Abstract
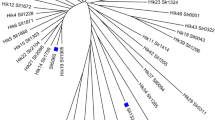
Phosphoglycerate mutase (PGM) is a widely distributed glycolytic enzyme. Two known distinct classes of PGM enzymes were identified, a cofactor-dependent one (dPGM) and a cofactor-independent one (iPGM). A complementary DNA (cDNA) encoding a PGM was cloned from a Clonorchis sinensis cDNA library by large-scale sequencing. This new cDNA contains 955 bp with a putative open reading frame of 256 amino acids, which has a high homology with dPGMs from a number of species. The putative peptide was produced in E. coli and was purified to electrophoretic homogeneity. Enzymatic assays showed that the product of this gene could catalyze the conversion of 3-phosphoglycerate to 2-phosphoglycerate when the cofactor was present and the enzyme activities could be inhibited by vanadate.



Similar content being viewed by others
References
Bond CS, White MF, Hunter WN (2002) Mechanistic implications for Escherichia coli cofactor-dependent phosphoglycerate mutase based on the high-resolution crystal structure of a vanadate complex. J Mol Biol 316:1071–1081
Bradford MM (1976) A rapid and sensitive method for the quantitation of microgram quantities of protein utilizing the principle of protein-dye binding. Anal Biochem 72:248–254
Campbell JW, Watson HC, Hodgson GI (1974) Structure of yeast phosphoglycerate mutase. Nature 250:301–303
Carreras J, Bartrons R, Grisolia S (1980) Vanadate inhibits 2,3-bisphosphoglycerate dependent phosphoglycerate mutases but does not affect the 2,3-bisphosphoglycerate independent phosphoglycerate mutases. Biochem Biophys Res Commun 96:1267–1273
Fothergill-Gilmore LA, Watson HC (1989) The phosphoglycerate mutases. Adv Enzymol Relat Areas Mol Biol 62:227–313
Fraser HI, Kvaratskhelia M, White MF (1999) The two analogous phosphoglycerate mutases of Escherichia coli. FEBS Lett 455:344–348
Galperin MY, Jedrzejas MJ (2001) Conserved core structure and active site residues in alkaline phosphatase superfamily enzymes. Proteins 45:318–324
Galperin MY, Bairoch A, Koonin EV (1998) A superfamily of metalloenzymes unifies phosphopentomutase and cofactor-independent phosphoglycerate mutase with alkaline phosphatases and sulfatases. Protein Sci 7:1829–1835
Grana X, Perez de la Ossa P, Broceno C, Stocker M, Garriga J, Puigdomenech P, Climent F (1995) 2,3-Bisphosphoglycerate-independent phosphoglycerate mutase is conserved among different phylogenic kingdoms. Comp Biochem Physiol B Biochem Mol Biol 112:287–293
Jedrzejas MJ (2000) Structure, function, and evolution of phosphoglycerate mutases: comparison with fructose-2,6-bisphosphatase, acid phosphatase, and alkaline phosphatase. Prog Biophys Mol Biol 73:263–287
Jedrzejas MJ, Chander M, Setlow P, Krishnasamy G (2000a) Structure and mechanism of action of a novel phosphoglycerate mutase from Bacillus stearothermophilu. EMBO J 19:1419–1431
Jedrzejas MJ, Chander M, Setlow P, Krishnasamy G (2000b) Mechanism of catalysis of the cofactor-independent phosphoglycerate mutase from Bacillus stearothermophilus. Crystal structure of the complex with 2-phosphoglycerate. J Biol Chem 275:23146–23153
Lee YH, Ogata C, Pflugrath JW, Levitt DG, Sarma R, Banaszak LJ, Pilkis SJ (1996) Crystal structure of the rat liver fructose-2,6-bisphosphatase based on selenomethionine multiwavelength anomalous dispersion phases. Biochemistry 35:6010–6019
Leyva-Vazquez MA, Setlow P (1994) Cloning and nucleotide sequences of the genes encoding triose phosphate isomerase, phosphoglycerate mutase, and enolase from Bacillus subtilis. J Bacteriol 176:3903–3910
Lindqvist Y, Schneider G, Vihko P (1993) Three-dimensional structure of rat acid phosphatase in complex with L(+)-tartrate. J Biol Chem 268:20744–20746
Morris VL, Jackson DP, Grattan M, Ainsworth T, Cuppels DA (1995) Isolation and sequence analysis of the Pseudomonas syringae pv. tomato gene encoding a 2,3-diphosphoglycerate-independent phosphoglyceromutase. J Bacteriol 177:1727–1733
Rigden DJ, Alexeev D, Phillips SE, Fothergill-Gilmore LA (1998) The 2.3 Å X-ray crystal structure of S. cerevisiae phosphoglycerate mutase. J Mol Biol 276:449–459
Rigden DJ, Mello LV, Setlow P, Jedrzejas MJ (2002) Structure and mechanism of action of a cofactor-dependent phosphoglycerate mutase homolog from Bacillus stearothermophilus with broad specificity phosphatase activity. J Mol Biol 315:1129–1143
Song L, Chen S, Yu X, Wu Z, Xu J, Yang G, Zheng N, Hu X, Guo L, Dai J, Xu J, Ji C, Gu S, Ying K (2004) Molecular cloning and characterization of cDNA encoding a ubiquitin-conjugating enzyme from Clonorchis sinensis. Parasitol Res 94:227–232
Sowadski JM, Handschumacher MD, Murthy HM, Foster BA, Wyckoff HW (1985) Refined structure of alkaline phosphatase from Escherichia coli at 2.8 Å resolution. J Mol Biol 186:417–433
Yang G, Jing C, Zhu P, Hu X, Xu J, Wu Z, Yu X (2006) Molecular cloning and characterization of a novel lactate dehydrogenase gene from Clonorchis sinensis. Parasitol Res 99:55–64
Zhang Y, Foster JM, Kumar S, Fougere M, Carlow CK (2004) Cofactor-independent phosphoglycerate mutase has an essential role in Caenorhabditis elegans and is conserved in parasitic nematodes. J Biol Chem 279:37185–37190
Zheng N, Xu J, Wu Z, Chen J, Hu X, Song L, Yang G, Ji C, Chen S, Gu S, Ying K, Yu X (2005) Clonorchis sinensis: molecular cloning and functional expression of novel cytosolic malate dehydrogenase. Exp Parasitol 109:220–227
Acknowledgment
This research was supported by grants from the Natural Science Foundation of Guangdong Province, China (No.2002B31005) and the Key Program of Science and Technology Department of Guangdong Province, China (No.200223-E4022). The experiments comply with the current laws of the country in which the experiments were performed.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Song, L., Xu, Z. & Yu, X. Molecular cloning and characterization of a phosphoglycerate mutase gene from Clonorchis sinensis . Parasitol Res 101, 709–714 (2007). https://doi.org/10.1007/s00436-007-0540-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00436-007-0540-9