidopsis thaliana. Curr Opin Plant Biol 4:447–456
Töpfer R, Matzeit V, Gronenborn B, Schell J, Steinbiss HH (1987) A set of plant expression vectors for transcriptional and translational fusions. Nucleic Acids Res 15:5890
Xiong H, Li J, Liu P, Duan J, Zhao Y, Guo X, Li Y, Zhang H, Ali J, Li Z (2014) Overexpression of OsMYB48-1, a novel MYB-related transcription factor, enhances drought and salinity tolerance in rice. PLoS ONE 9(3):e92913
Yang A, Dai X, Zhang W-H (2012) A R2R3-type MYB gene, OsMYB2, is involved in salt, cold, and dehydration tolerance in rice. J Exp Bot 63:2541–2556
Zhang L, Zhao G, Jia J, Liu X, Kong X (2012) Molecular characterization of 60 isolated wheat MYB genes and analysis of their expression during abiotic stress. J Exp Bot 63:203–214
Acknowledgments
CSIR-CSMCRI Communication No. 019/2015. The authors are thankful to Department of Science and Technology (DST) and Council of Scientific and Industrial Research (CSIR), New Delhi, India for financial assistance. P.S. Shukla is thankful to AcSIR for enrolment in Ph.D. and CSIR for financial support.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
425_2015_2366_MOESM1_ESM.pptx
Supplementary material 1 Fig. S1 Schematic representation of gradual exposure to treatments of N. tabacum WT and transgenic lines in the greenhouse(PPTX 1055 kb)
425_2015_2366_MOESM2_ESM.pptx
Supplementary material 2 Fig. S2 Sequence logos of R2–R3 repeats of SbMYB15. The overall height of each stack indicates the conservation of the sequence at particular position. The height of letters within each stack represents the relative frequency of the corresponding amino acid. The triangle indicates the positions of the conserved tryptophan (W) and phenylalanine (F) amino acids, which are identical in other MYB proteins (PPTX 284 kb)
425_2015_2366_MOESM3_ESM.pptx
Supplementary material 3 Fig. S3 Nucleotide and deduced amino acid sequences of the SbMYB15 gene. R2 and R3 repeats are marked by black arrows. Conserved tryptophan residues are shown by (*). The presence of SANT domains in SbMYB15 is demarcated by a red box. Linker region of R2 and R3 repeat is marked by a red dotted line. Blue and brown arrows represent a serine- and histidine-rich region, respectively. Green bars represent alpha helices and a red arrow represents the presence of a loop (PPTX 135 kb)
425_2015_2366_MOESM4_ESM.pptx
Supplementary material 4 Fig. S4 a Secondary structure prediction of SbMYB15 shows the presence of alpha helices, strands and coils. b Secondary structure composition. c Solvent accessibility of SbMYB15 protein (PPTX 205 kb)
425_2015_2366_MOESM6_ESM.pptx
Supplementary material 6 Fig. S6 Transactivation assay of SbMYB15. a Transformed yeast cells (AH109) with pGBKT7-SbMYB15 and pGBKT7 alone were grown on SD/-Trp/-His/-Ura medium. b Schematic representation of the plating. c Yeast cells transferred on filter paper showed β-galactosidase (encoded by LacZ gene) activity using X-gal staining (PPTX 137 kb)
425_2015_2366_MOESM7_ESM.pptx
Supplementary material 7 Fig. S7 a Diagrammatic representation of the pCAMBIA1301-35S:SbMYB15 construct. Screening of T1 transgenic plants. b–c PCR confirmation of T1 transgenics with SbMYB15 gene specific primers and hptII primers. d GUS staining of seedlings of N. tabacum WT (i), VA (pCAMBIA1301 alone) (ii), and 35S:SbMYB15 overexpressing transgenic lines (iii–vii). e GUS:NRA ratio for determination of copy number of transgene insertion in transgenics (PPTX 777 kb)
425_2015_2366_MOESM8_ESM.pptx
Supplementary material 8 Fig. S8 a Morphological analysis of N. tabacum transgenic lines (L-79, L-86, L-87, L-123, L-127, L-158 and VA) and WT plants in 0, 100 and 200 mM NaCl concentrations in hydroponic medium. b RT-PCR analysis of the three SbMYB15 representative overexpression lines. c Growth of N. tabacum WT and transgenic lines (L-87, L-123, L-127, L-158 and VA) in 0 and 10 % PEG in hydroponic medium (PPTX 754 kb)
Rights and permissions
About this article
Cite this article
Shukla, P.S., Gupta, K., Agarwal, P. et al. Overexpression of a novel SbMYB15 from Salicornia brachiata confers salinity and dehydration tolerance by reduced oxidative damage and improved photosynthesis in transgenic tobacco. Planta 242, 1291–1308 (2015). https://doi.org/10.1007/s00425-015-2366-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00425-015-2366-5