Abstract
Purpose
We conducted a meta-analysis assessing the association between the p.P240L (c.C719T) variant and the risk of nonsyndromic hearing loss (NSHL).
Methods
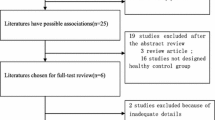
Literatures that reported prevalence rates were identified using PubMed, EMBASE, OVID, Cochrane library, Web of Science, Chinese National Knowledge Infrastructure, Wanfang Databases for the period from inception to August 2017. Random and fixed effects models were used to generate pooled ORs and I2 values. The heterogeneity assumption decided the effect model.
Results
A total of four relevant studies were included in the meta-analysis. The results of meta-analysis indicated that the p.P240L variant was correlated with the risk of NHSL in Asian populations (OR = 10.17, 95% CI = 2.74–37.82, P = 0.001). The T allele of p.P240L was associated with a 12-fold higher risk of NSHL than the C allele (OR = 11.68; 95% CI = 3.16–43.24, P < 0.001). Specifically, p.P240L heterozygotes (OR = 8.49; 95% CI = 2.28–31.59, P = 0.001), had a significantly higher risk of NSHL. Publication bias of our meta-analysis was attributed to the limited availability of relevant results and the number of studies included in our meta-analysis was relatively small.
Conclusions
The p.P240L variant increased the risk of NHSL in Asian populations, suggesting a remarkable ethnic specificity linked with susceptibility to this mutation.



Similar content being viewed by others
References
World Health Organization (2018) Global estimates on prevalence of hearing loss. http://www.who.int/pbd/deafness/estimates/en/. Accessed 25 May 2018
Friedman TB, Griffith AJ (2003) Human nonsyndromic sensorineural deafness. Annu Rev Genom Hum Genet 4:341–402
Astuto LM, Bork JM, Weston MD, Askew JW, Fields RR, Orten DJ et al (2002) CDH23 mutation and phenotype heterogeneity: a profile of 107 diverse families with Usher syndrome and nonsyndromic deafness. Am J Hum Genet 71:262–275
Mizutari K, Mutai H, Namba K, Miyanaga Y, Nakano A, Arimoto Y et al (2005) High prevalence of CDH23 mutations in patients with congenital high-frequency sporadic or recessively inherited hearing loss. Orphanet J Rare Dis 10:60
Kim BJ, Kim AR, Lee C, Kim SY, Kim NK, Chang MY et al (2016) Discovery of CDH23 as a significant contributor to progressive postlingual sensorineural hearing loss in Koreans. PLoS One 11:e0165680
Stang A (2010) Critical evaluation of the Newcastle–Ottawa scale for the assessment of the quality of nonrandomized studies in meta-analyses. Eur J Epidemiol 25:603–605
DerSimonian R, Laird N (1986) Meta-analysis in clinical trials. Control Clin Trials 7:177–188
Mantel N, Haenszel W (1959) Statistical aspects of the analysis of data from retrospective studies of disease. J Natl Cancer Inst 22:719–748
Wagatsuma M, Kitoh R, Suzuki H, Fukuoka H, Takumi Y, Usami S (2007) Distribution and frequencies of CDH23 mutations in Japanese patients with non-syndromic hearing loss. Clin Genet 72:339–344
Miyagawa M, Nishio SY, Usami S (2012) Prevalence and clinical features of hearing loss patients with CDH23 mutations: a large cohort study. PLoS One 7:e40366
Mutai H, Suzuki N, Shimizu A, Torii C, Namba K, Morimoto N et al (2013) Diverse spectrum of rare deafness genes underlies early-childhood hearing loss in Japanese patients: a cross-sectional, multi-center next-generation sequencing study. Orphanet J Rare Dis 8:172
Kim SY, Kim AR, Kim NK, Kim MY, Jeon EH, Kim BJ et al (2015) Strong founder effect of p.P240L in CDH23 in Koreans and its significant contribution to severe-to-profound nonsyndromic hearing loss in a Korean pediatric population. J Transl Med 13:263
Nele H, Smith JH, Van Camp G (2009) Function and expression pattern of nonsyndromic deafness genes. Curr Mol Med 9:546–564
Morton NE (1991) Genetic epidemiology of hearing impairment. Ann N Y Acad Sci 630:16–31
Nollet F, Kools P, van Roy F (2000) Phylogenetic analysis of the cadherin superfamily allows identification of six major subfamilies besides several solitary members. J Mol Biol 299:551–572
Bolz H, von Brederlow B, Ramirez A, Bryda EC, Kutsche K, Nothwang HG et al (2001) Mutation of CDH23, encoding a new member of the cadherin gene family, causes Usher syndrome type 1D. Nat Genet 27:108–112
Bork JM, Peters LM, Riazuddin S, Bernstein SL, Ahmed ZM, Ness SL et al (2001) Usher syndrome 1D and nonsyndromic autosomal recessive deafness DFNB12 are caused by allelic mutations of the novel cadherin-like gene CDH23. Am J Hum Genet 68:26–37
Wu H, Feng Y, Jiang L, Pan Q, Liu Y, Liu C et al (2016) Application of a new genetic deafness microarray for detecting mutations in the deaf in China. PLoS One 11:e0151909
Nishio SY, Usami S (2015) Deafness gene variations in a 1120 nonsyndromic hearing loss cohort: molecular epidemiology and deafness mutation spectrum of patients in Japan. Ann Otol Rhinol Laryngol 124(Suppl 1):49S–60S
Duman D, Sirmaci A, Cengiz FB, Ozdag H, Tekin M (2011) Screening of 38 genes identifies mutations in 62% of families with nonsyndromic deafness in Turkey. Genet Test Mol Biomark 15(1–2):29–33
Shahin H, Walsh T, Rayyan AA, Lee MK, Higgins J, Dickel D et al (2010) Five novel loci for inherited hearing loss mapped by SNP-based homozygosity profiles in Palestinian families. Eur J Hum Genet 18:407–413
Yang T, Wei X, Chai Y, Li L, Wu H (2013) Genetic etiology study of the non-syndromic deafness in Chinese Hans by targeted next-generation sequencing. Orphanet J Rare Dis 8:85
Ganapathy A, Pandey N, Srisailapathy CR, Jalvi R, Malhotra V, Venkatappa M et al (2014) Non-syndromic hearing impairment in India: high allelic heterogeneity among mutations in TMPRSS3, TMC1, USHIC, CDH23 and TMIE. PLoS One 9:e84773
Mori K, Moteki H, Miyagawa M, Nishio SY, Usami S (2016) Social health insurance-based simultaneous screening for 154 mutations in 19 deafness genes efficiently identified causative mutations in Japanese hearing loss patients. PLoS One 11:e0162230
Miyagawa M, Naito T, Nishio SY, Kamatani N, Usami S (2013) Targeted exon sequencing successfully discovers rare causative genes and clarifies the molecular epidemiology of Japanese deafness patients. PLoS One 8:e71381
Brownstein Z, Friedman LM, Shahin H, Oron-Karni V, Kol N, Abu Rayyan A et al (2011) Targeted genomic capture and massively parallel sequencing to identify genes for hereditary hearing loss in Middle Eastern families. Genome Biol 12(9):R89
Lebeko K, Sloan-Heggen CM, Noubiap JJ, Dandara C, Kolbe DL, Ephraim SS et al (2016) Targeted genomic enrichment and massively parallel sequencing identifies novel nonsyndromic hearing impairment pathogenic variants in Cameroonian families. Clin Genet 90:288–290
Woo HM, Park HJ, Park MH, Kim BY, Shin JW, Yoo WG et al (2014) Identification of cdh23 mutations in korean families. BMC Med Genet 15:46
Park JH, Kim NK, Kim AR, Rhee J, Oh SH, Koo JW et al (2014) Exploration of molecular genetic etiology for Korean cochlear implantees with severe to profound hearing loss and its implication. Orphanet J Rare Dis 9:167
Acknowledgements
The authors exclude any relevant financial activities outside the submitted work.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no conflict of interest. This research did not receive any specific grant from funding agencies in the public, commercial, or not-for-profit sectors.
Ethical approval
All procedures performed in this study involving human participants were in accordance with the ethical standards of the institutional and/or national research committee and with the 1964 Helsinki declaration and its later amendments or comparable ethical standards. There was no evaluation or vote of an Ethical Committee necessary because this study was designed as a meta-analysis.
Rights and permissions
About this article
Cite this article
Xu, T., Zhu, W. & Wang, P. The p.P240L variant of CDH23 and the risk of nonsyndromic hearing loss: a meta-analysis. Eur Arch Otorhinolaryngol 276, 11–16 (2019). https://doi.org/10.1007/s00405-018-5160-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00405-018-5160-8