Abstract
Background
The role of forkhead-box A1 (FOXA1) and Androgen receptor (AR) in breast cancer (BC) has been extensively studied. However, the prognostic role of their co-expression in Estrogen receptor positive (ER+) BC has not been investigated so far. The aim of the present study was thus to assess the co-expression (protein and mRNA) of FOXA1 and AR in BC patients, in order to evaluate their prognostic impact according to ER status.
Methods
Immunohistochemical expression of AR and FOXA1 was evaluated on 479 consecutive BC, with complete clinical-pathological and follow up data. Fresh-frozen tissues from 65 cases were available. The expression of AR and FOXA1 with ER was validated using mRNA analyses. Survival and Cox proportional hazard analyses were used to evaluate the relationship between FOXA1, AR and prognosis.
Results
Expression of ER, AR and FOXA1 was observed in 78, 60 and 85% of cases respectively. Most AR+ cases (97%) were also FOXA1+. The level of FOXA1 mRNA positively correlated with level of both AR mRNA (r = 0.8975; P < 0.001) and ER mRNA (r = 0.7326; P < 0.001). In ER+ BC, FOXA1 was associated with a good prognosis independently of AR expression in the three subgroups analyzed (FOXA1+/AR+; FOXA1+/AR-; FOXA1−/AR-). Multivariate analyses confirmed that FOXA1 may provide more information than AR in Disease-Free Interval (DFI) of ER+ BC patients.
Conclusion
Our results suggest that in BC the expression of FOXA1 is directly related to the expression of AR. Despite that, FOXA1 is found as superior predicting marker of recurrences compared to AR in ER+ BC patients.
Similar content being viewed by others
Background
In breast cancer (BC), Estrogen (ER) and Androgen Receptors (AR) regulate cell proliferation and differentiation. They are frequently co-expressed, however AR may be expressed in ER-negative (ER-) BC, where it modulates gene transcription by using regulatory molecules and pathways normally activated by ER [1]. As a result, in ER- BC cells, androgens activate cell proliferation [2], whereas in ER-positive (ER+) cells, androgens inhibit cell proliferation [3, 4]. In line with these data, we demonstrated that patients with AR+/ER+ BC have a better prognosis compared to those affected by AR−/ER+ BC [5, 6].
FOXA1, a member of the forkhead family protein [7], is an important regulator of ER DNA binding and transcription of its target genes [8]. In addition, in both ER+ and ER- BC cells, FOXA1 promotes AR DNA binding [1, 9, 10]. Several studies [11,12,13,14,15,16,17,18,19] evaluated the prognostic role of FOXA1 in BC, and demonstrated that in ER+ BC the expression of FOXA1 is positively correlated with a better prognosis. Indeed, the role of FOXA1/AR co-expression in ER+ BC has not been investigated, although it has been suggested that the relative ratio among FOXA1, ER and AR could influence growth and aggressiveness of cancer cells [20].
The aim of the present study was first to assess the co-expression, at both protein and mRNA levels, of FOXA1 and AR in BC, and then to evaluate their prognostic impact in ER+ BC patients.
Methods
Case series
We collected a series of 479 female patients that underwent surgery for BC from June 1994 to December 2012 at the Breast Unit of the Città della Salute e della Scienza Hospital of Turin, Italy. All patients were treated with surgery, either mastectomy or wide local excision, followed by radiotherapy.
Clinical-pathological data such as age at time of diagnosis, surgery (conserving surgery vs radical mastectomy), type of therapy (hormonal therapy, chemotherapy), type and site of recurrences, histological types, tumor size (< 15 mm vs ≥ 15 mm), nodal involvement, histologic grade and vascular invasion were collected. Medical charts of all patients were reviewed to confirm accuracy of previously recorded data. Tumor slides were re-evaluated to select representative blocks that were used to construct multicore tissue microarrays (TMAs, tissue arrayer Galileo TMA CK 3500, Integrated Systems Engineering Srl, Milan, Italy), as previously described [21].
Immunohistochemistry
To confirm the results of the diagnostic reports, immunohistochemistry (IHC) was performed on TMA sections using an automated slide processing platform (Ventana BenchMark AutoStainer, Ventana Medical Systems, Tucson, AZ, USA) and the following primary antibodies were used: prediluted anti-ER rabbit monoclonal antibody (SP1, Ventana-Roche, Tucson, AZ, USA); prediluted anti-Progesterone receptor (PgR) rabbit monoclonal antibody (1E2, Ventana-Roche); anti-Ki67 monoclonal antibody (MIB1, diluted 1:100 Dako); anti-human c-erbB2 oncoprotein (Ventana Pathway HER-2/Neu-4B5). In addition, AR and FOXA1 expression were tested using anti-AR mouse monoclonal antibody (AR441, diluted 1:50, Dako, Glostrup, Denmark) and prediluted anti-FOXA1 mouse monoclonal antibody (2F83, Ventana-Roche). Positive and negative controls (omission of the primary antibody and IgG-matched serum) were included for each IHC run.
The cut-off value for ER and PgR expression was set at 1%, as suggested by St Gallen Consensus meeting [22], and the same cut-off was also adopted for AR and FOXA1 expression [5]. The percentage of Ki67-positive cells was recorded and the cut-off for dichotomizing tumors with low and high proliferative fraction was established at 20% according to 2013 St Gallen Consensus meeting [23] and also on the basis of the median Ki67 value of our local laboratory [24, 25]. HER2 status was classified as negative (score 0, 1+ and 2+ not amplified) or positive (when scored 3+ by IHC or HER2 amplified by FISH) according to the recommended guidelines for invasive carcinoma [26].
Real-time PCR (qPCR) analysis
To determine the specificity of AR and FOXA1 antibodies, we compared gene expression levels (using qPCR) with IHC results. The relationship between AR and FOXA1 was validated using relative quantification mRNA analyses.
qPCR for AR and FOXA1 mRNA was performed on 65 fresh-frozen BC samples (Fig. 1). Total RNA was extracted from tissues using TRIzol Reagent (Invitrogen Ltd., Paisley, UK) following manufacturer’s instructions. DNase I was added to remove remaining genomic DNA. 1 μg of total RNA was reverse-transcribed with iScript cDNA Synthesis Kit (Bio-Rad Laboratories Inc., Hercules, CA, USA), following manufacturer protocol. Primers (Additional file 1: Table S1) were designed using Beacon Designer 5.0 software according to parameters outlined in the Bio-Rad iCycler Manual. Specificity of primers was confirmed by BLAST analysis. qPCR was performed using a BioRad iQ iCycler Detection System (Bio-Rad Laboratories Inc., Hercules, CA, USA) with SYBR green fluorophore. Reactions were performed in a total volume of 25 μl containing 12.5 μl of IQ SYBR Green Supermix (Bio-Rad Laboratories Inc., Hercules, CA, USA), 1 μl of each primer at 10 μM concentration, and 5 μl of the previously reverse-transcribed cDNA template. The protocol used was as follows: denaturation (95 °C for 5 min) and amplification repeated 40 times (95 °C for 15 s, 60 °C for 30 s). At each run, a melting curve analysis was performed to ensure a single specific amplified product for every reaction. Results were normalized using the Delta-Ct (Δct) method, using β-actin as housekeeping gene. Samples with a Δct ≤ 6 were defined as positive.
Statistical and survival analyses
Pearson’s Chi square test and Student’s t-test were preliminary performed to compare respectively categorical and continuous variables, and to evaluate potential differences in the variable distribution among groups. Test for median and means (Analysis of Variance-ANOVA) were performed. For more than two groups Tukey HSD post-hoc test was performed. Disease-Free Interval (DFI) was calculated from the date of surgical excision of the primary tumor to the date of first disease relapse or last check-up. Disease-specific survival (DSS) was calculated from the surgical excision date of the primary tumor to the date of BC death or last check-up [24, 27]. Survival distribution curves were plotted using the Kaplan-Meier method and the statistical comparisons were performed using the log-rank test. Cox regression analyses were carried out on DFI and DSS to calculate crude and adjusted HRs and 95% confidence intervals (CIs) for the different study group. Cases lost to follow up and cases with a non-BC related cause of death were censored at the last follow up control. The step-wise model selection method was used to determine the final Cox regression model. Akaike Information Criterion test (AIC) and likelihood ratio test (LRT) were carried out to measure how selected variables improve parsimony and goodness of fit of the selected model. The proportional hazard assumption was assessed with the Schoenfeld residuals. This did not give reasons to suspect violation of this assumption. All statistical tests were two sided. P-values < 0.05 were considered significant. Statistical analyses were performed using Stata/SE12.0 Statistical Software (STATA, College Station, TX).
Results
Association of FOXA1 and AR IHC expression with clinical-pathological characteristics
Clinical and histopathological characteristics of the whole population are reported in Table 1. The median follow up was 10.1 years (7,7-12,7). The majority of patients was over 50 years (> 80%) of age and underwent conservative surgery. Positive expression of ER, AR and FOXA1 was observed in 78, 60 and 85% of cases respectively. As previously reported [13, 15], in our cohort FOXA1 positivity was associated with small tumor size (< 15 mm), absence of lymph node metastases, low histological grade, no special type (NST) histotype, low level of Ki67, as well as, with ER+ and PgR+ tumors (Additional file 1: Table S2). In the consecutive series of patients, 58% of cases showed AR+/FOXA1+ (Table 2), while 14% presented AR-/FOXA1- immunophenotype and only 1.7% of cases were AR+/FOXA1-. This latter subgroup, did not show specific features. Compared to the other subgroups, FOXA1-/AR- BC phenotype was more frequently associated with high histological grade, large tumor size, no expression of ER and PgR and high proliferation index (P < 0.001) (Additional file 1: Table S3).
qPCR analysis: Correlation between mRNA and protein levels of FOXA1 and AR in BC
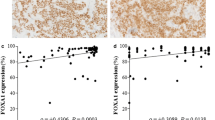
We found a strict correlation of FOXA1 and AR mRNA and protein expression (Fig. 2). To correlate the expression of ER, AR and FOXA1, we decided to use qPCR results, because this procedure allows quantifying more precisely the level of expression of each molecule. As shown in Fig. 3, there was a linear correlation (Spearman’s correlation) of the level of FOXA1 mRNA with the level of AR (r = 0.8975; P < 0.001) (Fig. 3a) and ER (r = 0.7326; P < 0.001) mRNA (Fig. 3b).
Protein (IHC) and mRNA (qPCR) expression for Androgen receptor (AR) and Forkhead box protein A1 (FOXA1). It can be observed that positive protein expression (AR and FOXA1) correlates with higher mRNA levels (low delta-Ct). Tukey’s multiple comparisons test showed significant differences between positive and negative cases, for both AR and FOXA1 (p < 0.0001). NT - Normal Tissue. *ANOVA analysis
Furthermore, FOXA1 mRNA was closely related to AR mRNA expression, regardless of ER status. Indeed, FOXA1 mRNA was expressed in all samples with ER+/AR+ (27 cases) and ER−/AR+ (3 cases) (Low delta-Ct. Fig. 4b and d), in 8/25 ER+/AR- cases and in only 1/10 ER−/AR- cases (High Delta-Ct. Fig. 4c and e).
FOXA1 mRNA expression in: a. NT - Normal Tissue; b. ER+/AR+ tumors; c. ER+/AR- tumors; d. ER−/AR+ tumors; e. ER−/AR- tumors. Independently of ER status, FOXA1 mRNA levels were higher (low delta-Ct) in AR+ tumors, compared to AR- cases. Tukey’s multiple comparisons test showed significant differences, mainly, between groups with AR+ and AR- cases (p < 0.0001. Additional file 1: Table S4). *ANOVA analysis
Impact of FOXA1 and AR IHC co-expression on prognosis
At univariate analysis performed on whole cohort, metastatic lymph nodes, histological grade, vascular invasion, ER and PR positivity, high Ki67 and HER2 overexpression were confirmed as significant prognostic factors. Additionally, the expression of AR and FOXA1 were associated with a better DFI and DSS (Table 3, Additional file 1: Figure S1).
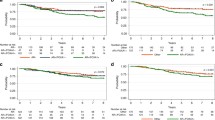
To analyze the impact of FOXA1 and AR in patients with BC (ER+ or ER-), we created three BC subgroups (FOXA1+/AR+; FOXA1+/AR-; FOXA1−/AR-). We were unable to perform any analyses on the FOXA1−/AR+ BC since only 8 patients carried this phenotype (Table 2). As shown in Fig. 5, in the consecutive series of patients, the lack of expression of both, FOXA1 and AR (FOXA1−/AR-), was related to a worse DFI and DSS compared to the other groups.
Finally, we investigated the relationship between FOXA1, AR and prognosis in BC patients stratified for ER expression. As shown in Fig. 6, in ER+ BC, FOXA1 expression was closely related to good prognosis independently of AR expression.
Multivariate analyses (Table 4) performed on ER+ BC confirmed that FOXA1 may provide more information than AR on DFI, but not on DSS. In the subset of patients with ER- BC, FOXA1, alone or in association with AR, did not show any relationship with outcome (data not shown).
Discussion
We assessed, for the first time, the expression of FOXA1 and AR in BC, evaluating their prognostic impact according to ER status. We found that (i) the expression (protein and mRNA) of FOXA1 and AR was closely related: the majority of cases expressing AR showed FOXA1 positivity, conversely, negative expression of FOXA1 correlates with very low level of AR; (ii) the expression of FOXA1 is strictly related to good outcome, and in the subgroup of patients with ER+ BC may provide more information on DFI than AR.
FOXA1 is a “winged helix” transcription factor. It was demonstrated that, by interacting with histones H3 and H4, FOXA1 is responsible for opening compacted chromatin [28], permitting efficient interaction of ER with its response elements. For this reason, the presence of FOXA1 suggests a functional ER complex, which probably will respond to endocrine therapy [11, 29, 30]. Moreover, FOXA1 seems to have a repressor effect on BC growth by promoting transcription of E-cadherin and cell cycle-dependent kinase inhibitor p27(Kip1), thus reducing the motility and invasion of BC cells [31, 32]. These findings suggest that FOXA1 expression in BC may be associated with a better clinical outcome. In our study we confirmed literature data, demonstrating that FOXA1 is mainly expressed in low grade, lymph node negative BC tumors, with size < 15 mm and low Ki67 index [15, 33, 34].
In addition, FOXA1 has been associated with recruitment of AR [7] and, it has been suggested that in prostate epithelium FOXA1 acts with AR in promoting differentiation [35].
ChIP-seq analysis of AR, ER, and FOXA1 in BC cell lines revealed a significant level of co-occupancy between these markers, presumably due to the presence of forkhead motif found at AR and ER binding sites [8, 10, 36, 37]. Furthermore, evidences of the relationship between AR and FOXA1 was supported by experiments demonstrating the co-localization of the two proteins on chromatin [1, 9, 37]. Our results support the evidence of those studies, showing that BC tumor with high mRNA level of FOXA1 are generally ER and AR enriched. On the contrary, tissues with low FOXA1 mRNA level present low level of hormonal receptors, especially of AR.
In several studies has been demonstrated that AR expression is a favorable prognostic marker of disease outcome in ER+ BC [5, 6]. This result has recently been confirmed in a meta-analysis conducted on 17,000 women with early-stage breast cancer [38]. The present work confirms the prognostic role of AR. However, the concurrent evaluation of the expression of both AR and FOXA1, shows that FOXA1 is superior to AR as prognostic marker in patients with BC, especially in ER+ cases. In fact, FOXA1 expression was always related to a better outcome even if AR was not detectable. Similar results were recently obtained in prostate cancer [39], in which it has been demonstrated that FOXA1 expression is closely related to prognosis independently of AR level. Hence, in FOXA1+ BC patients, similar results regarding prognosis were found in AR- and AR+ cases. Thus, we suppose that in ER+ BC patients, FOXA1 could be more important than AR as a marker of better prognosis. Actually, several studies suggested that functionality of AR as well as ER may depend on FOXA1 activity [1, 8, 11].
Sahu, B et al. suggested that in prostate cancers FOXA1 level may contribute to select specific AR binding sites on DNA, activating different gene expression signatures [39]. In our case series we observed very low number of AR+/FOXA1- cases; moreover, as shown in Fig. 4, the expression of these markers seems to correlate. Therefore, we hypothesize that FOXA1 in ER+ BC may control the level of AR expression.
Conclusions
Our results suggest that in BC the expression of FOXA1 is directly proportional to the expression of AR. Despite that, FOXA1 is found as a superior predicting marker of recurrences compared to AR in ER+ BC patients. Therefore, FOXA1 expression evaluated by IHC on ER+ BC specimens could be considered in routine diagnosis as an additional support to oncologists in the definition of the patient prognosis.
Abbreviations
- AIC:
-
Akaike information criterion
- AR:
-
Androgen receptor
- DFI:
-
Disease-free interval
- DSS:
-
Disease-specific survival
- ER:
-
Estrogen receptor
- FOXA1:
-
Forkhead-box A1 protein
- HER2:
-
Human epidermal growth factor receptor 2
- PgR:
-
Progesterone receptor
- qPCR:
-
Real-time PCR
References
Robinson JL, Macarthur S, Ross-Innes CS, Tilley WD, Neal DE, Mills IG, Carroll JS. Androgen receptor driven transcription in molecular apocrine breast cancer is mediated by FoxA1. EMBO J. 2011;30(15):3019–27.
Lehmann BD, Bauer JA, Chen X, Sanders ME, Chakravarthy AB, Shyr Y, Pietenpol JA. Identification of human triple-negative breast cancer subtypes and preclinical models for selection of targeted therapies. J Clin Invest. 2011;121(7):2750–67.
Cops EJ, Bianco-Miotto T, Moore NL, Clarke CL, Birrell SN, Butler LM, Tilley WD. Antiproliferative actions of the synthetic androgen, mibolerone, in breast cancer cells are mediated by both androgen and progesterone receptors. J Steroid Biochem Mol Biol. 2008;110(3–5):236–43.
Ortmann J, Prifti S, Bohlmann MK, Rehberger-Schneider S, Strowitzki T, Rabe T. Testosterone and 5 alpha-dihydrotestosterone inhibit in vitro growth of human breast cancer cell lines. Gynecol Endocrinol. 2002;16(2):113–20.
Castellano I, Allia E, Accortanzo V, Vandone AM, Chiusa L, Arisio R, Durando A, Donadio M, Bussolati G, Coates AS, et al. Androgen receptor expression is a significant prognostic factor in estrogen receptor positive breast cancers. Breast Cancer Res Treat. 2010;124(3):607–17.
Castellano I, Chiusa L, Vandone AM, Beatrice S, Goia M, Donadio M, Arisio R, Muscara F, Durando A, Viale G, et al. A simple and reproducible prognostic index in luminal ER-positive breast cancers. Ann Oncol. 2013;24(9):2292–7.
Augello MA, Hickey TE, Knudsen KE. FOXA1: master of steroid receptor function in cancer. EMBO J. 2011;30(19):3885–94.
Hurtado A, Holmes KA, Ross-Innes CS, Schmidt D, Carroll JS. FOXA1 is a key determinant of estrogen receptor function and endocrine response. Nat Genet. 2011;43(1):27–33.
Ni M, Chen Y, Lim E, Wimberly H, Bailey ST, Imai Y, Rimm DL, Liu XS, Brown M. Targeting androgen receptor in estrogen receptor-negative breast cancer. Cancer Cell. 2011;20(1):119–31.
Carroll JS, Liu XS, Brodsky AS, Li W, Meyer CA, Szary AJ, Eeckhoute J, Shao W, Hestermann EV, Geistlinger TR, et al. Chromosome-wide mapping of estrogen receptor binding reveals long-range regulation requiring the forkhead protein FoxA1. Cell. 2005;122(1):33–43.
Badve S, Turbin D, Thorat MA, Morimiya A, Nielsen TO, Perou CM, Dunn S, Huntsman DG, Nakshatri H. FOXA1 expression in breast cancer--correlation with luminal subtype A and survival. Clin Cancer Res. 2007;13(15 Pt 1):4415–21.
Habashy HO, Powe DG, Rakha EA, Ball G, Paish C, Gee J, Nicholson RI, Ellis IO. Forkhead-box A1 (FOXA1) expression in breast cancer and its prognostic significance. Eur J Cancer. 2008;44(11):1541–51.
He K, Zeng H, Xu X, Li A, Cai Q, Long X. Clinicopathological significance of forkhead box protein A1 in breast cancer: a meta-analysis. Exp Ther Med. 2016;11(6):2525–30.
Horimoto Y, Arakawa A, Harada-Shoji N, Sonoue H, Yoshida Y, Himuro T, Igari F, Tokuda E, Mamat O, Tanabe M, et al. Low FOXA1 expression predicts good response to neo-adjuvant chemotherapy resulting in good outcomes for luminal HER2-negative breast cancer cases. Br J Cancer. 2015;112(2):345–51.
Mehta RJ, Jain RK, Leung S, Choo J, Nielsen T, Huntsman D, Nakshatri H, Badve S. FOXA1 is an independent prognostic marker for ER-positive breast cancer. Breast Cancer Res Treat. 2012;131(3):881–90.
Ross-Innes CS, Stark R, Teschendorff AE, Holmes KA, Ali HR, Dunning MJ, Brown GD, Gojis O, Ellis IO, Green AR, et al. Differential oestrogen receptor binding is associated with clinical outcome in breast cancer. Nature. 2012;481(7381):389–93.
Shou J, Lai Y, Xu J, Huang J. Prognostic value of FOXA1 in breast cancer: a systematic review and meta-analysis. Breast. 2016;27:35–43.
Thorat MA, Marchio C, Morimiya A, Savage K, Nakshatri H, Reis-Filho JS, Badve S. Forkhead box A1 expression in breast cancer is associated with luminal subtype and good prognosis. J Clin Pathol. 2008;61(3):327–32.
Wolf I, Bose S, Williamson EA, Miller CW, Karlan BY, Koeffler HP. FOXA1: growth inhibitor and a favorable prognostic factor in human breast cancer. Int J Cancer. 2007;120(5):1013–22.
Robinson JL, Holmes KA, Carroll JS. FOXA1 mutations in hormone-dependent cancers. Front Oncol. 2013;3:20.
Sapino A, Marchio C, Senetta R, Castellano I, Macri L, Cassoni P, Ghisolfi G, Cerrato M, D’Ambrosio E, Bussolati G. Routine assessment of prognostic factors in breast cancer using a multicore tissue microarray procedure. Virchows Arch. 2006;449(3):288–96.
Goldhirsch A, Wood WC, Coates AS, Gelber RD, Thurlimann B, Senn HJ, Panel members. Strategies for subtypes--dealing with the diversity of breast cancer: highlights of the St. Gallen International Expert Consensus on the Primary Therapy of Early Breast Cancer 2011. Ann Oncol. 2011;22(8):1736–47.
Goldhirsch A, Winer EP, Coates AS, Gelber RD, Piccart-Gebhart M, Thurlimann B, Senn HJ, Panel members. Personalizing the treatment of women with early breast cancer: highlights of the St Gallen International Expert Consensus on the Primary Therapy of Early Breast Cancer 2013. Ann Oncol. 2013;24(9):2206–23.
Bustreo S, Osella-Abate S, Cassoni P, Donadio M, Airoldi M, Pedani F, Papotti M, Sapino A, Castellano I. Optimal Ki67 cut-off for luminal breast cancer prognostic evaluation: a large case series study with a long-term follow-up. Breast Cancer Res Treat. 2016;157(2):363–71.
Coates AS, Winer EP, Goldhirsch A, Gelber RD, Gnant M, Piccart-Gebhart M, Thurlimann B, Senn HJ, Panel M. Tailoring therapies--improving the management of early breast cancer: St Gallen International Expert Consensus on the Primary Therapy of Early Breast Cancer 2015. Ann Oncol. 2015;26(8):1533–46.
Wolff AC, Hammond ME, Hicks DG, Dowsett M, McShane LM, Allison KH, Allred DC, Bartlett JM, Bilous M, Fitzgibbons P, et al. Recommendations for human epidermal growth factor receptor 2 testing in breast cancer: American Society of Clinical Oncology/College of American Pathologists clinical practice guideline update. J Clin Oncol. 2013;31(31):3997–4013.
Rangel N, Rondon-Lagos M, Annaratone L, Osella-Abate S, Metovic J, Mano MP, Bertero L, Cassoni P, Sapino A, Castellano I. The role of the AR/ER ratio in ER-positive breast cancer patients. Endocr Relat Cancer. 2018;25(3):163–72.
Cirillo LA, Lin FR, Cuesta I, Friedman D, Jarnik M, Zaret KS. Opening of compacted chromatin by early developmental transcription factors HNF3 (FoxA) and GATA-4. Mol Cell. 2002;9(2):279–89.
Hisamatsu Y, Tokunaga E, Yamashita N, Akiyoshi S, Okada S, Nakashima Y, Aishima S, Morita M, Kakeji Y, Maehara Y. Impact of FOXA1 expression on the prognosis of patients with hormone receptor-positive breast cancer. Ann Surg Oncol. 2012;19(4):1145–52.
Hisamatsu Y, Tokunaga E, Yamashita N, Akiyoshi S, Okada S, Nakashima Y, Taketani K, Aishima S, Oda Y, Morita M, et al. Impact of GATA-3 and FOXA1 expression in patients with hormone receptor-positive/HER2-negative breast cancer. Breast Cancer. 2015;22(5):520–8.
Liu YN, Lee WW, Wang CY, Chao TH, Chen Y, Chen JH. Regulatory mechanisms controlling human E-cadherin gene expression. Oncogene. 2005;24(56):8277–90.
Williamson EA, Wolf I, O’Kelly J, Bose S, Tanosaki S, Koeffler HP. BRCA1 and FOXA1 proteins coregulate the expression of the cell cycle-dependent kinase inhibitor p27(Kip1). Oncogene. 2006;25(9):1391–9.
Ademuyiwa FO, Thorat MA, Jain RK, Nakshatri H, Badve S. Expression of Forkhead-box protein A1, a marker of luminal A type breast cancer, parallels low Oncotype DX 21-gene recurrence scores. Mod Pathol. 2010;23(2):270–5.
Oh DS, Troester MA, Usary J, Hu Z, He X, Fan C, Wu J, Carey LA, Perou CM. Estrogen-regulated genes predict survival in hormone receptor-positive breast cancers. J Clin Oncol. 2006;24(11):1656–64.
Gao N, Ishii K, Mirosevich J, Kuwajima S, Oppenheimer SR, Roberts RL, Jiang M, Yu X, Shappell SB, Caprioli RM, et al. Forkhead box A1 regulates prostate ductal morphogenesis and promotes epithelial cell maturation. Development. 2005;132(15):3431–43.
Jia L, Berman BP, Jariwala U, Yan X, Cogan JP, Walters A, Chen T, Buchanan G, Frenkel B, Coetzee GA. Genomic androgen receptor-occupied regions with different functions, defined by histone acetylation, coregulators and transcriptional capacity. PLoS One. 2008;3(11):e3645.
Wang Q, Li W, Liu XS, Carroll JS, Janne OA, Keeton EK, Chinnaiyan AM, Pienta KJ, Brown M. A hierarchical network of transcription factors governs androgen receptor-dependent prostate cancer growth. Mol Cell. 2007;27(3):380–92.
Bozovic-Spasojevic I, Zardavas D, Brohee S, Ameye L, Fumagalli D, Ades F, de Azambuja E, Bareche Y, Piccart M, Paesmans M, et al. The prognostic role of androgen receptor in patients with early-stage breast cancer: a meta-analysis of clinical and gene expression data. Clin Cancer Res. 2017;23(11):2702–12.
Sahu B, Laakso M, Ovaska K, Mirtti T, Lundin J, Rannikko A, Sankila A, Turunen JP, Lundin M, Konsti J, et al. Dual role of FoxA1 in androgen receptor binding to chromatin, androgen signalling and prostate cancer. EMBO J. 2011;30(19):3962–76.
Acknowledgments
The authors would like to acknowledge technical support in immunohistochemical procedures to Mrs. Stefania Bolla and Mr. Marco Cupo.
Funding
This study was funded by “Lega Italiana per la Lotta contro i Tumori” – LILT and by the Ministry of University bando ricerca locale ex-60% anno 2014 to IC. NR was supported by Colciencias Grant (call 617, Colombia). The funding body had no role in the design of the study and collection, analysis, and interpretation of data and in writing the manuscript.
Availability of data and materials
All data generated or analysed during this study are included in this published article [and its supplementary information files].
Author information
Authors and Affiliations
Contributions
Study conception and design (Conceptualization-Methodology) NR, NF, SOA, LA, AS and IC. Acquisition of data (Investigation) NR, NF, LA, CI, MGC, LR, JN, RB, DB, PF, FM, PC and IC. Analysis and interpretation of data (Data curation-Formal analysis) NR, NF, SOA, LA, MGC, LR, RB, DB, PF, FM, PC, AS and IC. Drafting of manuscript (Writing – original draft preparation) NF, CI, MGC, LR, JM, RB, DB, PF, FM and IC. Critical revision (Writing – review and editing) NR, SOA, LA, JM, PC, AS and IC. Final approval of the version to be submitted NR, NF, SOA, LA, CI, MGC, LR, JM, RB, DB, PF, FM, PC, AS and IC. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Ethics approval and consent to participate
The project provided an informed consent, obtained from the patients at the time of surgery due to the retrospective approach of the study, which did not impact on their treatment. The procedure for collecting the consent was approved by the Committee for human Biospecimen Utilization (Department of Medical Sciences – ChBU). “All the cases were anonymously recorded, and data were accessed anonymously”. The study was conducted in compliance with the Helsinki Declaration.
Consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing interests.
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Additional file
Additional file 1:
Table S1. Primers for real-time PCR. Table S2. Patients’ clinical and histopathological characteristics according to FOXA1 expression. Table S3. Clinical and histopathological characteristics of BC patients according to FOXA1 and AR status. Table S4. Multiple comparisons of FOXA1 mRNA expression in tumors classified according to ER and AR status. Figure S1. Kaplan–Meier estimates of a) disease free interval and b) disease-specific survival according to FOXA1 status in 479 breast tumors. (DOCX 662 kb)
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The Creative Commons Public Domain Dedication waiver (http://creativecommons.org/publicdomain/zero/1.0/) applies to the data made available in this article, unless otherwise stated.
About this article
Cite this article
Rangel, N., Fortunati, N., Osella-Abate, S. et al. FOXA1 and AR in invasive breast cancer: new findings on their co-expression and impact on prognosis in ER-positive patients. BMC Cancer 18, 703 (2018). https://doi.org/10.1186/s12885-018-4624-y
Received:
Accepted:
Published:
DOI: https://doi.org/10.1186/s12885-018-4624-y