Abstract
The Cape Fold Ecoregion (CFE) is one of southern Africa’s unique aquatic ecoregions and its freshwater fish fauna is characterized by high levels of endemism. As with many other Mediterranean-type ecosystems, the region is also a hotspot for threatened and range-restricted freshwater fish. Many of the CFE’s endemic species are at risk for extinction, with declines in population sizes and distribution ranges. The Clanwilliam sandfish Labeo seeberi is an example of such a species and is considered one of South Africa’s most endangered large migratory cyprinids. This species is endemic to the Olifants/Doring river system in the CFE and has been subject to a major population decline, mainly as a result of invasive alien fish and adverse climate events. Little is known of the genetics of the Clanwilliam sandfish, thus this study aimed to provide basic population genetic parameters to inform future conservation interventions. Both microsatellite and mitochondrial DNA (mtDNA) markers were used to assess populations from three sites within the Olifants/Doring river system. Genetic diversity was moderate to low and did not reflect the drastic decline expected on the basis of previous relative abundance data. This is likely due to a lag effect between ecological/life history demographics (due to juvenile recruitment failures) and population genetic composition. Furthermore, there was limited genetic differentiation between the sampling locations, suggesting a single breeding population, but mtDNA haplotype distribution and slight divergence of the smaller populations does suggest that the population might have become recently fragmented. The results show that the effective population size of the current breeding population might still be sufficient to maintain evolutionary potential in the short term, which could act as a buffer until conservation strategies focusing on protecting breeding animals and maximizing juvenile survival can restore population numbers.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Freshwater fishes are some of the most threatened animals globally. This is largely due to fragmentation, degradation, and loss of habitat; water flow modification and pollution; introduction of invasive alien species (as predators and competitors); the increasing threat of climate change; and overfishing (Reid et al. 2013; Dudgeon 2019; Jordaan et al. 2020). The Cape Floristic Region, located primarily within the Western Cape Province of South Africa, is one of the six floral kingdoms of the world and recognized as a global biodiversity hotspot (Myers et al. 2000). The geographical extent of the Cape Floristic Region corresponds to the Cape Fold Ecoregion (CFE), one of the eight aquatic ecoregions of southern Africa (Abell et al. 2008; Chakona et al. 2022). The CFE is moderately diverse in terms of freshwater fish taxa (n = 40), but a hotspot for freshwater fish endemism with 92% of these taxa being endemic to the region (Chakona et al. 2022). The number of endemic and threatened taxa continue to increase as a result of ongoing morphological and genetic studies (e.g., Swartz et al. 2007, 2009; Chakona et al. 2013a; Bronaugh et al. 2020) and confirms the suggestion by Linder et al. (2010) that the current taxonomy vastly underestimates the diversity of freshwater fishes of the region. In terms of the International Union for Conservation of Nature (IUCN) Red List status, 70% of freshwater fish taxa of the CFE are threatened, with five taxa listed as critically endangered, fourteen as endangered and nine as vulnerable (Chakona et al. 2022).
Alien invasive fish and habitat degradation has been highlighted as the major drivers for the declines in the freshwater fishes in the CFE (Tweddle et al. 2009; De Moor and Day 2013; Weyl et al. 2013, 2014; Chakona et al. 2022). Many populations have become fragmented and are now largely confined to the headwater reaches of streams (Swartz et al. 2004; Tweddle et al. 2009; Chakona et al. 2013b). Conservation management for the persistence and protection of the unique freshwater biodiversity in this region has thus become of increasing importance (Paxton et al. 2012). This is especially important in the case of freshwater fish, as the current protected area network was shown to be largely ineffective for protecting the majority of indigenous freshwater fish species of the region (Jordaan et al. 2020).
The Clanwilliam sandfish Labeo seeberi is a large cyprinid species endemic to the Olifants/Doring river system in the CFE. It favors pools or deep runs of larger rivers and migrates upstream into tributary habitat during spawning, which occurs during the austral spring (Paxton et al. 2002, 2012). Hontela and Stacey (1990) reported that the changes in water chemistry, brought on by heavy rain and subsequent flooding of terrestrial vegetation, are the ultimate spawning cue for cyprinids. Potts et al. (2005) reported that flooding has been recognized as the primary factor regulating spawning in the closely related Labeo umbratus. Poor rainfall can thus negatively affect spawning cues, causing female fish to retain their eggs until conditions are optimal or reabsorb eggs altogether if conditions do not improve, as was illustrated in L. umbratus (Gaigher 1984; Potts et al. 2005).
Labeo seeberi was historically widespread within the Olifants/Doring river system, but currently persists as a fragmented population confined to the middle and northern reaches of the Doring River and its isolated tributaries, namely the Oorlogskloof/Koebee, Gif, Kransgat, Biedouw, Tra-Tra, and Matjies rivers (Paxton et al. 2012; Lubbe et al. 2015). It has been estimated that the species has experienced a more than 90% decline in relative abundance since 2013. This is partly attributed to an estimated 99% decline in annual juvenile recruitment in one of the main spawning areas (Oorlogskloof River) as a consequence of aberrant weather patterns, highlighting the threat of climate change (Cerrilla et al. 2022). The species is currently listed as endangered (Jordaan et al. 2017) and while there is a relatively good understanding of threats to the species, there has not been any study of the current genetic status of the sandfish. This paucity of knowledge may preclude the effective identification and implementation of conservation strategies.
An understanding of the genetic composition of the population will assist in determining conservation units, the extent of genetic diversity and evolutionary potential of the species, and inform captive breeding and release strategies for population recovery and augmentation (Kardos 2021). Riverine environments are often characterized by significant population genetic structuring of aquatic animals due to the system’s unique in-stream features (natural and anthropogenic) and hydrodynamics that might act as barriers, restricting animal movement and subsequent gene flow (e.g., Abbas et al. 2010; Peacock et al. 2016; Coleman et al. 2018; Rougemont et al. 2021; Shelley et al. 2022). Using the COI mitochondrial DNA (mtDNA) region, Modeel et al. (2023) argued that there was little population genetic structure among Labeo rohita (Rohu) populations from south and southeast Asia. However, this mtDNA marker, commonly used as a species identifying barcoding gene due to its low mutation rate, is expected to show high homogeneity within a species (Bhattacharya et al. 2016). Other studies for L. rohita, (based on nuclear microsatellite markers and the mtDNA marker, cyt b) have presented mixed results, with evidence for limited genetic differentiation and significant genetic divergence between populations depending on the river systems (Luhariya et al. 2012; Sahoo et al. 2014; Behera et al. 2018). Other Indian Labeo species, including Labeo dero (Chaturvedi et al. 2011) and Labeo fimbriatus (Swain et al. 2022) also demonstrated evidence for population genetic stratification. Across the various Asian Labeo species, population genetic diversity seems to be wide-ranging depending on the conservation and/or commercial exploitation status of the species.
African Labeo species form a distinct clade from the Asian species (Ramoejane 2016; Kebede and Harris 2019), and there are limited studies on population genetic diversity and structure for these species. However, an extensive study using mtDNA and nuclear gene sequences on L. umbratus, a closely related species to L. seeberi, does suggest that significant population genetic structure could persist within African Labeo species on a broad spatial scale (Ramoejane et al. 2021). However, unlike L. umbratus,, which has a fairly large distribution range across various river systems (Ramoejane 2016), L. seeberi has a restricted distribution within the Olifants/Doring river system. Other cyprinid species within the Olifants/Doring river system seem to exhibit varying degrees of genetic differentiation (Swartz et al. 2004). Hence, the genetic constitution of species, even if restricted to a single river system, might be highly dependent on species-specific life history characteristics and adaptations, breeding behavior, and dispersal ability. The aim of the study was therefore to investigate population genetic parameters, using microsatellite and mtDNA markers, to gain insight into the population dynamics, genetic health, and status of the remaining L. seeberi population to inform optimal conservation strategies.
Materials and methods
Sample collection and DNA extraction
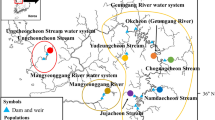
The bulk of the sample collection was done by staff from relevant provincial conservation agencies (CapeNature and the Northern Cape Department Nature and Environment Conservation) while conducting routine freshwater fish surveys. Additional samples were collected and donated by staff from the Endangered Wildlife Trust (permit number AAA008-00022). In total, samples were collected from 128 adult fish (> 200 mm) that were sampled from three different localities, two of which were located on the Oorlogskloof River. The upstream site is located within the Oorlogskloof Nature Reserve (n = 82, OKNR), while the downstream site is located on Rietkuil Farm at the confluence of the Oorlogskloof and Klein/Kobee rivers (n = 36, Riet). These two sites are geographically close, but functionally separated by a waterfall that is a complete barrier to any upstream fish movement. Additional samples were collected from the Bos River (n = 10, Bos), a tributary of the Doring River. All samples were taken during a 2013 survey of the greater Doring River system; these sampling locations presented the only sites along the river system where the fish was found during the survey (Fig. 1; Table S1).
Map of the Olifants/Doring river system in the Northern and Western Cape of South Africa. The Clanwilliam sandfish is restricted to the northern reaches of the Doring River with recruitment restricted to the Oorlogskloof/Koebee River tributary. Dots indicate sampling locations. The upstream site (above the waterfall barrier) is located within the Oorlogskloof Nature Reserve (n = 82, OKNR), the downstream Rietkuil site is located at the confluence of the Oorlogskloof and Klein/Kobee rivers (n = 36, Riet), and the Bos River (n = 10, Bos) is a tributary of the Doring River
Fish were captured using a combination of sein nets and large fyke nets. Fin clips measuring a maximum size of 0.5 cm × 0.5 cm were collected from one of the paired anal fins of each individual and stored in a 2.5 ml tube filled with 100% ethanol. All fish were released back at their site of capture post sample collection. Samples were stored at 4 °C until DNA extractions could be performed using an adjusted cetyl trimethylammonium bromide (CTAB) protocol as described by Justesen et al. 2002.
Microsatellite loci and mitochondrial control region amplification
Six microsatellite markers were sourced from literature based on two sister species, Labeo rohita (Patel et al. 2011) and Labeo fimbriatus (Swain et al. 2012, 2013) (Table S2) and used to genotype the fish from three sampling locations. The polymerase chain reaction (PCR) conditions were as follow: final reaction volume of 10 μl consisted of 1× KAPA Taq Ready Mix (Roche, Basel, Switzerland), 100 ng of DNA and 0.4 μM of each primer. Cycling conditions included an initial denaturation phase at 95 °C for 5 min, followed by 30 cycles of 95 °C for 45 s, 1 min at the annealing temperature (TA; Table S1), 72 °C for 2 min, with a final extension step at 72 °C for 7 min. Amplified samples were submitted to the Central Analytical Facility (CAF), Stellenbosch University for capillary electrophoresis on a 3730XL Genetic Analyzer (Life Technologies, Carlsbad, California, USA). GeneMapper v4.0 (Life Technologies) was used for binning, allele scoring, and manually curated where needed.
A total of 36 individuals were randomly subsampled to represent the three sampling locations (OKNR—13; Riet—13; Bos—10) for the amplification of the mitochondrial control region (mtCR). A 736 bp fragment of the mtCR was PCR amplified using the primer pair by Lin et al. (2010): Fish G (F): 5′-GCATGGGTCTTGTAATCCGA-3′ and Fish F (R): 5′-TAGTAAGGTCGGGACCATGC-3′, with a total reaction volume of 20 µl Kappa Taq Ready Mix (Roche), using the same reaction composition as for the microsatellite markers. The thermal cycling conditions of these reactions are as follows: an initial denaturation step at 95 °C for 3 min, followed by 35 cycles at 95 °C for 30 s, an annealing temperature of 57 °C for 30 s, 72 °C for 1 min, and a final extension step at 72 °C for 7 min. PCR products were run on a 1.5% agarose gel at 100 V for 30 min to confirm successful amplification, after which the products were sent to CAF, Stellenbosch University for sample purification using the QIAGEN gel clean-up system and then sequenced using Sanger sequencing chemistry (BigDye® termninator v3.1 cycle sequencing kit, Life Technologies). Sequencing products were purified using Sephadex spin columns (Princeton Separation, Adelphia, NJ) and analyzed via capillary electrophoresis on a 3730XL Genetic Analyzer (Life Technologies). Sequences were aligned using the ClustalW algorithm in MEGA v7 (Kumar et al. 2018), manually edited and trimmed to equal lengths.
Genetic diversity, relatedness, and population demographic analyses
Microsatellite genotypes were evaluated for stuttering, allelic dropout, and the presence of null alleles, with the frequency of null alleles per locus per sampling location calculated with Microchecker v2.2.3 (Van Oosterhout et al. 2004). Genepop on the web v4.2 (Rousset 2008) was used to test for between-loci linkage disequilibrium (LD), within and across sampling locations, as well as for loci deviating from Hardy–Weinberg equilibrium (HWE) expectations (10,000 dememorizations, 100 batches, and 10,000 iterations per batch). Genetic diversity indices were calculated for each sampling location, which included: the average number of alleles per locus (AN), observed and unbiased expected heterozygosities (HO and uHE), Shannon’s information index (I), and fixation indices (FIS), as well as standard error (SE) for each mean, in GenAlEx v6.501 (Peakall and Smouse 2012). Polymorphism information content (PIC) was calculated using MSTools v3.0 (Park 2001). The rarefied allelic richness per locus (AR) was estimated using HP-Rare (Kalinowski 2005). The non-parametric Kruskal–Wallis test was performed in XL Statistics v2016.5 (https://software.deakin.edu.au/2017/03/24/xlstatistics) to evaluate the significance (P < 0.05) of differences in genetic diversity estimates between sampling locations. Mean relatedness per population (i.e., sampling location) was calculated in GenAlEx, using the relatedness estimator, r (Queller and Goodnight 1989). Significance testing by 999 bootstrap replicates for differences between populations was done.
Effective population sizes (Ne) were estimated for each sampling location, using the linkage disequilibrium method as implemented in NeEstimator v2.0.1 (minimum allele frequency of 0.02, assuming a random mating model) (Do et al. 2014), with significance tests set at upper and lower 95% confidence intervals based on bootstrapping. To test for recent bottlenecks, a Wilcoxon signed rank test (Luikart et al. 1998) was performed in Bottleneck v1.2.02 (Piry et al. 1999). Three mutational models were implemented, specifically the infinite allele model (IAM), stepwise mutation model (SMM), and the two-phase model (TPM). Analyses were done using 10,000 replications at the 5% nominal level.
Mitochondrial DNA diversity was determined in DnaSP v5.0 (Librado and Rozas 2009), including the total number of haplotypes (H), hapotype diversity (h), and nucleotide diversity (π) for all sampling locations. The evolutionary relationships among haplotypes was inferred by constructing a median-joining inference network (Bandelt et al. 1999) as implemented in Network v5.0.0.3 (https://www.flexus-engineering.com).
Genetic structuring analyses
The following microsatellite analyses were performed to assess genetic differentiation between the sampling locations, pairwise FST estimates, analysis of molecular variance (AMOVA), and multivariate discriminant analysis of principal components (DAPC). Pairwise FST and an AMOVA were performed (at significance level P < 0.05) in Arlequin v3.5 (Excoffier and Lischer 2010). Finally, a DAPC plot was constructed using the R packages ade4 and adegenet (Jombart 2008). Prior to running the DAPC, cross-validation was done to determine the optimal number of principal components (PCs) to retain that allowed for the most accurate assignment of individuals (> 80%) to specific genetic clusters. For the mtCR, genetic differentiation among sampling locations was determined in Arlequin, by means of pairwise ɸST estimates and AMOVA (P < 0.05). Additionally, the nucleotide substitution model for the alignment was determined in jModelTest v2.0 (Darriba et al. 2012), with the HKY + G model being the best fit for the dataset. A maximum likelihood (ML) phylogenetic tree was then constructed in MEGA v7 and was bootstrapped with 1000 runs, assuming this substitution model.
Results
Genetic diversity
Microsatellite diversity
A total of 128 individuals were successfully genotyped across six loci, with the average number of alleles ranging from 2 to 23 per maker (Table S3 for detail on per marker diversity across sampling locations). Across all sampling locations, two markers deviated from HWE (Table S3), and no significant linkage disequilibrium was observed. However, locus Lr_41 displayed evidence for null alleles and deviated from HWE in all populations. Since no significant differences were observed when the locus was excluded from analyses and given the limited number of loci available for analyses, the locus was ultimately retained for all downstream analyses.
Overall, the number of alleles per sampling location was moderate to low (Table 1), ranging from 6.50 (Bos) to 11.17 (OKNR). However, allelic richness was more comparable between the sampling locations (Table 1). Of the three localities, the Bos sample displayed the lowest number of alleles, observed, and unbiased expected heterozygosities (AN—6.50; HO—0.583; uHE—0.043; Table 1). Additionally, the inbreeding coefficients of all three groups were not statistically significantly different from one another, and the low values point to less inbred populations of L. seeberi, and none of the populations deviated significantly from HWE on the whole. Low levels of inbreeding were further supported by the mean relatedness (r) estimates that did not deviate significantly from zero for all populations (Fig. 2).
Estimates of mean relatedness for each of the sampling locations of Labeo seeberi. Error bars represent standard error of the mean. Upper (U) and lower (L) 95% confidence intervals for differences among the populations [Oorlogskloof Nature Reserve (n = 82, OKNR); Rietkuil (n = 36, Riet); and the Bos River (n = 10, Bos)]
Mitochondrial diversity
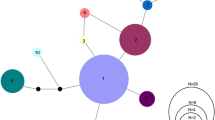
A 736 bp fragment of the mtCR was amplified in 37 individuals. Analysis resulted in eleven polymorphic sites, which consisted of ten transitions and one transversion. A total of ten haplotypes were identified, comprised of three high frequency haplotypes (H1, 16.20%; H3, 27.03%; H4, 32.43%; Fig. 3). Notably, haplotype 1 was absent from the Riet population, while the lower frequency haplotypes appeared to be unique to specific populations. Overall, mitochondrial diversity was relatively high, with haplotype diversity (h) and nucleotide diversity (π) ranging from 0.782 to 0.821 and from 0.00233 to 0.00300, respectively (Table 1).
Median-joining haplotype network of L. seeberi mitochondrial control region haplotypes. Haplotypes are separated by black lines, with black dots indicating mutated positions between haplotypes. The size of the haplotype circles is proportional to the number of individuals possessing said haplotype. A total of ten haplotypes were identified, composed of three high frequency haplotypes (H1, 16.20%; H3, 27.03%; H4, 32.43%). The Bos River (Bos; blue, n = 10); Oorlogskloof Nature Reserve (OKNR; orange, n = 82); Rietkuil (Riet; green, n = 36) (color figure online)
Population differentiation
Nuclear structure
Pairwise FST estimates were low, ranging from 0.002 to 0.007, with no pairwise comparisons reaching statistical significance at the 1% nominal level (Table 2), indicating little to no genetic differentiation between the sampling locations. Furthermore, the AMOVA (Table 3) supported the lack of genetic differentiation with less than 1% of variation explained by among populations differences (global FST = 0.005, P > 0.05). While the DAPC plot reveals some separation of the Bos population from the OKNR and Rietkuil populations, this is most likely a result of few samples from the Bos area (Fig. 4).
Multivariate discriminant analysis of principal components (DAPC) plot for Labeo seeberi using six microsatellite markers, with sampling locations represented by different symbols and colors [Oorlogskloof Nature Reserve (OKNR; blue triangle, n = 82); Rietkuil (Riet; yellow diamond, n = 36); and the Bos River (Bos; red circle, n = 10)] (color figure online)
Mitochondrial structure
Pairwise ɸST analyses revealed no significant differentiation between the populations, (Table 2), with additional support from the AMOVA results (ɸST = 0.045, P > 0.05; Table 3). The majority of the variance was the diversity of haplotypes within each population contributing 94.45% of the total variance (Table 3). Furthermore, the maximum likelihood tree supports this lack of structure, with no discernible formation of groups within the entirety of the dataset (Fig. 5).
Maximum likelihood tree of the Labeo seeberi populations based on a fragment of the mitochondrial control region. Numbers at nodes indicate bootstrap support for the placement of individuals (only nodes that reached a bootstrap value higher than 50% are indicated), with the tree rooted using a representative control region sequence from a sister species, Labeo fimbriatus
Population demography
The infinite allele model (IAM) showed statistically significant (P < 0.05; Table 4) evidence of recent population bottlenecks, although this was not supported by the two-phase model (TPM) or the stepwise mutational model (SMM).
Estimations of effective population size (Ne) using the linkage disequilibrium method indicated very low estimates for the lower 95% confidence interval for the Riet and Bos populations, while the OKNR population was higher (Table 4). However, all point estimates using the linkage disequilibrium method were infinite, which is likely a result of using few markers.
Discussion
The cumulative threats present in the CFE have resulted in a decline in both numbers and distribution in almost all of the endemic fish species of the region, including the Olifants/Doring river system (Paxton et al. 2002; Van der Walt et al. 2016; Chakona et al. 2022). These threats include loss of habitat as a result of water overabstraction, water infrastructure such as dams disrupting migration pathways, predatory invasive fish species, and the destruction of riparian zones. This causes fish populations to become fragmented, thereby reducing the population sizes and potentially serving as barriers to gene flow (Paxton et al. 2012; Lubbe et al. 2015; Chakona et al. 2020). Small, isolated populations are intrinsically vulnerable to the loss of genetic diversity through genetic drift and further homogenization of the population through inbreeding (Palstra and Ruzzante 2008; Hare et al. 2011; Chakona et al. 2020). If unchecked, this could impair the evolutionary potential and adaptive capacity of a species, decreasing fitness and leading to eventual extinction (Frankham 2005; Charlesworth and Willis 2009; Hare et al. 2011). This study provides some insight into the population genetics of the disjunct populations of the Clanwilliam sandfish.
Genetic diversity at the microsatellite loci was moderate to low (Table 1) as has previously been reported by Sahoo et al. (2014) and Singh et al. (2012) for Labeo rohita and Labeo calbasu, respectively. The number of optimized markers and marker utility evaluation was also similar to Mohindra et al. (2005) who tested microsatellite marker transfer to Labeo dyocheilus. This suggested the utility of the cross-species markers that are often criticized for creating ascertainment bias leading to underestimation of genetic parameters in relation to the source species (Barbara et al. 2007). The Bos population did show marginally reduced estimates of diversity, most likely a consequence of the limited sample size, in comparison with the OKNR and Riet populations. However, AR correcting for unbalanced sampling does suggest that nuclear genetic diversity was similar across sampling locations (Kalinowski 2005). There was a slight homozygous excess for all three sampling locations, with evidence for limited inbreeding (significant FIS and FIT from the AMOVA) but was negligible, as the general population relatedness was practically zero (Fig. 2). This homozygous excess could also be explained by possible null alleles. Alam et al. (2009) and Sahoo et al. (2014) reported very similar diversity estimates for Labeo rohita using microsatellite markers. Nonetheless, the genetic data is indicating a population at the initial phases of decline with some loss of genetic diversity (Leberg 2002; Foulley and Ollivier 2006), further supported by evidence for a recent genetic bottleneck under the infinite alleles model (Table 4). This could be attributed to the impacts of invasive black bass (Micropterus spp.) that have become established in more than 80% of the river system, with deleterious consequences for native fish biota, resulting in the extirpation of most indigenous fishes from invaded river reaches (Van der Walt et al. 2016). Larger bodied species such as the sandfish persist in these habitats, but at low numbers and in the form of relatively isolated populations. Unusual and erratic climate events, associated with climate change, altering waterscapes, and impacting on fish breeding behavior and juvenile recruitment can also not be excluded (Cerrilla et al. 2022).
Interestingly, estimates of effective population size (Ne) remained high across all three sampling locations. However, the point estimates, based the confidence intervals are imprecise, likely due to the small number of microsatellite markers and limited sample sizes; some caution in the interpretation of the values are thus needed (Waples et al. 2016). This is a common problem for species of conservation concern due to the scarcity and high value of animals (that limits sampling opportunities) and limited genomic resources for an understudied organism that further compounds the estimation of a parameter well known to be difficult to determine and is dependent on various organismal life-history characteristics and demographics (Serbezov et al. 2012). Nonetheless, it has been argued that despite large confidence intervals for estimates of Ne, the LD method remains robust, and that lower confidence bound could still provide a fair assessment for making conservation management judgements (Waples and Do 2010). In their revised recommendations, Frankham et al. (2014) suggested to increase the lower limit for Ne from 50 to 100 in aim of mitigating the immediate effects of inbreeding, and to 500 to ensure at least short-term protection for loss of fitness over the next five generations. Looking at the lower bound Ne estimate for the OKNR population (Table 4), which is considered one of the last major reproducing populations of sandfish, it is already at that 500 cusp. The other two populations seem to have significantly reduced Ne with the Riet population at about the 100 point and the Bos population in critical danger of local extinction with an Ne less than 50. While there is no geographical barrier separating the Riet and the Bos localities, it is likely that the established black bass population in the system has reduced survival of juvenile sandfish to negligible levels, which has effectively fragmented these two populations. Wang et al. (2019) argued that minimum viable population size (MVP) in freshwater fish can be highly variable and that long lived fish with late sexual maturity, long generation intervals, and high fecundity typically require larger MVPs. The authors estimated an upper bound for such species at an MVP of 320 individuals. It has been suggested that MVP should be at least five to ten times greater than the Ne (Frankham 1995; Rosenfeld 2014) and this fits sandfish relative abundance data (Cerrilla et al. 2022). These estimates are also congruent with the patterns of genetic diversity (Baek et al. 2018; Coleman et al. 2018; Martinez et al. 2018).
Cerrilla et al. (2022) documented a significant and drastic decline of more than 92% in the relative abundance of Clanwilliam sandfish in the Oorlogskloof River between 2013 and 2018. This decline was largely driven by a more than 99% decline in young of year individuals, which in turn was attributed to severe weather events including prolonged drought spells. The trend of this severe decline is not reflected in the genetic data, especially for the OKNR population. This is primary because the authors attribute the decline to recruitment failure among the young-of-year, as catch per unit effort for adult fish has remained fairly consistent over the study period. The generation interval is not known for Clanwilliam sandfish, but the closely related L. umbratus was reported to reach sexual maturity at 3 and 4 years, respectively, for males and females (Mulder 1973). Estimates based on simulations for other Labeo species, L. niloticus, L. roita, and L. coubie, also suggest that they have fairly long generational times, between 3 and 5 years (www.fishbase.se). Therefore, as genetic analyses are particularly sensitive to changes in the genetic composition of populations over generations (Serbezov et al. 2012), the genetic consequences of the population contraction remain “hidden” if new members are not recruited into the breeding population to form the next generation. This creates a known lag time between ecological and life history demographics of populations and its corresponding genetic composition and structure (Epps and Keyghobadi 2015). This may also explain the relatively high observed mitochondrial diversity (Table 1) that tend to reflect more historical population dynamics versus the microsatellite data that gives a contemporary analysis (relative to the lag time) (Wan et al. 2004).
Given the critical interplay between Ne, genetic diversity, evolutionary potential, and fitness (Reed and Frankham 2003; Ellegren and Galtier 2016), it is worth noting that there could be considerable differences between short- and long-term Ne, especially for animals with mass spawning and r-selected reproductive strategies (Martinez et al. 2018; Barry et al. 2022). Their reproductive life history characteristics are defined by large variances in breeding success among individuals and mass larval/juvenile mortality rates that leads to reduced Ne in the short-term (Rhode et al. 2017; Martinez et al. 2018; Monteiro et al. 2022). These animals might therefore have some natural resilience to short term larval recruitment failures, as long as the breeding population remains genetically stable and long-lived. Also, high fecundity leads to high mutation rates, that could replenish lost diversity and bolster evolutionary potential relatively quickly, in comparison with animals with k-selected strategies, with effective conservation management (e.g., restoring gene flow, and ensuring equal reproductive success of all breeding animals) and restoration of the natural habitat (Frankham 2015; Ellegren and Galtier 2016; Pavlova et al. 2017; Martinez et al. 2018; Prunier et al. 2023). Given the sampling strategy of this study (only adult fish were sampled) and the stable abundance data for adult fish (from Cerrilla et al. 2022), the current genetic analyses suggest that there is a genetically stable, mature population of Clanwilliam sandfish, with an Ne (approximately 500) that could provide short- to mid-term evolutionary potential and act as a buffer until the natural replenishment of genetic diversity (Martinez et al. 2018). However, it may be concerning that this population is potentially aging, due to the lack of juvenile recruits (Cerrilla et al. 2022). Conservation strategies should thus focus on protecting breeding animals and natural spawning sites and maximizing juvenile survival and broodstock contributions, perhaps through a captive breeding and restocking program or through head-starting initiatives aimed at increasing juvenile survival. Care should, however, be taken to reduce the unintentional impact of hatchery effects leading to genetic erosion (Klütsch et al. 2019; Monteiro et al. 2022).
Neither microsatellite nor mitochondrial data could support sufficient evidence for genetic differentiation within or between the three sampling locations (Tables 2, 3, Fig. 5). However, there was some minor separation of the Riet and Bos population from OKNR and each other, observable on the DAPC plot (Fig. 4). This is likely a sampling artifact of the small populations. But, considering several private mitochondrial haplotypes, and the absence of a high frequency haplotype from the Riet population (H1, Fig. 3), it might suggest that these populations were recently separated. A large waterfall on the Oorlogskloof River serves as a barrier to upstream fish movement, but prior to the establishment of black bass throughout the system juvenile fish from above the waterfall would have migrated downstream toward the mainstream Doring River. At present sandfish do persist downstream of the waterfall but the population is composed almost exclusively of large and old fish. It is thus recommended that all isolated populations of sandfish be managed as a single unit for conservation to maximize population viability and simplify conservation strategies and planning (Funk et al. 2012; Frankham 2015; Pavlova et al. 2017; Coates et al. 2018).
Conclusion
Labeo seeberi is one of the most threatened large cyprinids in southern Africa and is endemic to the Olifants/Doring river system in the CFE. It has been extirpated from much of its historical distribution range and is presently listed as endangered. Contemporary estimates of census population size suggest a more than 90% decline in population numbers in one of the main breeding tributaries, the Oorlogskloof River. Population declines for the greater Doring River system is difficult to quantify given the paucity of monitoring data, but is believed to be severe (Jordaan et al. 2017; Paxton et al. unpublished data). However, the genetic data does not reflect this drastic decline, most likely due to a time-lag between ecological/life history demographics and genetic structure. Nonetheless, the current breeding population, which consists of a single population across three sampling locations, does seem to harbor sufficient genetic diversity to serve as a reservoir for evolutionary potential in the short term (five generations). Conservation strategies must therefore focus on protecting breeding animals and maximizing juvenile survival. Given the high fecundity of the species, there is scope for the “self-restoration” of lost genetic diversity if conservation strategies are successful in securing juvenile recruitment. Conservation interventions that should be implemented with urgency include habitat restoration through management of water abstraction in sensitive catchments and the targeted removal of alien invasive piscivorous fish in priority tributaries where sandfish are known to breed. Removal of undesirable predatory fishes generally require the construction of instream barriers to prevent reinvasion, but this should be implemented within the context of the migratory nature of sandfish, which may prove challenging.
Data availability
All data generated in this project was based on previously published protocols and resources and is referenced appropriately either in the main text or supplementary data. The microsatellite genotypic data (Supplementary file 2) and mtCR sequence alignment file (Supplementary file 3) generated and analyzed during the current study are available as supplementary data.
Change history
15 April 2024
A Correction to this paper has been published: https://doi.org/10.1007/s00027-024-01078-7
References
Abbas K, Zhou X, Li Y, Gao Z, Wang W (2010) Microsatellite diversity and population genetic structure of yellowcheek, Elopichthys bambusa (Cyprinidae) in the Yangtze River. Biochem Syst Ecol 38(4):806–812
Abell R, Thieme ML, Revenga C, Bryer M, Kottelat M, Bogutskaya N, Coad B, Mandrak N, Balderas SC, Bussing W, Stiassny ML (2008) Freshwater ecoregions of the world: a new map of biogeographic units for freshwater biodiversity conservation. Bioscience 58(5):403–414. https://doi.org/10.1641/B580507
Alam MS, Jahan M, Hossain MM, Islam MS (2009) Population genetic structure of three major river populations of rohu, Labeo rohita (Cyprinidae: Cypriniformes) using microsatellite DNA markers. Genes Genomics 31:43–51. https://doi.org/10.1007/BF03191137
Baek SY, Kang JH, Jo SH, Jang JE, Byeon SY, Wang JH, Lee HG, Choi JK, Lee HJ (2018) Contrasting life histories contribute to divergent patterns of genetic diversity and population connectivity in freshwater sculpin fishes. BMC Evol Biol 18:1–14. https://doi.org/10.1186/s12862-018-1171-8
Bandelt HJ, Forster P, Rohl A (1999) Median-joining networks for inferring intraspecific phylogenies. Mol Biol Evol 16(1):37–48. https://doi.org/10.1093/oxfordjournals.molbev.a026036
Barbara T, Palma-Silva C, Paggi GM, Bered F, Fay MF, Lexer C (2007) Cross-species transfer of nuclear microsatellite markers: potential and limitations. Mol Ecol 16(18):3759–3767. https://doi.org/10.1111/j.1365-294x.2007.03439.x
Barry P, Broquet T, Gagnaire PA (2022) Age-specific survivorship and fecundity shape genetic diversity in marine fishes. Evol Lett 6(1):46–62. https://doi.org/10.1002/evl3.265
Behera BK, Baisvar VS, Kunal SP, Meena DK, Panda D, Pakrashi S, Paria P, Das P, Bhakta D, Debnath D, Roy S (2018) Population structure and genetic diversity of Indian Major Carp, Labeo rohita (Hamilton, 1822) from three phylo-geographically isolated riverine ecosystems of India as revealed by mtDNA cytochrome b region sequences. Mitochondrial DNA Part A 29(2):199–205. https://doi.org/10.1080/24701394.2016.1267156
Bhattacharya M, Sharma AR, Patra BC, Sharma G, Seo EM, Nam JS, Chakraborty C, Lee SS (2016) DNA barcoding to fishes: current status and future directions. Mitochondrial DNA Part A 27(4):2744–2752. https://doi.org/10.3109/19401736.2015.1046175
Bronaugh WM, Swartz ER, Sidlauskas BL (2020) Between an ocean and a high place: coastal drainage isolation generates endemic cryptic species in the cape kurper Sandelia capensis (Anabantiformes: Anabantidae), cape region, South Africa. J Fish Biol 96(5):1087–1099. https://doi.org/10.1111/jfb.14182
Cerrilla C, Afrika J, Impson D, Jordaan MS, Kotze N, Paxton BR, Reed C, Schumann M, van der Walt JA, Shelton JM (2022) Rapid population decline in one of the last recruiting populations of the endangered Clanwilliam sandfish (Labeo seeberi): the roles of climate change and non-native fish. Aquat Conserv Mar Freshw Ecosyst 32:781–796. https://doi.org/10.1002/aqc.3785
Chakona A, Swartz ER, Gouws G, Bloomer P (2013a) A freshwater fish defies ancient mountain ranges and drainage divides: extrinsic and intrinsic influences on the evolutionary history of a recently identified galaxiid. J Biogeogr 40:1399–1412. https://doi.org/10.1111/jbi.12104
Chakona A, Swartz ER, Gouws G (2013b) Evolutionary drivers of diversification and distribution of a southern temperate stream fish assemblage: testing the role of historical isolation and spatial range expansion. PLoS ONE 8(8):e70953. https://doi.org/10.1371/journal.pone.0070953
Chakona A, Gouws G, Kadye WT, Jordaan MS, Swartz ER (2020) Reconstruction of the historical distribution ranges of imperilled stream fishes from a global endemic hotspot based on molecular data: implications for conservation of threatened taxa. Aquat Conserv Mar Freshw 30(1):144–158. https://doi.org/10.1002/aqc.3251
Chakona A, Jordaan MS, Raimondo DC, Bills RI, Skelton PH, Van der Colff D (2022) Diversity, distribution and extinction risk of native freshwater fishes of South Africa. J Fish Biol 100(4):1044–1061. https://doi.org/10.1111/jfb.15011
Charlesworth D, Willis JH (2009) The genetics of inbreeding depression. Nat Rev Genet 10:783–796. https://doi.org/10.1038/nrg2664
Chaturvedi A, Mohindra V, Singh RK, Lal KK, Punia P, Bhaskar R, Mandal A, Narain L, Lakra WS (2011) Population genetic structure and phylogeography of cyprinid fish, Labeo dero (Hamilton, 1822) inferred from allozyme and microsatellite DNA marker analysis. Mol Biol Rep 38:3513–3529. https://doi.org/10.1007/s11033-010-0462-y
Coates DJ, Byrne M, Moritz C (2018) Genetic diversity and conservation units: dealing with the species-population continuum in the age of genomics. Front Ecol Evol 6:165. https://doi.org/10.3389/fevo.2018.00165
Coleman RA, Gauffre B, Pavlova A, Beheregaray LB, Kearns J, Lyon J, Sasaki M, Leblois R, Sgro C, Sunnucks P (2018) Artificial barriers prevent genetic recovery of small isolated populations of a low-mobility freshwater fish. Heredity 120(6):515–532. https://doi.org/10.1038/s41437-017-0008-3
Darriba D, Taboada GL, Doallo R, Posada D (2012) jModelTest 2: more models, new heuristics and parallel computing. Nat Methods 9(8):772–772. https://doi.org/10.1038/nmeth.2109
De Moor FC, Day JA (2013) Aquatic biodiversity in the mediterranean region of South Africa. Hydrobiologia 719(1):237–268. https://doi.org/10.1007/s10750-013-1488-7
Do C, Waples RS, Peel D, Macbeth GM, Tillett BJ, Ovenden JR (2014) NeEstimator v2: re-implementation of software for the estimation of contemporary effective population size (Ne) from genetic data. Mol Ecol Resour 14:209–214. https://doi.org/10.1111/1755-0998.12157
Dudgeon D (2019) Multiple threats imperil freshwater biodiversity in the Anthropocene. Curr Biol 29(19):R960–R967. https://doi.org/10.1016/j.cub.2019.08.002
Ellegren H, Galtier N (2016) Determinants of genetic diversity. Nat Rev Genet 17(7):422–433. https://doi.org/10.1038/nrg.2016.58
Epps CW, Keyghobadi N (2015) Landscape genetics in a changing world: disentangling historical and contemporary influences and inferring change. Mol Ecol 24(24):6021–6040. https://doi.org/10.1111/mec.13454
Excoffier L, Lischer HEL (2010) Arlequin suite ver 3.5: a new series of programs to perform population genetics analyses under Linux and Windows. Mol Ecol Res 10:564–567. https://doi.org/10.1111/j.1755-0998.2010.02847.x
Foulley JL, Ollivier L (2006) Estimating allelic richness and its diversity. Livest Sci 101(1–3):150–158. https://doi.org/10.1016/j.livprodsci.2005.10.021
Frankham R (1995) Effective population size/adult population size ratios in wildlife: a review. Genet Res 66(2):95–107. https://doi.org/10.1017/S0016672300034455
Frankham R (2005) Genetics and extinction. Biol Conserv 126:131–140. https://doi.org/10.1016/j.biocon.2005.05.002
Frankham R (2015) Genetic rescue of small inbred populations: meta-analysis reveals large and consistent benefits of gene flow. Mol Ecol 24:2610–2618. https://doi.org/10.1111/mec.13139
Frankham R, Bradshaw CJA, Brook BW (2014) Genetics in conservation management: revised recommendations for the 50/500 rules, Red List criteria and population viability analyses. Biol Conserv 170:56–63. https://doi.org/10.1016/j.biocon.2013.12.036
Funk WC, Mckay JK, Hohenlohe PA, Allendorf FW (2012) Harnessing genomics for delineating conservation units. Trends Ecol Evol 27:489–496. https://doi.org/10.1016/j.tree.2012.05.012
Gaigher IG (1984) Reproduction of Labeo umbratus (Pisces, Cyprinidae) in Wuras Dam, a shallow, turbid impoundment. S Afr J Zool 19:105–108. https://doi.org/10.1080/02541858.1984.11447866
Hare MP, Nunney L, Schwartz MK, Ruzzante DE, Burford M, Waples RS, Reugg K, Palstra F (2011) Understanding and estimating effective population size for practical application in marine species management. Conserv Biol 25(3):438–449. https://doi.org/10.1111/j.1523-1739.2010.01637.x
Hontela A, Stacey NE (1990) Cyprinidae. In: Munro AD, Scott AP, Lam TT (eds) Reproductive seasonality in teleosts: environmental influences. CRC Press, Boca Raton, pp 53–78
Jombart T (2008) adegenet: a R package for the multivariate analysis of genetic markers. Bioinformatics 24(11):1403–1405. https://doi.org/10.1093/bioinformatics/btn129
Jordaan MS, Chakona A, Van der Colff D (2020) Protected areas and endemic freshwater fishes of the Cape Fold Ecoregion: missing the boat for fish conservation? Front Environ Sci 8:502042. https://doi.org/10.3389/fenvs.2020.502042
Jordaan M, Lubbe A, Bragg C, Paxton BR, Schumann M, Van der Walt R, Ngobela T (2017) Labeo seeberi (errata version published in 2020). The IUCN Red List of threatened species 2017, p e.T11071A174778890. https://doi.org/10.2305/IUCN.UK.2017-3.RLTS.T11071A174778890.en
Justesen AF, Ridout CJ, Hovmøller MS (2002) The recent history of Puccinia striiformis f. sp. tritici in Denmark as revealed by disease incidence and AFLP markers. Plant Pathol 51(1):13–23. https://doi.org/10.1046/j.0032-0862.2001.00651.x
Kalinowski S (2005) HP-RARE 1.0: a computer program for performing rarefaction on measures of allelic richness. Mol Ecol Notes 5:187–189. https://doi.org/10.1111/j.1471-8286.2004.00845.x
Kardos M (2021) Conservation genetics. Curr Biol 31(19):R1185–R1190. https://doi.org/10.1016/j.cub.2021.08.047
Kebede AB, Harris PM (2019) New insight into the phylogeny and biogeography of the Cyprinid fishes Labeo (Cyprinidae; Cypriniformes) in Africa with evidence for cryptic diversity in Ethiopia. Ethiop J Biol Sci 18(2):153–178
Klütsch CF, Maduna SN, Polikarpova N, Forfang K, Aspholm PE, Nyman T, Eiken HG, Amundsen PA, Hagen SB (2019) Genetic changes caused by restocking and hydroelectric dams in demographically bottlenecked brown trout in a transnational subarctic riverine system. Ecol Evol 9(10):6068–6081. https://doi.org/10.1002/ece3.5191
Kumar S, Stecher G, Li M, Knyaz C, Tamura K (2018) MEGA X: molecular evolutionary genetics analysis across computing platforms. Mol Biol Evol 35(6):1547. https://doi.org/10.1093/molbev/msy096
Leberg PL (2002) Estimating allelic richness: effects of sample size and bottlenecks. Mol Ecol 11(11):2445–2449. https://doi.org/10.1046/j.1365-294x.2002.01612.x
Librado P, Rozas J (2009) DnaSP v5: a software for comprehensive analysis of DNA polymorphism data. Bioinformatics 25:1451–1452. https://doi.org/10.1093/bioinformatics/btp187
Lin CJ, Lin HD, Wang JP, Chao SC, Chiang TY (2010) Phylogeography of Hemibarbus labeo (Cyprinidae): secondary contact of ancient lineages of mtDNA. Zoo Scr 39(1):23–35. https://doi.org/10.1111/j.1463-6409.2009.00407.x
Linder HP, Johnson SD, Kuhlmann M, Matthee CA, Nyffeler R, Swartz ER (2010) Biotic diversity in southern African winter-rainfall region. Environ Sustain 2:109–116. https://doi.org/10.1016/j.cosust.2010.02.001
Lubbe A, Jordaan M, Bragg C, Paxton BR, Schumann M, Van der Walt R, Ngobela T (2015) Labeo seeberi. The IUCN Red List of Threatened Species 2015, p e.T11071A76889617. https://doi.org/10.2305/IUCN.UK.2015-2.RLTS.T11071A76889617.en
Luhariya RK, Lal KK, Singh RK, Mohindra V, Punia P, Chauhan UK, Gupta A, Lakra WS (2012) Genetic divergence in wild population of Labeo rohita (Hamilton, 1822) from nine Indian rivers, analyzed through MtDNA cytochrome b region. Mol Biol Rep 39:3659–3665. https://doi.org/10.1007/s11033-011-1140-4
Luikart G, Allendorf FW, Cornuet JM, Sherwin WB (1998) Distortion of allele frequency distributions provides a test for recent population bottlenecks. J Hered 89(3):238–247. https://doi.org/10.1093/jhered/89.3.238
Martinez AS, Willoughby JR, Christie MR (2018) Genetic diversity in fishes is influenced by habitat type and life-history variation. Ecol Evol 8(23):12022–12031. https://doi.org/10.1002/ece3.4661
Modeel S, Joshi BD, Yadav S, Bharti M, Negi RK (2023) Mitochondrial DNA reveals shallow population genetic structure in economically important Cyprinid fish Labeo rohita (Hamilton, 1822) from South and Southeast Asia. Mol Biol Rep 50:4759–4767. https://doi.org/10.1007/s11033-023-08386-5
Mohindra V, Narain L, Punia P, Gopalakrishnan A, Mandal A, Kapoor D, Ponniah AG, Lal KK (2005) Microsatellite DNA markers for population-genetic studies of Labeo dyocheilus (McClelland, 1839). J Appl Ichthyol 21(6):478–482. https://doi.org/10.1111/j.1439-0426.2005.00668.x
Monteiro HJA, Saurel C, Jacobsen MB, Hemmer-Hansen J, Bekkevold D (2022) Genetic parentage reconstruction as a practical tool applied to aquaculture and restoration programs for the European flat. Aquat Living Resour 35:18. https://doi.org/10.1051/alr/2022018
Mulder PFS (1973) Aspects on the ecology of Labeo capensis and Labeo umbratus in the Vaal River. Afr Zool 8(1):15–24. https://doi.org/10.1080/00445096.1973.11447462
Myers N, Mittermeier RA, Mittermeier CG, Da Fonseca GA, Kent J (2000) Biodiversity hotspots for conservation priorities. Nature 403(6772):853–858. https://doi.org/10.1038/35002501
Palstra FP, Ruzzante DE (2008) Genetic estimates of contemporary effective population size: what can they tell us about the importance of genetic stochasticity for wild population persistence? Mol Ecol 17:3428–3447. https://doi.org/10.1111/j.1365-294x.2008.03842.x
Park S (2001) The Excel add-in MS_tools.xla contains Microsoft Excel visual basic macros for use with autosomal microsatellite data. Genetics Department, Trinity College, Dunblin, Ireland
Patel A, Das P, Meher PK, Barat A, Sarangi N (2011) Test of Mendelian segregation and linkage relationships among 69 microsatellite loci in rohu (Labeo rohita). Indian J Anim Sci 81(8):128
Pavlova A, Beheregaray LB, Coleman R, Gilligan D, Harrisson KA, Ingram BA, Kearns J, Lamb AM, Lintermans M, Lyon J, Nguyen TT (2017) Severe consequences of habitat fragmentation on genetic diversity of an endangered Australian freshwater fish: a call for assisted gene flow. Evol Appl 10(6):531–550. https://doi.org/10.1111/eva.12484
Paxton BR, Clark BM, Brown CA (2002) An assessment of the effects of habitat degradation and exotic fish species invasions on the distribution of three endemic cyprinids: Barbus capensis, Barbus serra and Labeo seeberi in the Olifants and Doring Rivers, Western Cape. Report prepared for the Department of Water Affairs and Forestry and Department of Agriculture. DWAF Report No. PB E000-00-1302
Paxton BR, Ramollo P, Schumann M, Jordaan M (2012) Draft biodiversity management plan for species for the Clanwilliam Sandfish (Labeo seeberi). The Freshwater Consulting Group (FCG), Northern Cape Department Nature and Environment Conservation and CapeNature
Peacock MM, Gustin MS, Kirchoff VS, Robinson ML, Hekkala E, Pizzarro-Barraza C, Loux T (2016) Native fishes in the Truckee River: are in-stream structures and patterns of population genetic structure related? Sci Total Environ 563:221–236. https://doi.org/10.1016/j.scitotenv.2016.04.056
Peakall R, Smouse PE (2012) GenAlEx 6.5: genetic analysis in Excel. Population genetic software for teaching and research—an update. Bioinformatics 28(19):2537–2539. https://doi.org/10.1093/bioinformatics/bts460
Piry S, Luikart G, Cornuet JM (1999) BOTTLENECK: a program for detecting recent effective population size reductions from allele data frequencies. J Hered 90(4):502–503. https://doi.org/10.1093/jhered/90.4.502
Potts WM, Booth AJ, Hecht T, Andrew TG (2005) Reproductive biology of a riverine cyprinid, Labeo umbratus (Teleostei: Cyprinidae), in small South African reservoirs. Afr J Aquat Sci 30:147–155. https://doi.org/10.2989/16085910509503849
Prunier JG, Loot G, Veyssiere C, Poulet N, Blanchet S (2023) Novel operational index reveals rapid recovery of genetic connectivity in freshwater fish species after riverine restoration. Conserv Lett 16(2):e12939. https://doi.org/10.1111/conl.12939
Queller DC, Goodnight KF (1989) Estimating relatedness using genetic markers. Evolution 43:258–275. https://doi.org/10.1111/j.1558-5646.1989.tb04226.x
Ramoejane M, Weyl OLF, Swartz ER, Gouws G (2021) Identifying multiple geographically restricted phylogeographic lineages of Moggel (Cyprinidae: Labeo umbratus) in South Africa. Afr J Aquat Sci 46(2):225–235
Ramoejane M (2016) Genetic diversity, evolutionary relationships and conservation of southern African Labeo fishes in relation to water management. PhD Thesis, Rhodes University, South Africa
Reed DH, Frankham R (2003) Correlation between fitness and genetic diversity. Conserv Biol 17:230–237. https://doi.org/10.1046/j.1523-1739.2003.01236.x
Reid GM, Contreras MacBeath T, Csatádi K (2013) Global challenges in freshwater-fish conservation related to public aquariums and the aquarium industry. Int Zoo Yearb 47:6–45. https://doi.org/10.1111/izy.12020
Rhode C, Bester-van der Merwe AE, Roodt-Wilding R (2017) An assessment of spatio-temporal genetic variation in the South African abalone (Haliotis midae), using SNPs: implications for conservation management. Conserv Genet 18:17–31. https://doi.org/10.1007/s10592-016-0879-5
Rosenfeld JS (2014) 50/500 or 100/1000? Reconciling short-and long-term recovery targets and MVPs. Biol Conserv 176:287–288. https://doi.org/10.1016/j.biocon.2014.05.005
Rougemont Q, Dolo V, Oger A, Besnard AL, Huteau D, Coutellec MA, Perrier C, Launey S, Evanno G (2021) Riverscape genetics in brook lamprey: genetic diversity is less influenced by river fragmentation than by gene flow with the anadromous ecotype. Heredity 126(2):235–250. https://doi.org/10.1038/s41437-020-00367-9
Rousset F (2008) GENEPOP’007: a complete re-implementation of the GENEPOP software for Windows and Linux. Mol Ecol Res 8(1):103–106. https://doi.org/10.1111/j.1471-8286.2007.01931.x
Sahoo L, Sahu BP, Das SP, Swain SK, Bej D, Patel A, Jayasankar P, Das P (2014) Limited genetic differentiation in Labeo rohita (Hamilton 1822) populations as revealed by microsatellite markers. Biochem Syst Ecol 57:427–431. https://doi.org/10.1016/j.bse.2014.09.014
Serbezov D, Jorde PE, Bernatchez L, Olsen EM, Vøllestad LA (2012) Short-term genetic changes: evaluating effective population size estimates in a comprehensively described brown trout (Salmo trutta) population. Genetics 191(2):579–592. https://doi.org/10.1534/genetics.111.136580
Shelley JJ, Holland OJ, Swearer SE, Dempster T, Le Feuvre MC, Sherman CD, Miller AD (2022) Landscape context and dispersal ability as determinants of population genetic structure in freshwater fishes. Freshw Biol 67(2):338–352. https://doi.org/10.1111/fwb.13844
Singh RK, Lal KK, Mohindra V, Punia P, Sah RS, Kuar R, Gupta A, Das R, Lakra WS, Ayyappan S (2012) Genetic diversity of Indian Major Carp, Labeo calbasu (Hamilton, 1822) populations inferred from microsatellite loci. Biochem Syst Ecol 44:307–316. https://doi.org/10.1016/j.bse.2012.02.002
Swain S, Das SP, Bej D, Patel A, Jayasankar P, Das PC, Das P (2012) Isolation and characterization of sixteen microsatellite loci for fringe-lipped carp, Labeo fimbriatus. Conserv Genet Res 4:913–915. https://doi.org/10.1007/s12686-012-9672-z
Swain S, Das SP, Bej D, Patel A, Jayasankar P, Chaudhary BK, Jena JK, Routray P, Swain SK, Das PC, Lakra WS (2013) Evaluation of genetic variation in Labeo fimbriatus (Bloch, 1795) populations using heterologous primers. Indian J Fish 60(1):29–35
Swain SK, Sahu BP, Das SP, Sahoo L, Das PC, Das P (2022) Population genetic structure of fringe-lipped carp, Labeo fimbriatus from the peninsular rivers of India. 3 Biotech 12(11):300. https://doi.org/10.1007/s13205-022-03369-y
Swartz ER, Flemming AF, Mouton PLN (2004) Contrasting genetic patterns and population histories in three threatened redfin species (Cyprinidae) from the Olifants River system, western South Africa. J Fish Biol 64:153–1167. https://doi.org/10.1111/j.0022-1112.2004.00375.x
Swartz ER, Skelton PH, Bloomer P (2007) Sea-level changes, river capture and the evolution of populations of the Eastern Cape and fiery redfins (Pseudobarbus afer and Pseudobarbus phlegethon, Cyprinidae) across multiple river systems in South Africa. J Biogeogr 34(12):2086–2099. https://doi.org/10.1111/j.1365-2699.2007.01768.x
Swartz ER, Skelton PH, Bloomer P (2009) Phylogeny and biogeography of the genus Pseudobarbus (Cyprinidae): shedding light on the drainage history of rivers associated with the Cape Floristic Region. Mol Phylogenet Evol 51(1):75–84. https://doi.org/10.1016/j.ympev.2008.10.017
Tweddle D, Bills R, Swartz ER, Coetzer W, Da Costa L, Engelbrecht J, Cambray JA, Marshall B, Impson ND, Skelton PH, Darwall WRT, Smith KG (2009) The status and distribution of freshwater fishes. In: Darwall WRT, Smith KG, Tweddle D, Skelton P (eds) The status and distribution of freshwater biodiversity in South Africa. IUCN, Gland, pp 21–37
Van Der Walt JA, Weyl OL, Woodford DJ, Radloff FG (2016) Spatial extent and consequences of black bass (Micropterus spp.) invasion in a Cape Floristic Region river basin. Aquat Conserv Mar Freshw 26(4):736–748. https://doi.org/10.1002/aqc.2589
Van Oosterhout C, Hutchinson WF, Wills DP, Shipley P (2004) MICRO-CHECKER: software for identifying and correcting genotyping errors in microsatellite data. Mol Ecol Notes 4(3):535–538. https://doi.org/10.1111/j.1471-8286.2004.00684.x
Wan QH, Wu H, Fujihara T, Fang SG (2004) Which genetic marker for which conservation genetics issue? Electrophoresis 25(14):2165–2176. https://doi.org/10.1002/elps.200305922
Wang T, Fujiwara M, Gao X, Liu H (2019) Minimum viable population size and population growth rate of freshwater fishes and their relationships with life history traits. Sci Rep 9(1):1–8. https://doi.org/10.1038/s41598-019-40340-z
Waples RS, Do C (2010) Linkage disequilibrium estimates of contemporary Ne using highly variable genetic markers: a largely untapped resource for applied conservation and evolution. Evol Appl 3:244–262. https://doi.org/10.1111/j.1752-4571.2009.00104.x
Waples R, Larson W, Waples R (2016) Estimating contemporary effective population size in non-model species using linkage disequilibrium across thousands of loci. Heredity 117:233–240. https://doi.org/10.1038/hdy.2016.60
Weyl OLF, Ellender BR, Woodford DJ, Jordaan MS (2013) Fish distributions in the Rondegat River, Cape Floristic Region, South Africa, and the immediate impact of rotenone treatment in an invaded reach. Afr J Aquat Sci 38:201–209. https://doi.org/10.2989/16085914.2012.753401
Weyl OLF, Finlayson B, Impson ND, Woodford DJ, Steikjer J (2014) Threatened endemic fishes in South Africa’s Cape Floristic Region: a new Beginning for the Rondegat River. Fisheries 39(6):270–278. https://doi.org/10.1080/03632415.2014.914924
Acknowledgements
The authors would like to thank the staff from the following conservation agencies: CapeNature, Northern Cape Department of Nature and Environment Conservation, and the Endangered Wildlife Trust for assisting in the acquisition of biological specimens. Ms Therese Forsythe from CapeNature is specifically thanked for assisting with drawing the map of sampling sites (Fig. 1). The DNA Sequencing Unit of the Central Analytical Facility of Stellenbosch University is thanked for rendering analytical services. Lastly, we would like to thank the reviewers for their inputs that helped improve the manuscript.
Funding
Open access funding provided by Stellenbosch University. This project was conducted through the in-kind contributions of the various research partners and collaborators.
Author information
Authors and Affiliations
Contributions
C. Rhode and R. Slabbert conceptualized and designed the study and provided overall project management, coordination, and supervision. S.F. Lesch conducted wet bench work, and preliminary data analysis. K.L. Hull contributed to data analysis and presentation of results. M.S. Jordaan assisted in the coordination of field work and biological sample acquisition. The original draft manuscript was written by C. Rhode and all authors contributed to the reviewing and editing of subsequent versions. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
The original online version of this article was revised due to change in river name as Oorlogskloof River in the introduction of page 2.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Rhode, C., Lesch, S.F., Hull, K.L. et al. Population genetics of the endangered Clanwilliam sandfish Labeo seeberi: considerations for conservation management. Aquat Sci 86, 5 (2024). https://doi.org/10.1007/s00027-023-01019-w
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00027-023-01019-w