Abstract
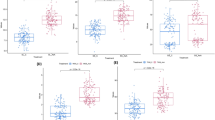
In cereals, a common salinity tolerance mechanism is to limit accumulation of Na+ in the shoot. In a cross between the barley variety Barque-73 (Hordeum vulgare ssp. vulgare) and the accession CPI-71284 of wild barley (H. vulgare ssp. spontaneum), the HvNax3 locus on chromosome 7H was found to determine a ~10–25 % difference in leaf Na+ accumulation in seedlings grown in saline hydroponics, with the beneficial exclusion trait originating from the wild parent. The Na+ exclusion allele was also associated with a 13–21 % increase in shoot fresh weight. The HvNax3 locus was delimited to a 0.4 cM genetic interval, where it cosegregated with the HVP10 gene for vacuolar H+-pyrophosphatase (V-PPase). Sequencing revealed that the mapping parents encoded identical HVP10 proteins, but salinity-induced mRNA expression of HVP10 was higher in CPI-71284 than in Barque-73, in both roots and shoots. By contrast, the expression of several other genes predicted by comparative mapping to be located in the HvNax3 interval was similar in the two parent lines. Previous work demonstrated roles for V-PPase in ion transport and salinity tolerance. We therefore considered transcription levels of HVP10 to be a possible basis for variation in shoot Na+ accumulation and biomass production controlled by the HvNax3 locus under saline conditions. Potential mechanisms linking HVP10 expression patterns to the observed phenotypes are discussed.


Similar content being viewed by others
Abbreviations
- CAPS:
-
Cleaved amplified polymorphic sequence
- HVP:
-
Hordeum vacuolar H+-pyrophosphatase
- MFS:
-
Major facilitator superfamily
- q-RT-PCR:
-
Quantitate reverse transcriptase polymerase chain reaction
- ORF:
-
Open reading frame
- SAM:
-
Sterile alfa motif
- V-PPase:
-
Vacuolar H+-pyrophosphatase
References
Abe K, Ruan ZS, Maloney PC (1996) Cloning, sequencing, and expression in Escherichia coli of OxlT, the oxalate: formate exchange protein of Oxalobacter formigenes. J Biol Chem 271:6789–6793
Ariyadasa R, Stein N (2012) Advances in BAC-based physical mapping and map integration strategies in plants. J Biomed Biotechnol. doi:10.1155/2012/184854
Bao AK, Wang SM, Wu GQ, Xi JJ, Zhang JL, Wang CM (2009) Overexpression of the Arabidopsis H+-PPase enhanced resistance to salt and drought stress in transgenic alfalfa (Medicago sativa L.). Plant Sci 176:232–240
Bayat F, Shiran B, Belyaev DV, Yur’eva NO, Sobol’kova GI, Alizadehe H, Khodambashi M, Babakov AV (2010) Potato plants bearing a vacuolar Na+/H+ antiporter HvNHX2 from barley are characterized by improved salt tolerance. Russ J Plant Physiol 57:696–706
Blumwald E, Gelli A (1997) Secondary inorganic ion transport at the tonoplast. Adv Bot Res 25:401–407
Burton RA, Jobling SA, Harvey AJ, Shirley HJ, Mather DE, Bacic A, Fincher GB (2008) The genetics and transcriptional profiles of the cellulose synthase-like HvCslF gene family in barley. Plant Physiol 146:1821–1833
Byrt CS, Platten JD, Spielmeyer W, James RA, Lagudah ES, Dennis ES, Tester M, Munns R (2007) HKT1;5-like cation transporters linked to Na+ exclusion loci in wheat, Nax2 and Kna1. Plant Physiol 143:1918–1928
Chen Z, Zhou M, Newman IA, Mendham NJ, Zhang G, Shabala S (2007) Potassium and sodium relations in salinised barley tissues as a basis of differential salt tolerance. Funct Plant Biol 34:150–162
Chhipa BR, Lal P (1995) Na/K ratios as the basis of salt tolerance in wheat. Aust J Agric Res 46:533–539
Chomczynski P, Sacchi N (1987) Single-step method of RNA isolation by acid guanidinium thiocyanate-phenol-chloroform extraction. Anal Biochem 162:156–159
Colmer TD, Flower TJ, Munns R (2006) Use of wild relatives to improve salt tolerance in wheat. J Exp Bot 57:1059–1078
Deng W, Nickle DC, Learn GH, Maust B, Mullins JI (2007) ViroBLAST: a stand-alone BLAST web server for flexible queries of multiple databases and user’s datasets. Bioinformatics 23:2334–2336
Ferjani A, Segami S, Horiguchi G, Muto Y, Maeshima M, Tsukaya H (2011) Keep an eye on PPi: the vacuolar-type H+-pyrophosphatase regulates postgerminative development in Arabidopsis. Plant Cell 23:2895–2908
Flowers TJ, Yeo AR (1995) Breeding for salinity resistance in crop plants: where next? Aust J Plant Physiol 22:875–884
Fukuda A, Chiba K, Maeda M, Nakamura A, Maeshima M, Tanaka Y (2004) Effect of salt and osmotic stresses on the expression of genes for the vacuolar H+- pyrophosphatase, H+-ATPase subunit A, and Na+/H+ antiporter from barley. J Exp Bot 55:585–594
Gao F, Gao Q, Duan XG, Yue GD, Yang AF, Zhang JR (2006) Cloning of an H+-PPase gene from Thellungiella halophile and its heterologous expression to improve tobacco salt tolerance. J Exp Bot 57:3259–3270
Garthwaite AJ, von Bothmer R, Colmer TD (2005) Salt tolerance in wild Hordeum species is associated with restricted entry of Na+ and Cl− into the shoots. J Exp Bot 56:2365–2378
Gaxiola RA, Li J, Undurraga S, Dang LM, Allen GJ, Alper SL, Fink GR (2001) Drought- and salt-tolerant plants result from overexpression of the AVP1 H+-pump. Proc Natl Acad Sci USA 98:11444–11449
Gaxiola RA, Fink GR, Hirschi KD (2002) Genetic manipulation of vacuolar proton pumps and transporters. Plant Physiol 129:967–973
Guo S, Yin H, Zhang X, Zhao F, Li P, Chen S, Zhao Y, Zhang H (2006) Molecular cloning and characterization of a vacuolar H+-pyrophosphatase gene, SsVP, from the halophyte Suaeda salsa and its overexpression increases salt and drought tolerance of Arabidopsis. Plant Mol Biol 60:41–50
Haydon MJ, Cobbett CS (2007) A novel major facilitator superfamily protein at the tonoplast influences zinc tolerance and accumulation in Arabidopsis. Plant Physiol 143:1705–1719
Huang SB, Spielmeyer W, Lagudah ES, James RA, Platten JD, Dennis ES, Munns R (2006) A sodium transporter (HKT7) is a candidate for Nax1, a gene for salt tolerance in durum wheat. Plant Physiol 142:1718–1727
James RA, Davenport RJ, Munns R (2006) Physiological characterization of two genes for Na+ exclusion in durum wheat, Nax1 and Nax2. Plant Physiol 142:1537–1547
Kane NA, Danyluk J, Tardif G, Ouellet F, Laliberté JF, Limin AE, Fowler DB, Sarhan F (2005) TaVRT-2, a member of the StMADS-11 clade of flowering repressors, is regulated by vernalization and photoperiod in wheat. Plant Physiol 138:2354–2363
Kane NA, Agharbaoui Z, Diallo AO, Adam H, Tominaga Y, Ouellet F, Sarhan F (2007) TaVRT2 represses transcription of the wheat vernalization gene TaVRN1. Plant J 51:670–680
Kim CA, Bowie JU (2003) SAM domains: uniform structure, diversity of function. Trends Biochem Sci 28:625–628
Krebs M, Beyhl D, Görlich E, Al-Rasheid KAS, Marten I, Stierhof YD, Hedrich R, Schumacher K (2010) Arabidopsis V-ATPase activity at the tonoplast is required for efficient nutrient storage but not for sodium accumulation. Proc Natl Acad Sci USA 107:3251–3256
Li J, Yang H, Peer WA, Richter G, Blakeslee J, Bandyopadhyay A, Titapiwantakun B, Undurraga S, Khodakovskaya M, Richards EL, Krizek B, Murphy AS, Gilroy S, Gaxiola R (2005) Arabidopsis H+-ATPase AVP1 regulates auxin-mediated organ development. Science 310:121–125
Li Z, Baldwin CM, Hu Q, Liu H, Luo H (2010) Heterologous expression of Arabidopsis H+-pyrophosphatase enhances salt tolerance in transgenic creeping bentgrass (Agrostis stolonifera L.). Plant, Cell Environ 33:272–289
Ligaba A, Katsuhara M (2010) Insights into the salt tolerance mechanism in barley (Hordeum vulgare) from comparisons of cultivars that differ in salt sensitivity. J Plant Res 123:105–118
Liu J, Zhu JK (1997) An Arabidopsis mutant that requires increased calcium for potassium nutrition and salt tolerance. Proc Natl Acad Sci USA 94:14960–14964
Liu J, Zhu JK (1998) A calcium sensor homolog required for plant salt tolerance. Science 280:1943–1945
Maeshima M (2000) Vacuolar H+-pyrophosphatase. Bioch Biophys Acta—Biomembranes 1465:37–51
Matsumoto T, Tanaka T, Sakai H et al (2011) Comprehensive sequence analysis of 24,783 barley full-length cDNAs derived from 12 clone libraries. Plant Physiol 156:20–28
Mayer KFX, Taudien S, Martis M et al (2009) Gene content and virtual gene order of barley chromosome 1H. Plant Physiol 151:496–505
Mayer KFX, Martis M, Hedley PE et al (2011) Unlocking the barley genome by chromosomal and comparative genomics. Plant Cell 23:1249–1263
Muchhal US, Pardo JM, Raghothama KG (1996) Phosphate transporters from the higher plant Arabidopsis thaliana. Proc Natl Acad Sci USA 93:10519–10523
Munns R (2007) Utilizing genetic resources to enhance productivity of salt-prone land. CAB Rev Perspect Agric Vet Sci Nutrit Nat Res 2:9. doi:10.1079/PAVSNNR20072009
Munns R, James RA (2003) Screening methods for salinity tolerance: a case study with tetraploid wheat. Plant Soil 253:201–218
Munns R, James RA, Xu B et al (2012) Wheat grain yield on saline soils is improved by an ancestral Na+ transporter gene. Nat Biotechnol 30:360–364
Pao SS, Paulsen IT, Saier MH Jr (1998) Major facilitator superfamily. Microbiol Mol Biol Rev 62:1–34
Pasapula V, Shen G, Kuppu S et al (2011) Expression of an Arabidopsis vacuolar H+-pyrophosphatase gene (AVP1) in cotton improves drought- and salt tolerance and increases fibre yield in the field conditions. Plant Biotechnol J 9:88–99
Poustini K, Siosemardeh A (2004) Ion distribution in wheat cultivars in response to salinity stress. Field Crops Res 85:125–133
Ren ZH, Gao JP, Li LG, Cai XL, Huang W, Chao DY, Zhu MZ, Wang ZY, Luan S, Lin HX (2005) A rice quantitative trait locus for salt tolerance encodes a sodium transporter. Nat Genet 37:1141–1146
Rivandi J, Miyazaki J, Hrmova M, Pallotta M, Tester M, Collins NC (2011) A SOS3 homologue maps to HvNax4, a barley locus controlling an environmentally sensitive Na+ exclusion trait. J Exp Bot 62:1201–1216
Saier MH Jr, Beatty JT, Goffeau A et al (1999) The major facilitator superfamily. J Mol Microbiol Biotechnol 1:257–279
Saier MH Jr, Tran CV, Barabote RD (2006) TCDB: the transporter classification database for membrane transport protein analyses and information. Nucleic Acids Res 34:D181–D186
Sauer N, Stolz J (1994) SUC1 and SUC2: two sucrose transporters from Arabidopsis thaliana; expression and characterization in baker’s yeast and identification of the histidine-tagged protein. Plant J 6:67–77
Schulte D, Ariyadasa R, Shi B et al (2011) BAC library resources for map-based cloning and physical map construction in barley (Hordeum vulgare L.). BMC Genomics 12:247. doi:1471-2164/12/247
Shavrukov Y, Gupta NK, Miyazaki J, Baho MN, Chalmers KJ, Tester M, Langridge P, Collins NC (2010a) HvNax3 – a locus controlling shoot sodium exclusion derived from wild barley (Hordeum vulgare ssp. spontaneum). Funct Integr Genomics 10:277–291
Shavrukov Y, Gupta NK, Chalmers KJ, Tester M, Langridge P (2010b) Identification of a QTL on chromosome 7H for sodium exclusion from wild barley, Hordeum spontaneum. In: Ceccarelli S, Grando S (eds) Proceedings of the 10th international barley genetics symposium. ICARDA, Aleppo, pp 241–247
Smith FW, Ealing PM, Dong B, Delhaize E (1997) The cloning of two Arabidopsis genes belonging to a phosphate transporter family. Plant J 11:83–92
Stolz J, Stadler R, Opekarova M, Sauer N (1994) Functional reconstitution of the solubilized Arabidopsis thaliana STP1 monosaccharide-H+ symporter in lipid vesicles and purification of the histidine tagged protein from transgenic Saccharomyces cerevisiae. Plant J 6:225–233
Szücs P, Karsai I, von Zitzewitz J, Mészáros K, Cooper LLD, Gu YQ, Chen THH, Haeys PM, Skinner JS (2006) Positional relationships between photoperiod response QTL and photoreceptor and vernalization genes in barley. Theor Appl Genet 112:1277–1285
Tanaka Y, Chiba K, Maeda M, Maeshima M (1993) Molecular cloning of cDNA for vacuolar membrane proton-translocating inorganic pyrophosphatase in Hordeum vulgare. Biochem Biophys Res Comm 190:1110–1114
Tsay YF, Schroeder JI, Feldmann KA, Crawford NM (1993) The herbicide sensitivity gene CHL1 of Arabidopsis encodes a nitrate-inducible nitrate transporter. Cell 72:705–713
Ueda A, Kathiresan A, Bennett J, Takabe T (2006) Comparative transcriptome analyses of barley and rice under salt stress. Theor Appl Genet 112:1286–1294
Vasekina AV, Yershov PV, Reshetove OS, Tikhonova TV, Lunin VG, Trofimova MS, Babakov AV (2005) Vacuolar Na+/H+ antiporter from barley: identification and response to salt stress. Biochem (Moscow) 70:100–107
Vincill ED, Szczyglowski K, Roberts DM (2005) GmN70 and LjN70. Anion transporters of the symbiosome membrane of nodules with a transport preference for nitrate. Plant Physiol 137:1435–1444
Yan L, Fu D, Li C, Blechl A, Tranquilli G, Bonafede M, Sanchez A, Valarik M, Yasuda S, Dubcovsky J (2006) The wheat and barley vernalization gene VRN3 is an orthologue of FT. Proc Natl Acad Sci USA 103:19581–19586
Yeo AR, Flowers TJ (1986) Salinity resistance in rice (Oryza sativa L.) and a pyramiding approach to breeding varieties for saline soils. Aust J Plant Physiol 13:161–173
Zhao FY, Zhang XJ, Li PH, Zhao YX, Zhang H (2006) Co-expression of the Suaeda salsa SsNHX1 and Arabidopsis AVP1 confer greater salt tolerance to transgenic rice than the single SsNHX1. Mol Breed 17:341–353
Zhou G, Johnson P, Ryan PR, Delhaize E, Zhou M (2012) Quantitative trait loci for salinity tolerance in barley (Hordeum vulgare L.). Mol Breed 29:427–436
Zhu GY, Kinet JM, Lutts S (2001) Characterization of rice (Oryza sativa L.) F3 populations selected for salt resistance I. Physiological behaviour during vegetative growth. Euphytica 121:251–263
Acknowledgments
This work was supported by funding to the ACPFG from the ARC, GRDC, and the South Australian government. We gratefully acknowledge Jason Eglinton (Barley Breeding Lab) and Ken Chalmers (Molecular Marker Lab) for the genetic resources used as the basis for this project.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Shavrukov, Y., Bovill, J., Afzal, I. et al. HVP10 encoding V-PPase is a prime candidate for the barley HvNax3 sodium exclusion gene: evidence from fine mapping and expression analysis. Planta 237, 1111–1122 (2013). https://doi.org/10.1007/s00425-012-1827-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00425-012-1827-3