Abstract
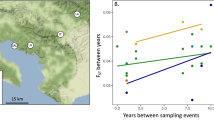
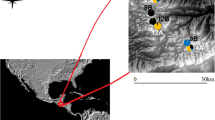
Fragmentation of habitat can decrease resource availability and restrict movement among geographic areas. Persistence in fragmented landscapes depends on the maintenance of connectivity among populations, without which genetic diversity may decrease and lead to population declines. Bees are particularly vulnerable to the negative effects of low genetic diversity so it is important to understand patterns of dispersal for native bees living in fragmented areas. I used population genetic techniques to characterize patterns of genetic diversity and dispersal for the orchid bee Euglossa imperialis within and among forest fragments in southern Costa Rica, in which the furthest two fragments were 226 km from one another. In addition, I compared results of population genetic analyses conducted with all bees sampled, and results from analyses conducted with a reduced dataset containing only one individual per full sibling family from each site. For both datasets genetic diversity was low within forest fragments, with expected heterozygosity averaging 0.28 for the full dataset and 0.29 for the dataset containing only one full sibling per site. I found no evidence that deforested areas restricted dispersal; pairwise estimates of genetic differentiation \(F_{\text{ST}}^{\prime }\) among forest fragments averaged 0.01 for the full dataset, and 0 for the dataset containing only one full sibling per site. Genetic distance among sites within forest fragments was significantly correlated to geographic distance for the full dataset, but there was no significant correlation for the dataset that contained only one individual from each full sibling family. This suggests that family structure can drive results of analyses of genetic structure, although reductions in sample sizes following removal of full siblings may have reduced power to detect genetic structure. Despite no evidence for restricted dispersal, the low genetic diversity found suggests that E. imperialis may be an important candidate for future conservation monitoring.

Similar content being viewed by others
References
Ackerman JD, Montaivo AM (1985) Longevity of Euglossine bees. Biotropica 17:79–81
Ackerman JD, Mesler MR, Lu KL, Montaivo AM (1982) Food-foraging behavior of male Euglossini (Hymenoptera: Apidae): vagabonds or trapliners? Biotropica 14:281–289
Aguiar W, Sofia SH, Melo GAR, Gaglianone MC (2015) Changes in orchid bee communities across forest-agroecosystem boundaries in Brazilian Atlantic forest landscapes. Environ Entomol 44:1465–1471
Augusto SC, Garofalo CA (2010) Task allocation and interactions among females in Euglossa carolina nests (Hymenoptera, Apidae, Euglossini). Apidologie 42:162–173
Becker P, Moure JS, Peralta JA (1991) More about Euglossine bees in Amazonian forest fragments. Biotropica 23:586–591
Beye M, Hasselman M, Fondrk MK, Page RP Jr, Om-Holt SW (2003) The gene csd is the primary signal for sexual development in the honeybee and encodes an SR-type protein. Cell 114:419–429
Boff S, Soro A, Paxton RJ, Alves-dos-Santos I (2013) Island isolation reduces genetic diversity and connectivity but does not significantly elevate diploid male production in a neotropical orchid bee. Con Gen 15:1123–1135
Briggs HM, Perfecto I, Brosi BJ (2013) The role of the agricultural matrix: coffee management and Euglossine bee (Hymenoptera: Apidae: Euglossini) communities in southern Mexico. Environ Entomol 42:1210–1217
Brosi BJ (2009) The effects of forest fragmentation on euglossine bee communities: (Hymenoptera: Apidea: Euglossini). Biol Conserv 142:414–423
Brosi B, Daily GC, Shih TM, Ovideo F, Dura G (2007) The effects of forest fragmentation on bee communities in tropical countryside. J Appl Ecol 45:773–783
Cerântola N, Oi CA, Cervini M, Del Lama MA (2010) Genetic differentiation of urban populations of Euglossa cordata from the state of São Paulo Brazil. Apidologie. doi:10.1051/apido/2010055
Clement R, Horn S (2001) Pre-Columbian land-use history in Costa Rica: a 3000-year record of forest clearance, agriculture and fires from Laguna Zoncho. Holocene 11:419–426
Crozier RH (1976) Counter-intuitive property of effective population size. Nature 262:384
Davis ES, Murray TE, Fitzpatrick U, Brown MJF, Paxton RJ (2010) Landscape effects on extremely fragmented populations of a rare solitary bee, Colletes floralis. Mol Ecol 19:4922–4935
Dressler RL (1982) Biology of the orchid bees (Euglossini). Annu Rev Ecol Syst 13:373–394
Earl DA, vonHoldt BM (2012) STRUCTURE HARVESTER: a website and program for visualizing STRUCTURE output and implementing the Evanno method. Conserv Genet Resour 4:259–361
Eltz T, Roubik DW, Lunau K (2005a) Experience-dependent choices ensure species-specific fragrance accumulation in male orchid bees. Behav Ecol Sociobiol 59:149–156
Eltz T, Sager A, Lunau K (2005b) Juggling with volatiles: exposure of perfumes by displaying male orchid bees. J Comp Physiol A 191:575–581
Ewers RM, Didham RK (2005) Confounding factors in the detection of species responses to habitat fragmentation. Biol Rev 81:117–142
Freiria GA, Ruim JB, Souza RF, Sofia SH (2012) Population structure and genetic diversity of the orchid bee Eufriesea violacea (Hymenoptera, Apidae, Euglossini) from Atlantic forest remnants in southern and southeastern Brazil. Apidologie 4:392–402
Gaggiotti OE, Hanski I (2004) Mechanisms of population extinction. In: Hanski I, Gaggiotti OE (eds) Ecology, genetics, and evolution of metapopulations. Elsevier Academic Press, San Diego, pp 337–366
Giangarelli DC, Freiria GA, Ferreira DG, Aguiar WM, Penha RES, Alves AN, Gaglionone MC, Sofia SH (2015) Orchid bees: a new assessment on the rarity of diploid males in populations of this group of neotropical pollinators. Apidologie 46:606–617
Goudet J, Jombart T (2015). Hierfstat: estimation and tests of hierarchical F-statistics. R package version 0.04-22
Greenleaf SS, Williams NM, Winfree R, Kremen C (2007) Bee foraging ranges and their relationship to body size. Oecologia 153:589–596
Guillot G, Rousset F (2013) Dismanteling the Mantel tests. Methods Ecol Evol 4:336–344
Hale ML, Burg TM, Steeves TE (2012) Sampling for microsatellite-based population genetic studies: 25 to 30 individuals per population Is enough to accurately estimate allele frequencies. PLoS ONE 7:e45170
Hedrick PW, Parker JD (1997) Evolutionary genetics and genetic variation of haplodiploids and x-lined genes. Annu Rev Ecol Syst 28:55–83
Hooke RL, Martín-Duque JF (2012) Land transformations by humans: a review. GSA Today 22:4–10
Hubisz MJ, Falush D, Stephens M, Pritchard JK (2009) Inferring weak population structure with the assistance of sample group information. Mol Ecol Resour 9:1322–1332
Janzen DH (1971) Euglossine bees as long-distance pollinators of tropical plants. Science 171:203–205
Janzen DH (1981) Reduction in euglossine bee species richness on Isla del Caño, a Costa Rican offshore island. Biotropica 13:238–239
Jost L (2008) GST and its relatives do not measure differentiation. Mol Ecol 17:4015–4026
Kremen C, Chaplin-Kramer R (2007) Insects as providers of crop pollination and pest control. In Stewart AJA, New TR, Lewis OT (eds.). Insect Conservation Biology, CABI, Cambridge, pp. 349–382
Kremen C, Williams NM, Thorp RW (2002) Crop pollination from native bees at risk from agricultural intensification. Proc Natl Acad Sci USA 99:16812–16816
Legendre P, Fortin M (2010) Comparison of the Mantel test and alternative approaches for detecting complex multivariate relationships in the spatial analysis of genetic data. Mol Ecol Resour 10:831–844
López‐Uribe MM, Zamudio KR, Cardoso CF, Danforth BN (2014) Climate, physiological tolerance and sex‐biased dispersal shape genetic structure of Neotropical orchid bees. Mol Ecol 23:1874–1890
Meirmans PG (2015) Seven common mistakes in population genetics and how to avoid them. Mol Ecol 13:3223–3231
Meirmans PG, Hedrick PW (2011) Assessing population structure: F ST and related measures. Mol Ecol Resour 11:5–18
Meirmans PG, Van Tienderen PH (2004) GENOTYPE and GENODIVE: two programs for the analysis of genetic diversity of asexual organisms. Mol Ecol Notes 4:792–794
Milet-Pinheiro Schlindwein C (2005) Do Euglossine males (Apidae, Euglossini) leave tropical rainforest to collect fragrances in sugarcane monocultures? Rev Bras Zool 22:853–858
Narum SR (2006) Beyond Bonferroni: less conservative analyses for landscape genetics. Conserv Genet 7:783–787
Nei M (1987) Molecular evolutionary genetics. Columbia University Press, New York
Nemésio A, Silveira FA (2010) Forest fragments with larger core areas better sustain diverse orchid bee faunas (Hymenoptera: Apidae: Euglossina). Neotrop Entomol 39:555–561
Oi CA, López-Uribe M, Cervini M, Del Lama MA (2013) Non-lethal method of DNA sampling in euglossine bees supported by mark-recapture experiments and microsatellite genotyping. J Insect Conserv 17:1071–1079
Ollerton J, Winfree R, Tarrant S (2011) How many flowering plants are pollinated by animals? Oikos 120:321–326
Paetkau D, Strobeck C (1995) The molecular basis and evolutionary history of a microsatellite null allele in bears. Mol Ecol 4:519–520
Paxton RJ, Zobel MU, Steiner J, Zillikens A (2009) Microsatellite loci for Euglossa annectans (Hymenoptera: Apidae) and their variability in other orchid bees. Mol Ecol Resour 9:1221–1223
Peakall R, Smouse PE (2006) Genalex 6: genetic analysis in excel. Population genetic software for teaching and research. Mol Ecol Notes 6:288–295
Penha RES, Gaglianone MC, Almeida FS, Boff SV, Sofia SH (2015) Mitochondrial DNA of Euglossa Iopoecila (Apidae, Euglossini) reveals two distinct lineages for this orchid bee species endemic to the Atlantic Forest. Apidologie 46:346–358
Pokorny T, Hannibal M, Quezada-Euan JJG, Hedenström E, Sjöberg N, Bång J, Eltz T (2013) Acquisition of species-specific perfume blends: influence of habitat-dependent compound availability on odour choices of male orchid bees (Euglossa spp.). Oecologia 172:417–425
Pokorny T, Loose D, Dyker G, Quezada-Euán Eltz T (2015) Dispersal ability of male orchid bees and direct evidence for long-range flights. Apidologie 46:224–237
Powell AH, Powell GVN (1987) Population dynamics of male euglossine bees in Amazonian forest fragments. Biotropica 19:176–179
Pritchard JM, Stephens M, Donnelly P (2000) Inference of population structure using multilocus genotype data. Genetics 155:945–959
Pritchard JK, Wen X, Falush D (2007) Documentation for structure software: version 2.2. University of Chicago, Chicago, pp 1–36
Prugh LR, Hodges KE, Sinclair ARE, Brashares JS (2008) Effects of habitat area and isolation on fragmented animal populations. Proc Natl Acad Sci 105:20770–20775
Ricketts TH, Regetz J, Steffan-Dewenter I, Cunningham SA, Kremen C, Bogdansk A, Gemmill-Herren B, Greenleaf SS, Klein AM, Mayfield MM, Morandin LA, Ochieng A, Potts SG, Viana BF (2008) Landscape effects on crop pollination services: are there general patterns? Ecol Lett 11:499–515
Rocha Filho LC, Cerântola NCM, Garófalo CA, Imperatriz-Fonseca VL, Del Lama MA (2013) Genetic differentiation of the Euglossini (Hymenoptera, Apidae) populations on a mainland coastal plain and an island in southeastern Brazil. Genetica 141:65–74
Rodriguez-Ramilo ST, Wang J (2012) The effect of close relatives on unsupervised Bayesian clustering algorithms in population genetic structure analysis. Mol Ecol Resour 12:873–874
Rosa JF, Ramalho M, Arias MC (2016) Functional connectivity and genetic diversity of Eulaema atleticana (Apidae, Euglossina) in the Brazilian atlantic forest corridor: assessment of gene flow. Biotropica. doi:10.1111/btp.12321
Roubik DW, Hanson PH (2004) Orchid bees of tropical America, 1st edn. INBio, Santo Domingo
Roubik DW, Weight LA, Bonilla MA (1996) Population genetics, diploid males, and limits to social evolution of Euglossine bees. Evolution 50:931–935
Rousset F (1997) Genetic differentiation and estimation of gene flow from F-statistics under isolation by distance. Genetics 145:1219–1228
Rousset F (2008) GENEPOP’007: a complete re-implementation of the GENEPOP software for windows and linux. Mol Ecol Res 8:103–106
Ryman N, Palm S, Andre C, Carvalho GR, Dahlgren TG, Jorde PE, Laikre L, Larsson LC, Palmé A, Ruzzante DE (2006) Power for detecting genetic divergence: differences between statistical methods and marker loci. Mol Ecol 15:2031–2045
Sambrook J, Fritsch EF, Maniatis T (1989) Molecular cloning: a laboratory manual, 2nd edn. Cold Spring Harbor Press, Cold Spring Harbor
Schwartz MK, McKelvey KS (2009) Why sampling scheme matters: the effect of sampling scheme on landscape genetic results. Conserv Genet 10:441–452
Silva DP, Marco JR (2014) No evidence of habitat loss affecting the orchid bees Eulaema nigrita Lepeletier and Eufriesea auriceps Friese (Apidae: Euglossini) in the Brazilian Cerrado Savanna. Neotrop Entomol 43:509–518
Silveira FA, Melo GAR, Almeida EAB (2002) Abelhas Brasileiras: sistemática e identificação, Belo Horizonte
Silviera GC, Freitas RF, Tosta THA, Rabelo LS, Gaglianone MC, Augusto SC (2015) The orchid bee fauna in the Brazilian savanna: do forest formations contribute to higher species diversity? Apidologie 46:197–208
Sofia SH, Paula FM, dos Santos AM, Almeida FS, Sodré LMK (2005) Genetic structure analysis of Eufriesea violacea (Hymenoptera, Apidae) populations from southern Brazilian Atlantic rainforest remnants. Genet Mol Biol 28:479–484
Souza RO, Cervini M, Del Lama MA, Paxton RJ (2007) Microsatellite loci for euglossine bees (Hymenoptera: Apidae). Mol Ecol Notes 7:1352–1356
Souza RO, Del Lama MA, Cervini M, Mortari N, Eltz T, Zimmermann Y, Bach C, Brosi BJ, Suni S, Quezada-Euán JJG, Paxton RJ (2010) Conservation genetics of neotropical pollinators revisited: microsatellite analysis suggests that diploid males are rare in orchid bees. Evolution 64:3318–3326
Stern DL (1991) Male territoriality and alternative male behaviors in the euglossine bee, Eulaema meriana (Hymenoptera: Apidae). J Kans Entomol Soc 64:421–437
Suni SS, Brosi BJ (2012) Population genetics of orchid bees in a fragmented tropical landscape. Conserv Genet 13:323–332
Suni SS, Brosi BJ, Bronstein J (2014) Spatio-temporal genetic structure suggests high dispersal over a fragmented landscape. Biotropica 46:202–209
Taylor A (2003) Assessing the consequences of inbreeding for population fitness: past challenges and future prospects. Cambridge University Press, Cambridge
Tonhasca A, Blackmer JL, Albuquerque GS (2002a) Abundance and diversity of Euglossine bees in the fragmented landscape of the Brazilian Atlantic forest. Biotropica 34:416–422
Tonhasca A, Blackmer JL, Albuquerque GS (2002b) Within-habitat heterogeneity of euglossine bee populations: a re-evaluation of the evidence. J Trop Ecol 18:929–933
van Wilgenburg E, Driessen G, Beukeboom LW (2006) Single locus complementary sex determination in Hymenoptera: an “unintelligent” design? Front Zool 3:1–15
Wang JL (2004) Sibship reconstruction from genetic data with typing errors. Genetics 166:1963–1979
Wang JL (2013) Examining the full effects of landscape heterogeneity on spatial genetic variation: a multiple matrix regression approach for quantifying geographic and ecological isolation. Evolution 67:3401–3411
Whiteley AR, Coombs JA, Hudy M, Robinson Z, Colton AR, Nislow KH, Letcher BH (2013) Fragmentation and patch size shape genetic structure of brook trout populations. Can J Fish Aquat Sci 70:678–688
Whiteley AR, McGarigal K, Schwartz MK (2014) Pronounced differences in genetic structure despite overall ecological similarity for two Ambystoma salamanders in the same landscape. Conserv Genet 15:573–591
Wikelski M, Moxley J, Eaton-Mordas A, López-Uribe MM, Holland R, Moskowitz D, Roubik DW, Kays R (2010) Large-range movements of neotropical orchid bees observed via radio telemetry. PLoS ONE 5:1–6
Winfree R, Aguilar R, Casquez DP, Lebuhn G, Aizen MA (2009) A meta-analysis of bees’ responses to anthropogenic disturbance. Ecology 90:2068–2076
Wright S (1951) The genetical structure of natural populations. Ann Eugen 15:323–354
Zayed A, Packer L (2005) Complementary sex determination substantially increases extinction proneness of haplodiploid populations. Proc Natl Acad Sci USA 102:10742–10746
Zimmermann Y, Schorkopf DLP, Mortiz RFA, Pemberton RW, Quezada-Euán JJG, Eltz T (2011) Population genetic structure of orchid bees (Euglossini) in anthropogenically altered landscapes. Conserv Genet 12:1183–1194
Acknowledgements
I thank the staff at the Las Cruces Biological Station near San Vito, Costa Rica, H. Woodard at Saladero Lodge in Costa Rica, M. Kaplan and the staff of University of Arizona’s genomics core facility for help with laboratory work, B. Brosi and T. Brookhart for fieldwork assistance, and R. Hopkins for comments on the manuscript. Fieldwork and genotyping were supported by the Center for Insect Science at the University of Arizona (NIH grant 5K12 GM000708).
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Suni, S.S. Dispersal of the orchid bee Euglossa imperialis over degraded habitat and intact forest. Conserv Genet 18, 621–630 (2017). https://doi.org/10.1007/s10592-016-0902-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10592-016-0902-x