Abstract
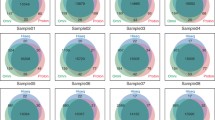
Although many quality control (QC) methods have been developed to improve the quality of single-nucleotide variants (SNVs) in SNV-calling, QC methods for use subsequent to single-nucleotide polymorphism-calling have not been reported. We developed five QC metrics to improve the quality of SNVs using the whole-genome-sequencing data of a monozygotic twin pair from the Korean Personal Genome Project. The QC metrics improved both repeatability between the monozygotic twin pair and reproducibility between SNV-calling pipelines. We demonstrated the QC metrics improve reproducibility of SNVs derived from not only whole-genome-sequencing data but also whole-exome-sequencing data. The QC metrics are calculated based on the reference genome used in the alignment without accessing the raw and intermediate data or knowing the SNV-calling details. Therefore, the QC metrics can be easily adopted in downstream association analysis.








Similar content being viewed by others
References
Cichon S, Craddock N, Daly M, Faraone SV, Gejman PV, Kelsoe J et al. Genomewide association studies: history, rationale, and prospects for psychiatric disorders. Am J Psychiatry 2009; 166: 540–556.
Marian AJ . Molecular genetic studies of complex phenotypes. Transl Res 2012; 159: 64–79.
Hong H, Jawaid A, Wang J, Catalano J, Fox JC, Hawkins TB . Combining genetic variations in CYP2C9 and VKORC1 with clinical factors for warfarin dosing determination improved clinical effectiveness. Pharmacogenomics 2013; 14: 459–460.
Klein RJ, Zeiss C, Chew EY, Tsai JY, Sackler RS, Haynes C et al. Complement factor H polymorphism in age-related macular degeneration. Science 2005; 308: 385–389.
Scott LJ, Mohlke KL, Bonnycastle LL, Willer CJ, Li Y, Duren WL et al. A genome-wide association study of type 2 diabetes in Finns detects multiple susceptibility variants. Science 2007; 316: 1341–1345.
Sladek R, Rocheleau G, Rung J, Dina C, Shen L, Serre D et al. A genome-wide association study identifies novel risk loci for type 2 diabetes. Nature 2007; 445: 881–885.
Wellcome Trust Case Control Consortium. Genome-wide association study of 14,000 cases of seven common diseases and 3,000 shared controls. Nature 2007; 447: 661–678.
Zeggini E, Weedon MN, Lindgren CM, Frayling TM, Elliott KS, Lango H et al. Replication of genome-wide association signals in UK samples reveals risk loci for type 2 diabetes. Science 2007; 316: 1336–1341.
Gold B, Kirchhoff T, Stefanov S, Lautenberger J, Viale A, Garber J et al. Genome-wide association study provides evidence for a breast cancer risk locus at 6q22.33. Proc Natl Acad Sci USA 2008; 105: 4340–4345.
Hindorff LA, Sethupathy P, Junkins HA, Ramos EM, Mehta JP, Collins FS et al. Potential etiologic and functional implications of genome-wide association loci for human diseases and traits. Proc Natl Acad Sci USA 2009; 106: 9362–9367.
Petersen GM, Amundadottir L, Fuchs CS, Kraft P, Stolzenberg-Solomon RZ, Jacobs KB et al. A genome-wide association study identifies pancreatic cancer susceptibility loci on chromosomes 13q22.1, 1q32.1 and 5p15.33. Nat Genet 2010; 42: 224–228.
Hong H, Xu L, Mendrick D, Tong W . Genome-Wide Association Studies of Type 2 Diabetes: Current Status, Open Challenges, and Future Perspectives. In: Barh D, Blum K, Madigan MA (eds). OMICS: Biomedical Perspectives and Applications. CRC Press Taylor & Francis Group, Boca Raton, Florida, USA, 2011, pp 401–430.
Rung J, Cauchi S, Albrechtsen A, Shen L, Rocheleau G, Cavalcanti-Proenca C et al. Genetic variant near IRS1 is associated with type 2 diabetes, insulin resistance and hyperinsulinemia. Nat Genet 2009; 41: 1110–1115.
Steinthorsdottir V, Thorleifsson G, Reynisdottir I, Benediktsson R, Jonsdottir T, Walters GB et al. A variant in CDKAL1 influences insulin response and risk of type 2 diabetes. Nat Genet 2007; 39: 770–775.
Hirschhorn JN . Genomewide association studies—illuminating biologic pathways. N Engl J Med 2009; 360: 1699–1701.
Kraft P, Hunter DJ . Genetic risk prediction—are we there yet? N Engl J Med 2009; 360: 1701–1703.
Hong H, Xu L, Su Z, Liu J, Ge W, Shen J et al. Pitfall of genome-wide association studies: Sources of inconsistency in genotypes and their effects. J Biomed Sci Eng 2012; 5: 557–573.
Pearson TA, Manolio TA . How to interpret a genome-wide association study. JAMA 2008; 299: 1335–1344.
Frayling TM, Timpson NJ, Weedon MN, Zeggini E, Freathy RM, Lindgren CM et al. A common variant in the FTO gene is associated with body mass index and predisposes to childhood and adult obesity. Science 2007; 316: 889–894.
McCarthy MI, Abecasis GR, Cardon LR, Goldstein DB, Little J, Ioannidis JP et al. Genome-wide association studies for complex traits: consensus, uncertainty and challenges. Nat Rev Genet 2008; 9: 356–369.
Hong H, Shi L, Su Z, Ge W, Jones WD, Czika W et al. Assessing sources of inconsistencies in genotypes and their effects on genome-wide association studies with HapMap samples. Pharmacogenomics J 2010; 10: 364–374.
Hong H, Su Z, Ge W, Shi L, Perkins R, Fang H et al. Evaluating variations of genotype calling: a potential source of spurious associations in genome-wide association studies. J Genet 2010; 89: 55–64.
Hong H, Su Z, Ge W, Shi L, Perkins R, Fang H et al. Assessing batch effects of genotype calling algorithm BRLMM for the Affymetrix GeneChip Human Mapping 500 K array set using 270 HapMap samples. BMC Bioinformatics 2008; 9: S17.
Hong H, Xu L, Liu J, Jones WD, Su Z, Ning B et al. Technical reproducibility of genotyping SNP arrays used in genome-wide association studies. PLoS One 2012; 7: e44483.
Hoheisel JD . Microarray technology: beyond transcript profiling and genotype analysis. Nat Rev Genet 2006; 7: 200–210.
Bentley DR, Balasubramanian S, Swerdlow HP, Smith GP, Milton J, Brown CG et al. Accurate whole human genome sequencing using reversible terminator chemistry. Nature 2008; 456: 53–59.
Drmanac R, Sparks AB, Callow MJ, Halpern AL, Burns NL, Kermani BG et al. Human genome sequencing using unchained base reads on self-assembling DNA nanoarrays. Science 2010; 327: 78–81.
Wheeler DA, Srinivasan M, Egholm M, Shen Y, Chen L, McGuire A et al. The complete genome of an individual by massively parallel DNA sequencing. Nature 2008; 452: 872–876.
Kim JI, Ju YS, Park H, Kim S, Lee S, Yi JH et al. A highly annotated whole-genome sequence of a Korean individual. Nature 2009; 460: 1011–1015.
Chung S, Low SK, Zembutsu H, Takahashi A, Kubo M, Sasa M et al. A genome-wide association study of chemotherapy-induced alopecia in breast cancer patients. Breast Cancer Res 2013; 15: R81.
Gudmundsson J, Sulem P, Gudbjartsson DF, Masson G, Agnarsson BA, Benediktsdottir KR et al. A study based on whole-genome sequencing yields a rare variant at 8q24 associated with prostate cancer. Nat Genet 2012; 44: 1326–1329.
Jonsson T, Atwal JK, Steinberg S, Snaedal J, Jonsson PV, Bjornsson S et al. A mutation in APP protects against Alzheimer's disease and age-related cognitive decline. Nature 2012; 488: 96–99.
Hong H, Zhang W, Shen J, Su Z, Ning B, Han T et al. Critical role of bioinformatics in translating huge amounts of next-generation sequencing data into personalized medicine. Sci China Life Sci 2013; 56: 110–118.
Parkinson NJ, Maslau S, Ferneyhough B, Zhang G, Gregory L, Buck D et al. Preparation of high-quality next-generation sequencing libraries from picogram quantities of target DNA. Genome Res 2012; 22: 125–133.
Thaitrong N, Kim H, Renzi RF, Bartsch MS, Meagher RJ, Patel KD . Quality control of next-generation sequencing library through an integrative digital microfluidic platform. Electrophoresis 2012; 33: 3506–3513.
Cabanski CR, Cavin K, Bizon C, Wilkerson MD, Parker JS, Wilhelmsen KC et al. ReQON: a Bioconductor package for recalibrating quality scores from next-generation sequencing data. BMC Bioinformatics 2012; 13: 221.
Patel RK, Jain M . NGS QC Toolkit: a toolkit for quality control of next generation sequencing data. PLoS One 2012; 7: e30619.
Li H . Improving SNP discovery by base alignment quality. Bioinformatics 2011; 27: 1157–1158.
Reumers J, De Rijk P, Zhao H, Liekens A, Smeets D, Cleary J et al. Optimized filtering reduces the error rate in detecting genomic variants by short-read sequencing. Nat Biotechnol 2012; 30: 61–68.
Forster M, Forster P, Elsharawy A, Hemmrich G, Kreck B, Wittig M et al. From next-generation sequencing alignments to accurate comparison and validation of single-nucleotide variants: the pibase software. Nucleic Acids Res 2013; 41: e16.
Ratan A, Miller W, Guillory J, Stinson J, Seshagiri S, Schuster SC . Comparison of sequencing platforms for single nucleotide variant calls in a human sample. PLoS One 2013; 8: e55089.
O'Rawe J, Jiang T, Sun G, Wu Y, Wang W, Hu J et al. Low concordance of multiple variant-calling pipelines: practical implications for exome and genome sequencing. Genome Med 2013; 5: 28.
Ahn SM, Kim TH, Lee S, Kim D, Ghang H, Kim DS et al. The first Korean genome sequence and analysis: full genome sequencing for a socio-ethnic group. Genome Res 2009; 19: 1622–1629.
Li H, Durbin R . Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics 2009; 25: 1754–1760.
Li H, Handsaker B, Wysoker A, Fennell T, Ruan J, Homer N et al. The Sequence Alignment/Map format and SAMtools. Bioinformatics 2009; 25: 2078–2079.
Li R, Yu C, Li Y, Lam TW, Yiu SM, Kristiansen K et al. SOAP2: an improved ultrafast tool for short read alignment. Bioinformatics 2009; 25: 1966–1967.
Li R, Li Y, Fang X, Yang H, Wang J, Kristiansen K . SNP detection for massively parallel whole-genome resequencing. Genome Res 2009; 19: 1124–1132.
McKenna A, Hanna M, Banks E, Sivachenko A, Cibulskis K, Kernytsky A et al. The Genome Analysis Toolkit: a MapReduce framework for analyzing next-generation DNA sequencing data. Genome Res 2010; 20: 1297–1303.
Wei Z, Wang W, Hu P, Lyon GJ, Hakonarson H . SNVer: a statistical tool for variant calling in analysis of pooled or individual next-generation sequencing data. Nucleic Acids Res 2011; 39: e132.
Clement NL, Snell Q, Clement MJ, Hollenhorst PC, Purwar J, Graves BJ et al. The GNUMAP algorithm: unbiased probabilistic mapping of oligonucleotides from next-generation sequencing. Bioinformatics 2010; 26: 38–45.
Patwari P, Lee RT . Mechanical control of tissue morphogenesis. Circ Res 2008; 103: 234–243.
Roberts NJ, Vogelstein JT, Parmigiani G, Kinzler KW, Vogelstein B, Velculescu VE . The predictive capacity of personal genome sequencing. Sci Transl Med 2012; 4: 133ra158.
Acknowledgements
This research was supported in part by an appointment to the Research Participation Program at the National Center for Toxicological Research (WZ and HWN) and at the Center for Biologics Evaluation and Research (VS) administered by the Oak Ridge Institute for Science and Education through an interagency agreement between the US Department of Energy and the US Food and Drug Administration (FDA). We thank Mike Mikailov of FDA’s Center for Devices and Radiological Health for his technical support on the data analysis conducted on the high-performance computation cluster Betsy in the White Oak Data Center of FDA. We acknowledge Critical Assessment of Massive Data Analysis consortium for providing us the raw data, as well as the mapping and SNVs calling results from KPGP. The findings and conclusions in this article have not been formally disseminated by the US FDA and should not be construed to represent any agency determination or policy.
Author Contributions
HH, WZ, WT and RP conceived the study and designed the experiments. WZ, VS (Valerii Soika), JM, ZS, WG, HWN and HH performed the data analysis. HH, WZ, RP and WT wrote the paper with assistance from the other authors. All authors have read and approved the manuscript for publication.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies the paper on the The Pharmacogenomics Journal website
Supplementary information
Rights and permissions
About this article
Cite this article
Zhang, W., Soika, V., Meehan, J. et al. Quality control metrics improve repeatability and reproducibility of single-nucleotide variants derived from whole-genome sequencing. Pharmacogenomics J 15, 298–309 (2015). https://doi.org/10.1038/tpj.2014.70
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/tpj.2014.70
- Springer Nature Limited