Abstract
The evolutionary origin of vertebrates included innovations in sensory processing associated with the acquisition of a predatory lifestyle1. Vertebrates perceive external stimuli through sensory systems serviced by cranial sensory ganglia, whose neurons arise predominantly from cranial placodes; however, the understanding of the evolutionary origin of placodes and cranial sensory ganglia is hampered by the anatomical differences between living lineages and the difficulty in assigning homology between cell types and structures. Here we show that the homeobox transcription factor Hmx is a constitutive component of vertebrate sensory ganglion development and that in the tunicate Ciona intestinalis, Hmx is necessary and sufficient to drive the differentiation programme of bipolar tail neurons, cells previously thought to be homologues of neural crest2,3. Using Ciona and lamprey transgenesis, we demonstrate that a unique, tandemly duplicated enhancer pair regulated Hmx expression in the stem-vertebrate lineage. We also show notably robust vertebrate Hmx enhancer function in Ciona, demonstrating that deep conservation of the upstream regulatory network spans the evolutionary origin of vertebrates. These experiments demonstrate regulatory and functional conservation between Ciona and vertebrate Hmx, and point to bipolar tail neurons as homologues of cranial sensory ganglia.




Similar content being viewed by others
Data availability
Cloned Hmx gene sequences have been deposited in Genbank accessions MN264670–MN264672. RNA-seq data have been deposited in SRA accession GSE141046. Original data underlying Fig. 4b, c of this manuscript can be accessed from the Stowers Original Data Repository at http://odr.stowers.org/websimr/.
References
Northcutt, R. G. & Gans, C. The genesis of neural crest and epidermal placodes: a reinterpretation of vertebrate origins. Q. Rev. Biol. 58, 1–28 (1983).
Horie, R. et al. Shared evolutionary origin of vertebrate neural crest and cranial placodes. Nature 560, 228–232 (2018).
Stolfi, A., Ryan, K., Meinertzhagen, I. A. & Christiaen, L. Migratory neuronal progenitors arise from the neural plate borders in tunicates. Nature 527, 371–374 (2015).
Shimeld, S. M. & Holland, P. W. Vertebrate innovations. Proc. Natl Acad. Sci. USA 97, 4449–4452 (2000).
Patthey, C. et al. Identification of molecular signatures specific for distinct cranial sensory ganglia in the developing chick. Neural Dev. 11, 3 (2016).
Adamska, M. et al. Five Nkx5 genes show differential expression patterns in anlagen of sensory organs in medaka: insight into the evolution of the gene family. Dev. Genes Evol. 211, 338–349 (2001).
Wang, W., Lo, P., Frasch, M. & Lufkin, T. Hmx: an evolutionary conserved homeobox gene family expressed in the developing nervous system in mice and Drosophila. Mech. Dev. 99, 123–137 (2000).
Feng, Y. & Xu, Q. Pivotal role of hmx2 and hmx3 in zebrafish inner ear and lateral line development. Dev. Biol. 339, 507–518 (2010).
Kelly, L. E. & El-Hodiri, H. M. Xenopus laevis Nkx5.3 and sensory organ homeobox (SOHo) are expressed in developing sensory organs and ganglia of the head and anterior trunk. Dev. Genes Evol. 226, 423–428 (2016).
Kiernan, A. E., Nunes, F., Wu, D. K. & Fekete, D. M. The expression domain of two related homeobox genes defines a compartment in the chicken inner ear that may be involved in semicircular canal formation. Dev. Biol. 191, 215–229 (1997).
Quina, L. A., Tempest, L., Hsu, Y. W., Cox, T. C. & Turner, E. E. Hmx1 is required for the normal development of somatosensory neurons in the geniculate ganglion. Dev. Biol. 365, 152–163 (2012).
Takahashi, H., Shintani, T., Sakuta, H. & Noda, M. CBF1 controls the retinotectal topographical map along the anteroposterior axis through multiple mechanisms. Development 130, 5203–5215 (2003).
Bayramov, A. V., Martynova, N. Y., Eroshkin, F. M., Ermakova, G. V. & Zaraisky, A. G. The homeodomain-containing transcription factor X-nkx-5.1 inhibits expression of the homeobox gene Xanf-1 during the Xenopus laevis forebrain development. Mech. Dev. 121, 1425–1441 (2004).
Takatori, N. et al. Comprehensive survey and classification of homeobox genes in the genome of amphioxus, Branchiostoma floridae. Dev. Genes Evol. 218, 579–590 (2008).
Wada, S. et al. A genomewide survey of developmentally relevant genes in Ciona intestinalis. II. Genes for homeobox transcription factors. Dev. Genes Evol. 213, 222–234 (2003).
Ryan, K., Lu, Z. & Meinertzhagen, I. A. The CNS connectome of a tadpole larva of Ciona intestinalis (L.) highlights sidedness in the brain of a chordate sibling. eLife 5, e16962 (2016).
Wang, W., Chan, E. K., Baron, S., Van de Water, T. & Lufkin, T. Hmx2 homeobox gene control of murine vestibular morphogenesis. Development 128, 5017–5029 (2001).
Wang, W., Grimmer, J. F., Van De Water, T. R. & Lufkin, T. Hmx2 and Hmx3 homeobox genes direct development of the murine inner ear and hypothalamus and can be functionally replaced by Drosophila Hmx. Dev. Cell 7, 439–453 (2004).
Wang, W., Van De Water, T. & Lufkin, T. Inner ear and maternal reproductive defects in mice lacking the Hmx3 homeobox gene. Development 125, 621–634 (1998).
Tang, W. J., Chen, J. S. & Zeller, R. W. Transcriptional regulation of the peripheral nervous system in Ciona intestinalis. Dev. Biol. 378, 183–193 (2013).
Sharma, S., Wang, W. & Stolfi, A. Single-cell transcriptome profiling of the Ciona larval brain. Dev. Biol. 448, 226–236 (2019).
Kim, K. et al. Regulation of neurogenesis by FGF signaling and neurogenin in the invertebrate chordate ciona. Front. Cell Dev. Biol. 8, 477 (2020).
Chacha, P. P. et al. Neuronal identities derived by misexpression of the POU IV sensory determinant in a protovertebrate. Proc. Natl Acad. Sci. USA 119, e2118817119 (2022).
Brozovic, M. et al. ANISEED 2017: extending the integrated ascidian database to the exploration and evolutionary comparison of genome-scale datasets. Nucleic Acids Res. 46, D718–D725 (2018).
Doglio, L. et al. Parallel evolution of chordate cis-regulatory code for development. PLoS Genet. 9, e1003904 (2013).
McEwen, G. K. et al. Early evolution of conserved regulatory sequences associated with development in vertebrates. PLoS Genet. 5, e1000762 (2009).
Shimeld, S. M. & Donoghue, P. C. Evolutionary crossroads in developmental biology: cyclostomes (lamprey and hagfish). Development 139, 2091–2099 (2012).
Parker, H. J., Bronner, M. E. & Krumlauf, R. A Hox regulatory network of hindbrain segmentation is conserved to the base of vertebrates. Nature 514, 490–493 (2014).
Scerbo, P. & Monsoro-Burq, A. H. The vertebrate-specific VENTX/NANOG gene empowers neural crest with ectomesenchyme potential. Sci. Adv. 6, eaaz1469 (2020).
Zalc, A. et al. Reactivation of the pluripotency program precedes formation of the cranial neural crest. Science 371, eabb4776 (2021).
Mazet, F. et al. Molecular evidence from Ciona intestinalis for the evolutionary origin of vertebrate sensory placodes. Dev. Biol. 282, 494–508 (2005).
Roure, A., Lemaire, P. & Darras, S. An otx/nodal regulatory signature for posterior neural development in ascidians. PLoS Genet. 10, e1004548 (2014).
Holland, L. Z. Tunicates. Curr. Biol. 26, R146–R152 (2016).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Brunetti, R. et al. Morphological evidence that the molecularly determined Ciona intestinalis type A and type B are different species: Ciona robusta and Ciona intestinalis. J. Zool. Syst. Evol. Res. 53, 186–193 (2015).
Adameyko, I. et al. Schwann cell precursors from nerve innervation are a cellular origin of melanocytes in skin. Cell 139, 366–379 (2009).
Adamska, M. et al. Inner ear and lateral line expression of a zebrafish Nkx5-1 gene and its downregulation in the ears of FGF8 mutant, ace. Mech. Dev. 97, 161–165 (2000).
Apostolova, G. et al. Neurotransmitter phenotype-specific expression changes in developing sympathetic neurons. Mol. Cell. Neurosci. 35, 397–408 (2007).
Bober, E., Baum, C., Braun, T. & Arnold, H. H. A novel NK-related mouse homeobox gene: expression in central and peripheral nervous structures during embryonic development. Dev. Biol. 162, 288–303 (1994).
Boisset, G. & Schorderet, D. F. Zebrafish hmx1 promotes retinogenesis. Exp. Eye Res. 105, 34–42 (2012).
Herbrand, H. et al. Two regulatory genes, cNkx5-1 and cPax2, show different responses to local signals during otic placode and vesicle formation in the chick embryo. Development 125, 645–654 (1998).
Munroe, R. J. et al. Mouse H6 homeobox 1 (Hmx1) mutations cause cranial abnormalities and reduced body mass. BMC Dev. Biol 9, 27 (2009).
Quina, L. A. et al. Deletion of a conserved regulatory element required for Hmx1 expression in craniofacial mesenchyme in the dumbo rat: a newly identified cause of congenital ear malformation. Dis. Model Mech. 5, 812–822 (2012).
Hartwell, R. D. et al. Anteroposterior patterning of the zebrafish ear through Fgf- and Hh-dependent regulation of hmx3a expression. PLoS Genet. 15, e1008051 (2019).
Liu, J. et al. Evolutionarily conserved regulation of hypocretin neuron specification by Lhx9. Development 142, 1113–1124 (2015).
Lara-Ramirez, R., Poncelet, G., Patthey, C. & Shimeld, S. M. The structure, splicing, synteny and expression of lamprey COE genes and the evolution of the COE gene family in chordates. Dev. Genes Evol. 227, 319–338 (2017).
Smith, J. J. et al. Sequencing of the sea lamprey (Petromyzon marinus) genome provides insights into vertebrate evolution. Nat. Genet. 45, 415–421 (2013).
Smith, J. J. et al. The sea lamprey germline genome provides insights into programmed genome rearrangement and vertebrate evolution. Nat. Genet. 50, 270–277 (2018).
Mehta, T. K. et al. Evidence for at least six Hox clusters in the Japanese lamprey (Lethenteron japonicum). Proc. Natl Acad. Sci. USA 110, 16044–16049 (2013).
Woolfe, A. et al. CONDOR: a database resource of developmentally associated conserved non-coding elements. BMC Dev. Biol. 7, 100 (2007).
Edgar, R. C. MUSCLE: multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res. 32, 1792–1797 (2004).
Stamatakis, A. RAxML version 8: a tool for phylogenetic analysis and post-analysis of large phylogenies. Bioinformatics 30, 1312–1313 (2014).
Marletaz, F. et al. Amphioxus functional genomics and the origins of vertebrate gene regulation. Nature 564, 64–70 (2018).
Lara-Ramirez, R., Patthey, C. & Shimeld, S. M. Characterization of two neurogenin genes from the brook lamprey Lampetra planeri and their expression in the lamprey nervous system. Dev. Dyn. 244, 1096–1108 (2015).
Boorman, C. J. & Shimeld, S. M. Pitx homeobox genes in Ciona and amphioxus show left–right asymmetry is a conserved chordate character and define the ascidian adenohypophysis. Evol. Dev. 4, 354–365 (2002).
Fuentes, M. et al. Insights into spawning behavior and development of the European amphioxus (Branchiostoma lanceolatum). J. Exp. Zool. B 308, 484–493 (2007).
Fuentes, M. et al. Preliminary observations on the spawning conditions of the European amphioxus (Branchiostoma lanceolatum) in captivity. J. Exp. Zool. B 302, 384–391 (2004).
Holland, P. W. H. Wholemount in situ hybridization to amphioxus embryos. Methods Mol. Biol. 97, 641–644 (1999).
Parker, H. J., Sauka-Spengler, T., Bronner, M. & Elgar, G. A reporter assay in lamprey embryos reveals both functional conservation and elaboration of vertebrate enhancers. PLoS ONE 9, e85492 (2014).
Schindelin, J. et al. Fiji: an open-source platform for biological-image analysis. Nat. Methods 9, 676–682 (2012).
Corbo, J. C., Levine, M. & Zeller, R. W. Characterization of a notochord-specific enhancer from the Brachyury promoter region of the ascidian, Ciona intestinalis. Development 124, 589–602 (1997).
Nakamura, M. J., Terai, J., Okubo, R., Hotta, K. & Oka, K. Three-dimensional anatomy of the Ciona intestinalis tailbud embryo at single-cell resolution. Dev. Biol. 372, 274–284 (2012).
Bolger, A. M., Lohse, M. & Usadel, B. Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics 30, 2114–2120 (2014).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Harafuji, N., Keys, D. N. & Levine, M. Genome-wide identification of tissue-specific enhancers in the Ciona tadpole. Proc. Natl Acad. Sci. USA 99, 6802–6805 (2002).
Chen, W. C. et al. Dissection of a Ciona regulatory element reveals complexity of cross-species enhancer activity. Dev. Biol. 390, 261–272 (2014).
Kari, W., Zeng, F., Zitzelsberger, L., Will, J. & Rothbacher, U. Embryo microinjection and electroporation in the chordate Ciona intestinalis. J. Vis. Exp. 16, 54313 (2016).
Stolfi, A., Gandhi, S., Salek, F. & Christiaen, L. Tissue-specific genome editing in Ciona embryos by CRISPR–Cas9. Development 141, 4115–4120 (2014).
Acknowledgements
We thank A. Stolfi and R. Zeller for sharing plasmids used in the Ciona CRISPR and overexpression studies, respectively; H. Escriva for hosting C.P. and for access to his amphioxus facility; and S. Green for lamprey husbandry assistance. V.P. was supported by a Natural Motion scholarship. V.P. and S.M.S. acknowledge the Elizabeth Hannah Jenkinson fund for financial support. V.P. also thanks T. Manousaki and C. Tsigenopoulos for their support while based in HCMR. A.P. was supported by the Accademia Nazionale dei Lincei while working in Oxford and by the H2020 Marie Sklodowska-Curie COFUND ARDRE to U.R. while working in Innsbruck. C.P. was supported by an EMBO Long Term Fellowship while working in Oxford. M.E.B. acknowledges support from award R35NS111564 from the NIH. H.J.P. was supported by funds from the Stowers Institute (grant no. 1001).
Author information
Authors and Affiliations
Contributions
V.P., C.P and S.M.S. conceived the study. V.P. conducted lamprey gene expression analysis, CNE identification, analysis of lamprey reporter gene experiments, Ciona Hmx expression analysis, Ciona Hmx overexpression analysis and RNA-seq and the molecular phylogenetic analyses. A.P. conducted Ciona CNE identification and reporter gene experiments, tests of lamprey CNE activity in Ciona, CRISPR and overexpression reporter analyses, and analysis of Ciona Hmx and Ngn gene expression. C.P. conducted the amphioxus in situ hybridization and participated in RNA-seq data analysis. H.J.P. conducted the lamprey reporter construct injections and analysis. U.R., M.E.B. and S.M.S. supervised the work. All authors contributing to drafting and editing the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature thanks Noriyuki Satoh and the other, anonymous reviewers for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
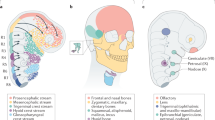
Extended Data Fig. 1 Expression of jawed vertebrate Hmx genes in neural derivatives.
The summaries show expression by gene cluster, by genome duplication paralogue (as in the associated diagram), or overall.
Extended Data Fig. 2 Schematics of experimental strategies for reporter assays and Ciona Hmx CNE identification.
a. Hmx overexpression in Ciona. b. Hmx or Ngn CRISPR Cas9 knockout in Ciona. c. Ciona Hmx CNE identification. Approximately 2 Kbp 5′ to the first Hmx exon in Ciona intestinalis (Type A) (Ciona robusta) scaffold KhS563 is shown, with conservation to the Hmx locus in Ciona savignyi shown below. d. Ciona Hmx CNE analysis in Ciona.
Extended Data Fig. 3 CRISPR-Cas9 knockout of Ciona Hmx and Ciona Ngn.
a. Placement of sgRNA guides for Hmx CRISPR knockout and primers used for validation, relative to gene structure. Guide and primer sequences in Methods and Supplementary Table 1. b. Predicted engineered outcome of Hmx CRISPR knockout. c. PCR amplification of Ciona intestinalis (Type B) genomic DNA from wild type, CRISPR control and Hmx CRISPR embryo DNA (as well as from additional sgRNAs that were tested but not used in further experiments). The guide used in further experiments is marked in yellow. Sizes of bands in the DNA ladder (100bp DNA-Ladder, extended: Carl Roth) are given in base pairs (bp). d. Sequencing of amplified bands with sequence identity matching the predicted outcomes in (b). e. Placement of sgRNA guides for Ngn CRISPR knockout and primers used for validation, relative to gene structure. Guide and primer sequences in Methods and Supplementary Table 1. f. Predicted engineered outcome of Ngn CRISPR knockout. g. PCR amplification of Ciona intestinalis (Type B) genomic DNA from CRISPR control and Ngn CRISPR embryo DNA (as well as from additional sgRNAs that were tested but not used in further experiments). The guide used in further experiments is marked in yellow. Sizes of bands in the DNA ladder (100 bp DNA-Ladder, extended: Carl Roth) are given in base pairs (bp). h. Sequencing of amplified bands with sequence identity matching the predicted outcomes in (f).
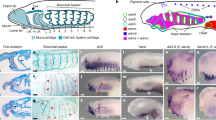
Extended Data Fig. 4 Early developmental expression of Hmx and Ngn in C. intestinalis (Type B).
Gene expression was analysed by whole mount in situ hybridisation. Only posterior BTNs (arrowheads) are marked by faint Hmx expression during neurula stages, while Ngn is expressed in posterior BTNs (arrowheads) and anterior BTNs (arrows) and the CNS. Scale bars 100 μM.
Extended Data Fig. 5 Molecular phylogenetic analysis of chordate Hmx sequences and alignment of lamprey Hmx sequences.
a. This phylogenetic analysis includes Hmx sequences from amphioxus and Ciona. The analysis was conducted using the Maximum Likelihood method and numbers indicate percentage node support out of 1000 bootstraps. b. Lamprey HmxA, HmxB and HmxC nucleotide sequence alignment. The translation shows the identical homeodomain amino-acid sequence encoded by all three genes. HmxA and HmxC share additional nucleotide sequence identity before and after the homeodomain encoding sequence. Nucleotide sequences are from the lamprey Lethenteron camtschaticum.
Extended Data Fig. 6 Model of evolution of vertebrate Hmx uCNE and dCNE from an ancestral udCNE.
CNE activity is shown in green on the embryo diagrams in the Central Nervous System (CNS) and Cranial Sensory Ganglia (CSG).
Extended Data Fig. 7 Assessment of background deriving from the vector used to generate lamprey transgenics.
Embryos were injected with vector only (which includes the zebrafish krt4 minimal promoter and reporter gene but no cloned enhancer), allowed to develop then fixed and labelled for DNA (DAPI, blue), GFP (green) and Hu/ELAV (red) before analysis by confocal microscopy. Each embryo was scored for expression in multiple tissues, as shown in the table at the top right of the picture, with a focus on tissues overlapping with Hmx expression. D and A indicate dorsal and anterior orientations for each image. CSG, cranial sensory ganglia. G, geniculate ganglion. VA, vestibuloacoustic ganglion. P, petrosal ganglion. Spinal cord expression was confined to isolated cells and distinct from the consistent column of expression generated by Hmx enhancers (see Fig. 4, main text). Brain expression appeared in the dorsal hindbrain and midbrain and was also distinct from Hmx and Hmx enhancer expression. In two embryos (1 and 2 below: also in high magnification in top left focused on the otic area) close examination revealed scattered cells around some cranial ganglia, including a few co-expressing Hu/ELAV. These differed from those labelled with Hmx enhancers in that GFP staining did not penetrate into axons. Scale bars 100μM except for the high magnification views of the otic region where they are 10μM.
Supplementary information
Supplementary Information cover sheet
This file contains table of contents for supplementary files.
Supplementary Table 1
Primers used in this study.
Supplementary Files 1–9
Supplementary Files 1–9
Rights and permissions
About this article
Cite this article
Papadogiannis, V., Pennati, A., Parker, H.J. et al. Hmx gene conservation identifies the origin of vertebrate cranial ganglia. Nature 605, 701–705 (2022). https://doi.org/10.1038/s41586-022-04742-w
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-022-04742-w
- Springer Nature Limited
This article is cited by
-
Ascidian embryonic cells with properties of neural-crest cells and neuromesodermal progenitors of vertebrates
Nature Ecology & Evolution (2024)
-
Comparative analysis of Hmx expression and the distribution of neuronal somata in the trigeminal ganglion in lamprey and shark: insights into the homology of the trigeminal nerve branches and the evolutionary origin of the vertebrate jaw
Zoological Letters (2023)