Abstract
O6-Methylguanine-DNA methyltransferase (MGMT) is a DNA repair protein that removes alkyl DNA adducts such as those induced by alkylating agents. Loss of MGMT expression through transcriptional silencing by hypermethylation of its CpG island (CGI) is found in diverse human cancers including glioblastomas. Glioblastomas that have MGMT methylation respond to temozolomide, an alkylating agent, resulting in improved survival. Consequently, assessment of MGMT methylation has become a therapy response and prognostic indicator. However, it is not clear whether the region of the MGMT CGI commonly analysed is the critical region involved in transcriptional control. We measured methylation levels at each CpG site for the entire MGMT CGI using bisulfite modification and pyrosequencing, and compared them with MGMT mRNA expression in glioblastoma cell lines, xenografts and normal brain tissues (41 samples). Two critical regions were identified (DMR1 and DMR2). DMR2 encompasses the commonly analysed region and was always methylated when DMR1 was methylated. A luciferase reporter assay showed that substitutions of several specific CpG sites within DMR2 significantly attenuated the promoter activity of the MGMT CGI. Our results indicate that several CpG sites within DMR2 play a critical role in the transcriptional control of MGMT, making DMR2 the optimal target for methylation testing. However, given the highly variable patterns of MGMT methylation associated with transcriptional silencing observed in this region among the tumours in this study, methylation levels need to be measured at a number of individual CpGs within DMR2 to confidently predict transcriptional silencing and thus sensitivity to alkylating agents.



Similar content being viewed by others
Abbreviations
- CGI:
-
CpG island
- DMR:
-
Differentially methylated region
- FDR:
-
Benjamini and Hochberg false discover rate
- MGMT:
-
O6-Methylguanine-DNA methyltransferase
- MSP:
-
Methylation-specific PCR
- PCR:
-
Polymerase chain reaction
- qPCR:
-
Real-time quantitative PCR
- TF:
-
Transcription factor
- TMZ:
-
Temozolomide
- TSS:
-
Transcription start site
References
Bhakat KK, Mitra S (2003) CpG methylation-dependent repression of the human O6-methylguanine-DNA methyltransferase gene linked to chromatin structure alteration. Carcinogenesis 24:1337–1345
Brada M, Stenning S, Gabe R et al (2010) Temozolomide versus procarbazine, lomustine, and vincristine in recurrent high-grade glioma. J Clin Oncol 28:4601–4608
Chodavarapu RK, Feng S, Bernatavichute YV et al (2010) Relationship between nucleosome positioning and DNA methylation. Nature 466:388–392
Costello JF, Futscher BW, Kroes RA, Pieper RO (1994) Methylation-related chromatin structure is associated with exclusion of transcription factors from and suppressed expression of the O-6-methylguanine DNA methyltransferase gene in human glioma cell lines. Mol Cell Biol 14:6515–6521
de la Iglesia N, Puram SV, Bonni A (2009) STAT3 regulation of glioblastoma pathogenesis. Curr Mol Med 9:580–590
Dunn J, Baborie A, Alam F et al (2009) Extent of MGMT promoter methylation correlates with outcome in glioblastomas given temozolomide and radiotherapy. Br J Cancer 101:124–131
Esteller M, Hamilton SR, Burger PC, Baylin SB, Herman JG (1999) Inactivation of the DNA repair gene O6-methylguanine-DNA methyltransferase by promoter hypermethylation is a common event in primary human neoplasia. Cancer Res 59:793–797
Esteller M, Herman JG (2004) Generating mutations but providing chemosensitivity: the role of O6-methylguanine DNA methyltransferase in human cancer. Oncogene 23:1–8
Everhard S, Tost J, El Abdalaoui H et al (2009) Identification of regions correlating MGMT promoter methylation and gene expression in glioblastomas. Neuro Oncol 11:348–356
Fuxe J, Akusjarvi G, Goike HM, Roos G, Collins VP, Pettersson RF (2000) Adenovirus-mediated overexpression of p15INK4B inhibits human glioma cell growth, induces replicative senescence, and inhibits telomerase activity similarly to p16INK4A. Cell Growth Differ 11:373–384
Gerson SL (2004) MGMT: its role in cancer aetiology and cancer therapeutics. Nat Rev Cancer 4:296–307
Goike HM, Asplund AC, Pettersson EH, Liu L, Ichimura K, Collins VP (2000) Cryopreservation of viable human glioblastoma xenografts. Neuropathol Appl Neurobiol 26:172–176
Harris LC, Potter PM, Tano K, Shiota S, Mitra S, Brent TP (1991) Characterization of the promoter region of the human O6-methylguanine-DNA methyltransferase gene. Nucleic Acids Res 19:6163–6167
Harris LC, Remack JS, Brent TP (1994) Identification of a 59 bp enhancer located at the first exon/intron boundary of the human O6-methylguanine DNA methyltransferase gene. Nucleic Acids Res 22:4614–4619
Harris LC, Remack JS, Brent TP (1994) In vitro methylation of the human O6-methylguanine-DNA methyltransferase promoter reduces transcription. Biochim Biophys Acta 1217:141–146
Hegi ME, Liu L, Herman JG et al (2008) Correlation of O6-methylguanine methyltransferase (MGMT) promoter methylation with clinical outcomes in glioblastoma and clinical strategies to modulate MGMT activity. J Clin Oncol 26:4189–4199
Herman JG, Graff JR, Myohanen S, Nelkin BD, Baylin SB (1996) Methylation-specific PCR: a novel PCR assay for methylation status of CpG islands. Proc Natl Acad Sci USA 93:9821–9826
Ichimura K, Bolin MB, Goike HM, Schmidt EE, Moshref A, Collins VP (2000) Deregulation of the p14ARF/MDM2/p53 pathway is a prerequisite for human astrocytic gliomas with G1-S transition control gene abnormalities. Cancer Res 60:417–424
Kaina B, Christmann M, Naumann S, Roos WP (2007) MGMT: key node in the battle against genotoxicity, carcinogenicity and apoptosis induced by alkylating agents. DNA Repair (Amst) 6:1079–1099
Kaplan N, Moore IK, Fondufe-Mittendorf Y et al (2009) The DNA-encoded nucleosome organization of a eukaryotic genome. Nature 458:362–366
Karayan-Tapon L, Quillien V, Guilhot J et al (2010) Prognostic value of O6-methylguanine-DNA methyltransferase status in glioblastoma patients, assessed by five different methods. J Neurooncol 97:311–322
Liu L, Backlund LM, Nilsson BR et al (2005) Clinical significance of EGFR amplification and the aberrant EGFRvIII transcript in conventionally treated astrocytic gliomas. J Mol Med 83:917–926
Mikeska T, Bock C, El-Maarri O et al (2007) Optimization of quantitative MGMT promoter methylation analysis using pyrosequencing and combined bisulfite restriction analysis. J Mol Diagn 9:368–381
Nakagawachi T, Soejima H, Urano T et al (2003) Silencing effect of CpG island hypermethylation and histone modifications on O6-methylguanine-DNA methyltransferase (MGMT) gene expression in human cancer. Oncogene 22:8835–8844
Patel SA, Graunke DM, Pieper RO (1997) Aberrant silencing of the CpG island-containing human O6-methylguanine DNA methyltransferase gene is associated with the loss of nucleosome-like positioning. Mol Cell Biol 17:5813–5822
Qian XC, Brent TP (1997) Methylation hot spots in the 5′ flanking region denote silencing of the O6-methylguanine-DNA methyltransferase gene. Cancer Res 57:3672–3677
Riemenschneider MJ, Jeuken JW, Wesseling P, Reifenberger G (2010) Molecular diagnostics of gliomas: state of the art. Acta Neuropathol 120:567–584
Sasai K, Nodagashira M, Nishihara H et al (2008) Careful exclusion of non-neoplastic brain components is required for an appropriate evaluation of O6-methylguanine-DNA methyltransferase status in glioma: relationship between immunohistochemistry and methylation analysis. Am J Surg Pathol 32:1220–1227
Segal E, Widom J (2009) What controls nucleosome positions? Trends Genet 25:335–343
Stupp R, Hegi ME, Mason WP et al (2009) Effects of radiotherapy with concomitant and adjuvant temozolomide versus radiotherapy alone on survival in glioblastoma in a randomised phase III study: 5-year analysis of the EORTC-NCIC trial. Lancet Oncol 10:459–466
Tabatabai G, Stupp R, van den Bent MJ et al (2010) Molecular diagnostics of gliomas: the clinical perspective. Acta Neuropathol 120:585–592
Tost J, El abdalaoui H, Gut IG (2006) Serial pyrosequencing for quantitative DNA methylation analysis. Biotechniques 40:721–722, 724, 726
Watts GS, Pieper RO, Costello JF, Peng YM, Dalton WS, Futscher BW (1997) Methylation of discrete regions of the O6-methylguanine DNA methyltransferase (MGMT) CpG island is associated with heterochromatinization of the MGMT transcription start site and silencing of the gene. Mol Cell Biol 17:5612–5619
Weller M, Felsberg J, Hartmann C et al (2009) Molecular predictors of progression-free and overall survival in patients with newly diagnosed glioblastoma: a prospective translational study of the German Glioma Network. J Clin Oncol 27:5743–5750
Zeng N, Liu L, McCabe MG, Jones DT, Ichimura K, Collins VP (2009) Real-time quantitative polymerase chain reaction (qPCR) analysis with fluorescence resonance energy transfer (FRET) probes reveals differential expression of the four ERBB4 juxtamembrane region variants between medulloblastoma and pilocytic astrocytoma. Neuropathol Appl Neurobiol 35:353–366
Zhao W, Soejima H, Higashimoto K et al (2005) The essential role of histone H3 Lys9 di-methylation and MeCP2 binding in MGMT silencing with poor DNA methylation of the promoter CpG island. J Biochem 137:431–440
Acknowledgments
This work was supported by funds from Samantha Dickson Brain Tumour Trust and Cambridge Fund for the Prevention of Disease (CAMPOD).
Conflict of interest
The authors declare that they have no conflict of interest.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
401_2011_803_MOESM1_ESM.ppt
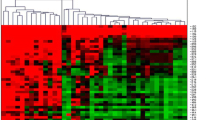
Supplementary Fig. 1. Two-way clustering and heat map of MGMT CGI methylation as determined by pyrosequencing (red, high methylation; green, low methylation). CpG sites clustered together are indicated on the right of the heat map. (PPT 231 kb)
401_2011_803_MOESM2_ESM.ppt
Supplementary Fig. 2. Superimposed plots of the multiple test-corrected p-values using different thresholds for expression to group samples (0.1%, blue; 1%, black; 30%, red) at each CpG site against their distance from TSS (=0, arrow) to indicate that the patterns of p-values are similar. Grid lines represent p=0.05, 0.01 and 0.001. The CpG numbers are indicated at selected sites. The positions of DMRs and MSP primers are shown with grey bars. (PPT 204 kb)
401_2011_803_MOESM3_ESM.ppt
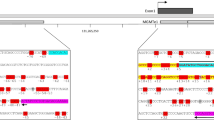
Supplementary Fig. 3. Nucleosome occupancy prediction within MGMT CGI. The posterior probability to start a nucleosome (P start, orange line, right axis) and the nucleosome occupancy prediction (P occupied, black line, left axis) within the MGMT CpG island (see Materials and Methods for the source) are plotted. A High P start value indicates the position nucleosome binding is likely to start. A High P occupied value indicates the position nucleosome is likely to be bound. Positions of CpGs are indicated by grey dots (the dots for CpG83, 86 and 89 are larger) and the numbers of selected CpGs are shown above the dots. Binding sites for various transcription factors predicted by TFSEARCH (http://molsun1.cbrc.aist.go.jp/research/db/TFSEARCH.html) are represented by green lines. Among them, the putative STAT3 binding site is indicated. AP2 and SP1 binding sites predicted by Harris et al. [13] and Costello et al. [4] are shown respectively as red and yellow lines. Frequently methylated regions such as CpG1-20, DMR1 and DMR2 are indicated by dark blue lines (see also Fig. 2) whereas pale blue lines correspond to the minimal promoter and enhancer. TSS, transcription start site. (PPT 147 kb)
Rights and permissions
About this article
Cite this article
Malley, D.S., Hamoudi, R.A., Kocialkowski, S. et al. A distinct region of the MGMT CpG island critical for transcriptional regulation is preferentially methylated in glioblastoma cells and xenografts. Acta Neuropathol 121, 651–661 (2011). https://doi.org/10.1007/s00401-011-0803-5
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00401-011-0803-5