Abstract
Background
GIST is the most common mesenchymal tumor of gastrointestinal tract and is more frequent in stomach. Its main mutations affect KIT and PDGFRA genes. Full genetic analysis panels are currently used to study mutations in GIST and other tumors. Considering that in gastric GIST KIT gene mutations in exon 11 are sensitive to IM whereas PDGFRΑ gene mutations in exon 18 (D842V) are resistant to the same drug, the aim of this study is to focus on these two molecular targets as a short alternative panel for predicting therapeutic response in gastric GIST which might optimize resources.
Methods
The genotypes of 38 cases of primary GIST were determined by performing bidirectional DNA sequencing.
Results
Exon 11 of KIT gene showed mutations in 65.3% and the exon 18 of PDGFRA gene showed 9% of cases. So it was possible to determine a subgroup of tumors which presented mutations in KIT exon 11 and PDGFRA exon 18.
Conclusion
Considering all of the foregoing analyzed globally, the application of short panel has impact on the cost and time of release of results to the physician, allowing a rapid approach to patients eligible for treatment with the target therapy.
Similar content being viewed by others
Introduction
Predictive markers of therapeutic response are biomarkers that provide upfront information to physicians as whether or not a patient can be beneficiary from a specific therapy. Such data is essential to clinical oncologists to guide therapy (Duffy et al. 2011).
Due the biological factors as heterogeneity (Almendro et al. 2013), only few patients with a particular type of tumor which show specific mutations can be treated with target therapy. Currently, it is known the response rate from several advanced tumors to target therapy.
Target therapies have efficacy in minority of patients. Therefore, detection of patients likely to respond to a specific treatment would be of great clinical value (Lasota and Miettinen 2006).
Gastrointestinal Stromal Tumor (GIST) is rare and represents less than 1% of all gastrointestinal tumors. It is the most frequent gastrointestinal mesenchymal visceral neoplasm (Miettinen et al. 2002). Approximately 60% of GISTs occur in stomach and have better prognosis than tumors located in other sites as small intestine, duodenum and rectum (Miettinen and Lasota 2011). Studies have shown that GISTs bear mutations in approximately 85% of cases, most of them related to kinase tyrosine KIT (70–80%) or PDGFRΑ (5–8%) genes (Cerski et al. 2011). About 15% of GISTs do not present detectable mutations (Rubin et al. 2001) (Fig. 1). These tumors may be sensitive or resistant to tyrosine kinase inhibitors depending on mutated exon. In fact, there is a link between gene, mutated exon, tumor location and prognostic factors (Dematteo et al. 2002). Most primary tumor mutations of KIT gene affect exon 11 (67–75%) and are related to poor prognosis. However, it has been demonstrated that GISTs show responsiveness of approximately 83.5% to tyrosine kinase inhibitor imatinib mesilate (IM) (Heinrich et al. 2008; Demetri et al. 2004),. Moreover, mutations in PDGFRΑ gene mainly occur in gastric tumors and almost exclusively in exon 18 followed by few mutations in exon 12 and exon 14 (Demetri 2001). The most commonly reported mutation in exon 18 is a substitution of a single nucleotide known as Asp842Val (D842V) which is primarily resistant to tyrosine kinase inhibitors (Cassier et al. 2012). Nowadays, several mutation detection methods have been used as conventional direct sequencing (Baskin et al. 2016) and next generation sequence (NGS). Both are sequenced gene panels involving several genes related to various tumors including GIST (Fisher et al. 2016). However, these procedures are very expensive, requiring high-cost equipment and qualified training for technical staff. Formalin-fixed paraffin-embedded samples (FFPE) are very common practices in pathology laboratories worldwide. This type of storage is extremely important for prospective studies and is the largest source of samples for pathology laboratories. Nevertheless, it often difficult the use of DNA for molecular assays (Greer et al. 1994).
Considering that in gastric GIST KIT gene mutations in exon 11 are sensitive to IM whereas PDGFRΑ gene mutations in exon 18 (D842V) are resistant to the same drug, the aim of this study is to focus on these two molecular targets as triage tool for detection of a subgroup of patients more likely to be responsive to IM (Fig. 1) (Gheorghe et al. 2014).
Materials and methods
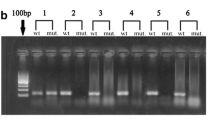
We reviewed 38 cases of primary GIST tumors (FFPE), all from gastric sites. The blocks were made available from the Pathology department of the Universidade Federal de São Paulo - UNIFESP/EPM, São Paulo, Brazil. The storage period ranged from 2000 to 2010. All tumors showed morphological characteristics typical of GIST, CD117 positive in immunohistochemistry and they had not received specific treatment before resection. It was analyzed 10-μm cuts from gastric GISTs fixed in formalin and embedded in paraffin. The sections were deparaffinized tissue was manually macrodissected. For DNA extraction, we used a Qiagen QIAamp® DNA Micro commercial kit, according to the manufacturer’s instructions. DNA quantification was calculated by Nanodrop 2000c Thermo scientific in A260. The PCR reactions were performed in a thermal cycler (Mastercycler® personal; Eppendorf, Hamburg, Germany). Three different primer combinations were used to study exon 11 of the KIT gene. Primers were used at a concentration of 0.4 μM per reaction, generating the following fragments: 174 bp (P1 and P2), 196 bp (P3 and P4) and 215 bp (P5 and P6) (Merkelbach-Bruse et al. 2010) (Fig. 2):
P1:F-5’GATCTATTTTTCCCTTTCTCC3’ and P2:R-5’ AGCCCCTGTTTCATACTGAC 3′; P3:F-5’GATCTATTTTTCCCTTTCTCC3’ and P4:R-5’TACCCAAAAAGGTGACATGG3’; P5:F-5’CCAGAGTGCTCTAATGACTG3’ and P6:R-5’AGCCCCTGTTTCATACTGAC3’. For amplification of the 218 bp fragment of the PDGFRA gene, we used the following forward (F) and reverse (R) sequences:
P7:F-5’CAGCTACAGATGGCTTGATC3‘ and R5’GAAGGAGGATGAGCCTGAC3‘. For all reactions, we used Master Mix (Qiagen®), and the concentration of DNA used in each reaction varied between 10 and 30 ng. Amplification of KIT gene exon 11 involved one cycle at 94 °C for one min, 94 °C for 40 s, 54 °C for 40 s, one extension period of 72 °C for 40 s and a final extension of 72 °C for 5 min. PDGFRA gene exon 18 amplification used the same PCR conditions as above, but at 57 °C. These reactions were adapted from Merkelbach-Bruse et al. (2010). The amplified fragments were visualized in 1.5% agarose gel stained with “Gel Red™ (Biotium®)”. Product purification was performed with a QIAquick PCR purification kit (Qiagen, Hilden, Germany). Tape sequencing (forward and reverse) was performed with Big Dye Terminator Cycle Sequencing Ready Reaction Kit v1.0 (Applied Biosystems, Foster City, CA, USA). The primer concentration was 0.4 μM each. We used between 1 and 4 μl of PCR product, depending on the intensity of the band observed. The PCR program conditions for the sequencing cycle of KIT gene exon 11 were: 96 °C for 1 min, 96 °C for 10 s, annealing at 55 °C for 5 s, and extension at 60 °C for 4 min. The conditions for PDGFRΑ gene exon 18 were the same except for the annealing temperature of 60 °C. Sequencing was performed on ABI 3100 Genetic Analyzer equipment (Applied Biosystems, Foster City, CA, USA).
In addition, duplicates were made and the two DNA strands (forward and reverse) were sequenced for validation of the technique.
Results
The results of the evaluation of purity of DNA extracted from the samples were close to 1.8. This indicates that DNA extraction by Qiagen QIAamp® DNA Micro commercial kit was adequate (Funabashi et al. 2012). A relationship between these values and the amplification and sequencing could be not established.
Exon 11 KIT gene PCR
Among the primer sets used, the one with the best response to the material was the one that produced 174 bp fragments in 37/38 (97.4%) of cases, followed by the 196 bp fragment in 18/38 (47.3%) and the 215 bp fragment in 20/38 (52.6%).
Thereby, 37 samples were amplified successfully; however, 31.5% could not be sequenced. One of the samples used had two simultaneous mutations so was considered 27 mutations in total. The number and types of mutations found in exon 11 of the KIT gene were 5/27 (18.5%) deletions, 9/27 (33.3%) substitutions, 1/27 (3.7%) insertions and 2/27 (7.5%) duplications, with no mutation in 9/27 (33.3%). The type of mutation, genomic location and the position of the codon in the 38 samples used in the study are shown in Table 1.
Exon 18 PCR PDGFRΑ gene
Of all 38 samples, 22/38 (58%) were amplified with the 213 bp primers. Of these, 11/22 (50%) had silent p.V824 V mutations e 2/22 (9.1%) of tumors carried mutations in codon 842. Finally, in 41% (9/22) of sample, there were no detected mutations (Table 1).
Sanger sequencing
The method used was the Sanger sequencing which is known as the gold standard method for GIST genotyping (Martin-Broto et al. 2017). The sensitivity and specificity of the Sanger sequencing method, as well in the complete panel as the short panel is the same, because it is the same method. In general, in terms of accuracy for determination of specific mutations, Sanger Sequencing presents similar accuracy to other methods (Zhang et al. 2015).
According to the criteria listed below, we mention some reasons why using the Short panel (Table 2).
Some authors argue about the weaknesses of the method and its implications on the results of the research (Table 3).
Although NGS methods have many advantages in terms of speed, cost, and parallelism, the accuracy and read length of Sanger sequencing is still superior and has confined the use of NGS mainly to sequencing genomes (Verma et al. 2017). Therefore, it is necessary that the tools that already implanted are not discarded but adapted or improved.
Discussion
The conventional Sanger sequencing used as a method of detecting mutations in GIST takes time, is laborious and requires specialized technical knowledge for the analysis of mutations (Agaimy et al. 2013; Schöffski et al. 2016).
Traditionally, full-panel sequencing approaches in GISTs involve KIT gene and PDGFRA gene (Miettinen and Lasota 2006). Multi-exon strategy allows tracking of different mutations, thus providing a broad spectrum of genetic profile of patients. However, such approach is very expensive and show several limitations in FFPE samples (Blanke and Huse 2010; Guerin et al. 2015).
Pre-analytic issues and high-cost equipment are limiting factors for large use of DNA sequencing for medical assistance purposes. Despite these difficulties, data provided by sequencing are essential to identify a subgroup of patients which will benefit from IM therapy.
Considering that the complete panel is extensive, involving mutations in four exons (9, 11, 13 and 17) of the KIT gene and three exons (12, 14 and 18) of the PDGFRA gene, the use of short sequencing panel may be a practical, fast and economical alternative of evaluating tumor sensitiveness to MI.
Furthermore, for each analyzed exon in addition to the duplicates, independent PCR reactions are required for both DNA strands of the sample. Therefore, only one GIST sample requires eight sequencing reactions for the KIT gene and six other sequence reactions if the PDGFRA gene exons are also analyzed.
For that reason, it would be important that laboratories with less demand and/or located outside the large urban centers, could benefit from simplified screening protocols that focus on the detection of mutations in specific exons to meet the need for therapeutic response (Zhang et al. 2015).
Currently, an alternative to Sanger sequencing is the next generation sequencing (NGS), this technique a solid potential as molecular test in several types of cancer (Gao et al. 2016). The large sequencing capability of NGS platforms allows detecting all mutation information in multiple samples at the same time. However, the NGS does not use the same work steps of the conventional Sanger method, and in addition, it presents a high cost, besides requiring many times the confirmation of the findings by the Sanger sequencing (Johnston et al. 2012; Pandey et al. 2016; Diekstra et al. 2015), making it difficult to use them in the diagnostic routine. In addition, the mutational spectrum is much broader generating a volume of information unnecessary for the therapeutic routine.
Different authors also discuss the limitations of Sanger’s method in relation to other methods and how this implies the results of the analysis. However, only knowing their limitations will it be possible to establish protocols that minimize them so that it can continue to be a benchmark for other methods.
Analyzing other sequencing methods, specifically the analytical validation tests on NGS platforms, is still extremely challenging for pathology laboratories, for although there are many affordable NGS instruments and easy to access, there are still no standardized guidelines for clinical validation trials NGS (Rathi et al. 2017).
In addition, patterns related to the input of FFPE material were defined, both for use in NGS and for Sanger sequencing. Thus, the reliability of mutation analysis could be improved by manual inspection of sequence data (de Leng et al. 2016).
It is possible that some institutions already select specific exons in the practice of diagnostic routine. However, this study provides a theoretical basis for this practice that would have sensitive implications in the management of GIST patients.
KIT gene: exon 11
Exon 11 of the KIT gene has a variable mutation profile that is distributed in different codons throughout the exon, sometimes hindering analysis and interpretation of results. Combined with pre-analytic issues as non-controlled fixation and embedding processes, full sequencing analysis becomes a challenging task. The type frequency and of mutations found in exon 11 of KIT gene were deletions (5/27–18.5%), substitutions (9/27–33.3%), insertions (1/27–3.7%) and duplications, (2/27–7.5%). No mutations were found in 9/27 of cases (33.3%).
Duplications have been reported among mutations found in this exon and are mainly associated with gastric site, female gender and benign behavior (Antonescu et al. 2003). We have found two cases of duplication in our set of cases (i.e., 7.5%), rate similar to values reported in literature (Palmirotta et al. 2013; Minárik et al. 2013).
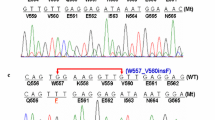
Deletions are often associated with worse prognosis, mainly if involving Trp557 and/or Lys558. These codons are regarded as the first hot spot of mutations in this exon and are associated to aggressive biological behavior (Wardelmann et al. 2003; Singer et al. 2002). Nevertheless, they present better response to IM. In this study, 16% (4/25) of cases had in-frame deletions. Three of them affected codons 557 and 558, one sample with p.W557_E561del5 and two samples with p.557_558del2. There was also a homozygous in-frame deletion in codon 579 (p.D579del; Fig. 3). The highest frequency of in-frame deletions in the terminal 5′ region of exon 11 involves codons 550–560 (Wardelmann et al. 2003; Wardelmann et al. 2007). The 3′ region of KIT gene exon 11 is considered the second most mutated hotspot (Antonescu et al. 2003). Insertions and duplications of large genomic regions most often occur in such region even though the number of nucleotides varies. Moreover, this type of mutation may be difficult to visualize as it does not present changes in the exon body (Lasota et al. 2003; Iesalnieks et al. 2005). We also observed a case like this in our sample: the insertion of 48 bp p.F591_G592ins16 at the 3′ terminal. Although this mutation is not directly involved in the body of exon 11, it could affect the protein structure due to its size and position. We found eight missense substitutions in the analyzed tumors with varied distribution. Substitutions have been found to be the most common mutations in this exon and most often occupy codons 557, 559, 560 and 576. The p.V559D mutation was found in two cases (Silva et al. 2010; Antonescu et al. 2005). When deleted, this codon may be associated with metastatic disease but the p.V559D substitution was not specifically associated with such behavior (Wardelmann et al. 2003). The missense substitution Y570F (c.1708_1710), also found in this study, was previously described in a study analyzing GIST mutations in a Portuguese population, but was not associated to gastric site (Gomes et al. 2008). Codon 572 is often cited in literature, presenting with several types of mutations. While other amino acids have been reported to occupy this codon in GIST and in other tumors (Rossi et al. 2010), the substitution of Aspartic acid (Asp/D) by Valine (Val/V) found in one of our patients had not been previously described (p.D572V). The p.P573M mutation has also not been described previously in gastric GIST tumors, but this codon is involved in several other mutations involving non gastric GIST (Antonescu et al. 2003) and melanoma (Omholt et al. 2011). The missense mutation p.F584 L c.1752 T > G has been described in GIST (Minárik et al. 2012), but it may also be present in other tumors, such as primary adenoid cystic carcinoma of salivary glands (Vila et al. 2009) and breast carcinoma (Hussain et al. 2012).
Finally, the p.P585T mutation was previously described in GIST in a study associating GIST mutations and VEGF expression (de Oliveira et al. 2011; Mazzola et al. 2008). One sample showed two mutations for KIT gene exon 11 -- p.E561L and p.Y578C – which had not been described to date. However, other substitutions in codon p.E561 involving different tissues, such as p.E561K in hematopoietic tissues (Hongyo et al. 2005) and GIST (Hanada et al. 2006), p.E561G in melanoma (Kong et al. 2011) and p.E561D in GIST (Nakajima et al. 2009), have already been described.
PDGFRA gene: exon 18
We obtained high-quality samples by using this primer designed for exon 18 of PDGFRA (Fig. 4). However, the size of fragment generated by this set of primers may also have affected the PCR amplification results (13/22, 59%), confirming the need for adjustments to PCR protocol and sequencing for this exon as it was done in KIT exon 11. In this study, 2/22 (9.1%) of tumors carried mutations in codon 842 of PDGFRA exon 18, which is considered a hotspot for this exon. No mutations were shown in 9/22 (40.9%) of cases.
Two samples (15 and 34) presented KIT exon 11 and PDGFRA exon 18 gene mutations. The explanation for such findings may be related to intratumoral heterogeneity and although some authors have previously reported that mutations in KIT and PDGFRA genes are mutually exclusive, morphological heterogeneity is sometimes evident. Therefore, further investigations are required to better elucidate this issue (Kumar et al. 2014; Alic et al. 2014).
However, a few points should be paid attention. First, our study was descriptive and is an initial study only demonstrates the feasibility of the method in FFPE samples. Second, the comparison between original panel and the panel will be proposed in future studies.
FFPE samples and quality
Noteworthy is the fact that our sample included tumors processed in different laboratories, which precluded the necessary quality control procedures. We have found that 18/38 (47.5%) of samples were sequenced for both exons studied. This finding indicates the quality and suitability of performed methods. However, there have been individual responses of different samples to the same protocol. Accordingly, it is difficult to obtain a well succeed sequencing procedure in diagnostic laboratory without making fine adjustments in protocols. The literature points that the average DNA length obtained after extraction is 300-400 bp but this value is lower in FFPE samples (Barcelos et al. 2008; Casale et al. 2010), which explains the increase in amplification from 52.6 to 97.4% obtained in this study. This increased value was only possible when we repositioned primers from 215 bp amplicons to 174 bp amplicons (Merkelbach-Bruse et al. 2010).
Reduction in fragment size does not guarantee PCR reaction efficiency since primers that generated 196 bp fragments presented 47.3% positivity, which was lower than presented by the 215 bp sample. That is true once there are regions in the genome that may be compromised by fixation. In the case of exon 11, mutations could be located in such regions, thus preventing primer annealing. Besides DNA extraction, PCR and sequencing protocols were the same. About 21% (8/38) of samples could not be sequenced for both genes and some samples could be sequenced only for one of them. This may be related to variations in pre-analysis step, such as batch and quality of reagents, since the study period involved years of collection and storage. This phenomenon also occurred with other authors who worked with FFPE samples that turned out to be unusable for molecular analysis (Origone et al. 2013).
Study limitations
Some limiting aspects deserve to be highlighted. The lack of amplification of some FFPE samples represents an important limitation of our study. In addition, studies involving a greater number of samples are necessary since our results are based on a rather reduced casuistry. In the future, an extended investigation produces relevant results to provide the basis for operating protocols.
Conclusion
The use of targeted therapies requires knowledge of specific mutations for the patient to truly benefit from the treatment. Establishing the genotypic profile of GIST can determine response to treatment and modify the entire treatment schedule.
Considering all the foregoing analyzed globally, we conclude that application of short panel has impact on the cost and time of release of results to the physician, allowing a rapid approach to patients eligible for treatment with the target therapy.
Abbreviations
- FFPE:
-
Formalin-fixed paraffin-embedded samples
- GIST:
-
Gastrointestinal Stromal Tumor
- IM:
-
Imatinib mesilate
- NGS:
-
Next generation sequence
References
Agaimy A et al (2013) Value of epithelioid morphology and PDGFRA immunostaining pattern for prediction of PDGFRA mutated genotype in gastrointestinal stromal tumors (GISTs). Int J Clin Exp Pathol 6(9):1839–1846
Alic L, Niessen WJ, Veenland JF (2014) Quantification of heterogeneity as a biomarker in tumor imaging: a systematic review. PLoS One 9(10):e110300
Almendro V, Marusyk A, Polyak K (2013) Cellular heterogeneity and molecular evolution in cancer. Annu Rev Pathol 8:277–302
Alvarez-Garcia V et al (2018) A simple and robust real-time qPCR method for the detection of PIK3CA mutations. Sci Rep 8(1):4290
Antonescu CR et al (2003) Association of KIT exon 9 mutations with nongastric primary site and aggressive behavior: KIT mutation analysis and clinical correlates of 120 gastrointestinal stromal tumors. Clin Cancer Res 9(9):3329–3337
Antonescu CR et al (2005) Acquired resistance to imatinib in gastrointestinal stromal tumor occurs through secondary gene mutation. Clin Cancer Res 11(11):4182–4190
Barcelos D, Franco MF, Leão SC (2008) Effects of tissue handling and processing steps on PCR for detection of Mycobacterium tuberculosis in formalin-fixed paraffin-embedded samples. Rev Inst Med Trop Sao Paulo 50:321–326
Baskin Y et al (2016) PDGFRA and KIT mutation status and its association with Clinicopathological properties, including DOG1. Oncol Res 24(1):41–53
Blanke CD, Huse DM (2010) Cost effectiveness of tyrosine kinase inhibitor therapy in metastatic gastrointestinal stromal tumors. J Med Econ 13(4):681–690
Casale V et al (2010) Optimisation of postmortem tissue preservation and alternative protocol for serotonin transporter gene polymorphisms amplification in SIDS and SIUD cases. Exp Mol Pathol 88(1):202–205
Cassier PA et al (2012) Outcome of patients with platelet-derived growth factor receptor alpha-mutated gastrointestinal stromal tumors in the tyrosine kinase inhibitor era. Clin Cancer Res 18(16):4458–4464
Cerski MR et al (2011) Exon 11 mutations, Ki67, and p16(INK4A) as predictors of prognosis in patients with GIST. Pathol Res Pract 207(11):701–706
de Leng WW et al (2016) Targeted next generation sequencing as a reliable diagnostic assay for the detection of somatic mutations in Tumours using minimal DNA amounts from formalin fixed paraffin embedded material. PLoS One 11(2):e0149405
de Oliveira AT et al (2011) Lymphangiogenic VEGF-C and VEGFR-3 expression in genetically characterised gastrointestinal stromal tumours. Histol Histopathol 26(12):1499–1507
Dematteo RP et al (2002) Clinical management of gastrointestinal stromal tumors: before and after STI-571. Hum Pathol 33(5):466–477
Demetri GD (2001) Targeting c-kit mutations in solid tumors: scientific rationale and novel therapeutic options. Semin Oncol 28(5 Suppl 17):19–26
Demetri GD et al (2004) Case records of the Massachusetts General Hospital. Weekly clinicopathological exercises. Case 32-2004. A 68-year-old man with a large retroperitoneal mass. N Engl J Med 351(17):1779–1787
Diekstra A et al (2015) Translating sanger-based routine DNA diagnostics into generic massive parallel ion semiconductor sequencing. Clin Chem 61(1):154–162
Duffy MJ, O'Donovan N, Crown J (2011) Use of molecular markers for predicting therapy response in cancer patients. Cancer Treat Rev 37(2):151–159
Fisher KE et al (2016) Clinical validation and implementation of a targeted next-generation sequencing assay to detect somatic variants in non-small cell lung, melanoma, and gastrointestinal malignancies. J Mol Diagn 18(2):299–315
Funabashi KS et al (2012) DNA extraction and molecular analysis of non-tumoral liver, spleen, and brain from autopsy samples: the effect of formalin fixation and paraffin embedding. Pathol Res Pract 208:584–591. https://doi.org/10.1016/j.prp.2012.07.001
Gao J et al (2016) Comparison of next-generation sequencing, quantitative PCR, and sanger sequencing for mutation profiling of EGFR, KRAS, PIK3CA and BRAF in clinical lung tumors. Clin Lab 62(4):689–696
Gheorghe M et al (2014) Clinical and therapeutic considerations of GIST. J Med Life 7(2):139–149
Giardina T et al (2018) Implementation of next generation sequencing technology for somatic mutation detection in routine laboratory practice. Pathology 50(4):389–401
Gomes AL et al (2008) Molecular alterations of KIT and PDGFRA in GISTs: evaluation of a Portuguese series. J Clin Pathol 61(2):203–208
Greer CE, Wheeler CM, Manos MM (1994) Sample preparation and PCR amplification from paraffin-embedded tissues. PCR Methods Appl 3(6):S113–S122
Guerin A et al (2015) The economic burden of gastrointestinal stromal tumor (GIST) recurrence in patients who have received adjuvant imatinib therapy. J Med Econ 18(3):241–248
Hanada N et al (2006) A case of recurrent GIST successfully treated with low-dose imatinib mesilate. Gan To Kagaku Ryoho 33(6):799–801
Heinrich MC et al (2008) Correlation of kinase genotype and clinical outcome in the north American intergroup phase III trial of imatinib mesylate for treatment of advanced gastrointestinal stromal tumor: CALGB 150105 study by Cancer and leukemia group B and southwest oncology group. J Clin Oncol 26(33):5360–5367
Hongyo T et al (2005) p53, K-ras, c-kit and beta-catenin gene mutations in sinonasal NK/T-cell lymphoma in Korea and Japan. Oncol Rep 13(2):265–271
Hussain SR et al (2012) Screening of the c-kit gene missense mutation in invasive ductal carcinoma of breast among north Indian population. Mol Biol Rep 39(9):9139–9144
Iesalnieks I et al (2005) Factors associated with disease progression in patients with gastrointestinal stromal tumors in the pre-imatinib era. Am J Clin Pathol 124(5):740–748
Ishige T et al (2016) Locked nucleic acid probe enhances sanger sequencing sensitivity and improves diagnostic accuracy of high-resolution melting-based KRAS mutational analysis. Clin Chim Acta 457:75–80
Johnston JJ et al (2012) Secondary variants in individuals undergoing exome sequencing: screening of 572 individuals identifies high-penetrance mutations in cancer-susceptibility genes. Am J Hum Genet 91(1):97–108
Kong Y et al (2011) Large-scale analysis of KIT aberrations in Chinese patients with melanoma. Clin Cancer Res 17(7):1684–1691
Kumar A et al (2014) Deep sequencing of multiple regions of glial tumors reveals spatial heterogeneity for mutations in clinically relevant genes. Genome Biol 15(12):530
Lasota J, Miettinen M (2006) KIT and PDGFRA mutations in gastrointestinal stromal tumors (GISTs). Semin Diagn Pathol 23(2):91–102
Lasota J et al (2003) Gastrointestinal stromal tumors with internal tandem duplications in 3′ end of KIT juxtamembrane domain occur predominantly in stomach and generally seem to have a favorable course. Mod Pathol 16(12):1257–1264
Martin-Broto J et al (2017) Gastrointestinal stromal tumors (GISTs): SEAP-SEOM consensus on pathologic and molecular diagnosis. Clin Transl Oncol 19(5):536–545
Martinuzzi C et al (2016) A combination of immunohistochemistry and molecular approaches improves highly sensitive detection of BRAF mutations in papillary thyroid cancer. Endocrine 53(3):672–680
Mazzola P et al (2008) Epidemiology and molecular biology of gastrointestinal stromal tumors (GISTs): a population-based study in the south of Switzerland, 1999-2005. Histol Histopathol 23(11):1379–1386
McEvoy AC et al (2018) Droplet digital PCR for mutation detection in formalin-fixed, paraffin-embedded melanoma tissues: a comparison with sanger sequencing and pyrosequencing. J Mol Diagn 20(2):240–252
Merkelbach-Bruse S et al (2010) Pitfalls in mutational testing and reporting of common KIT and PDGFRA mutations in gastrointestinal stromal tumors. BMC Med Genet 11:106
Miettinen M, Lasota J (2006) Gastrointestinal stromal tumors: review on morphology, molecular pathology, prognosis, and differential diagnosis. Arch Pathol Lab Med 130(10):1466–1478
Miettinen M, Lasota J (2011) Histopathology of gastrointestinal stromal tumor. J Surg Oncol 104(8):865–873
Miettinen M et al (2002) Evaluation of malignancy and prognosis of gastrointestinal stromal tumors: a review. Hum Pathol 33(5):478–483
Minárik G et al (2012) Spectrum of mutations in gastrointestinal stromal tumor patients - a population-based study from Slovakia. APMIS. https://doi.org/10.1111/apm.12019
Minárik G et al (2013) Spectrum of mutations in gastrointestinal stromal tumor patients - a population-based study from Slovakia. APMIS 121(6):539–548
Nakajima T et al (2009) Interstitial cells of Cajal do not harbor c-kit or PDGFRA gene mutations in patients with sporadic gastrointestinal stromal tumors. J Gastroenterol 44(5):426–431
Ofner R et al (2017) Non-reproducible sequence artifacts in FFPE tissue: an experience report. J Cancer Res Clin Oncol 143(7):1199–1207
Omholt K et al (2011) KIT pathway alterations in mucosal melanomas of the vulva and other sites. Clin Cancer Res 17(12):3933–3942
Origone P et al (2013) Molecular characterization of an Italian series of sporadic GISTs. Gastric Cancer 16(4):596–601
Palmirotta R et al (2013) Mutational analysis of gastrointestinal stromal tumors (GISTs): procedural approach for diagnostic purposes. Cancer Genomics Proteomics 10(3):115–123
Pandey RV et al (2016) MutAid: sanger and NGS based integrated pipeline for mutation identification, validation and annotation in human molecular genetics. PLoS One 11(2):e0147697
Rathi V et al (2017) Clinical validation of the 50 gene AmpliSeq Cancer panel V2 for use on a next generation sequencing platform using formalin fixed, paraffin embedded and fine needle aspiration tumour specimens. Pathology 49(1):75–82
Rossi S et al (2010) Molecular and clinicopathologic characterization of gastrointestinal stromal tumors (GISTs) of small size. Am J Surg Pathol 34(10):1480–1491
Rubin BP et al (2001) KIT activation is a ubiquitous feature of gastrointestinal stromal tumors. Cancer Res 61(22):8118–8121
Sabour L, Sabour M, Ghorbian S (2017) Clinical applications of next-generation sequencing in Cancer diagnosis. Pathol Oncol Res 23(2):225–234
Schöffski P et al (2016) Overcoming cost implications of mutational analysis in patients with gastrointestinal stromal tumors: a pragmatic approach. Oncol Res Treat 39(12):811–816
Silva M et al (2010) Chromosome copy number changes carry prognostic information independent of KIT/PDGFRA point mutations in gastrointestinal stromal tumors. BMC Med 8:26
Singer S et al (2002) Prognostic value of KIT mutation type, mitotic activity, and histologic subtype in gastrointestinal stromal tumors. J Clin Oncol 20(18):3898–3905
Tzou PL et al (2018) Comparison of an in vitro Diagnostic Next-Generation Sequencing Assay with Sanger Sequencing for HIV-1 Genotypic Resistance Testing. J Clin Microbiol 56(6). https://doi.org/10.1128/JCM.00105-18
Verma M, Kulshrestha S, Puri A (2017) Genome Sequencing. Methods Mol Biol 1525:3–33
Vila L et al (2009) Identification of c-kit gene mutations in primary adenoid cystic carcinoma of the salivary gland. Mod Pathol 22(10):1296–1302
Wardelmann E et al (2003) Deletion of Trp-557 and Lys-558 in the juxtamembrane domain of the c-kit protooncogene is associated with metastatic behavior of gastrointestinal stromal tumors. Int J Cancer 106(6):887–895
Wardelmann E et al (2007) Mutation analysis of gastrointestinal stromal tumors: increasing significance for risk assessment and effective targeted therapy. Virchows Arch 451(4):743–749
Zhang L et al (2015) Genotyping on ALDH2: comparison of four different technologies. PLoS One 10(3):e0122745
Acknowledgements
The authors would like to thank CAPES Foundation for funding part of the research. The authors also thank Dr. Laercio Gomes Lourenço presented the biopsies studied and the dedicated and skilled staff CTCMol laboratory Cellular and Molecular Therapy Center.
“This manuscript was reviewed by a professional science editor and by a native English-speaking copy editor to improve readability”.
Funding
The Capes Foundation funded this research through the research grant.
Availability of data and materials
Data sharing not applicable to this article as no datasets were generated or analysed during the current study.
Author information
Authors and Affiliations
Contributions
DB was responsible for the design, study design and implementation of the project and drafted the manuscript. RI and ESMI conceived of the study, and participated in its design and coordination and helped to draft the manuscript. KF, MF, AC held the molecular genetic studies, participated in the sequence alignment and drafted the manuscript. FC and LC participated in the histopathological analysis of biopsies and drafted the manuscript. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Ethics approval and consent to participate
The study was approved by the Ethical Committee of Universidade Federal de São Paulo - Plataforma Brasil, reference number 16183.
Consent for publication
“Not applicable”.
Competing interests
The authors declare that they have no competing interests.
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The Creative Commons Public Domain Dedication waiver (http://creativecommons.org/publicdomain/zero/1.0/) applies to the data made available in this article, unless otherwise stated.
About this article
Cite this article
Barcelos, D., Neto, R.A., Cardili, L. et al. KIT exon 11 and PDGFRA exon 18 gene mutations in gastric GIST: proposal of a short panel for predicting therapeutic response. Surg Exp Pathol 1, 8 (2018). https://doi.org/10.1186/s42047-018-0021-8
Received:
Accepted:
Published:
DOI: https://doi.org/10.1186/s42047-018-0021-8