Abstract
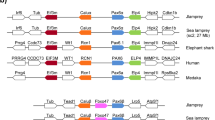
COE genes encode transcription factors that have been found in all metazoans examined to date. They possess a distinctive domain structure that includes a DNA-binding domain (DBD), an IPT/TIG domain and a helix-loop-helix (HLH) domain. An intriguing feature of the COE HLH domain is that in jawed vertebrates it is composed of three helices, compared to two in invertebrates. We report the isolation and expression of two COE genes from the brook lamprey Lampetra planeri and compare these to COE genes from the lampreys Lethenteron japonicum and Petromyzon marinus. Molecular phylogenetic analyses do not resolve the relationship of lamprey COE genes to jawed vertebrate paralogues, though synteny mapping shows that they all derive from duplication of a common ancestral genomic region. All lamprey genes encode conserved DBD, IPT/TIG and HLH domains; however, the HLH domain of lamprey COE-A genes encodes only two helices while COE-B encodes three helices. We also identified COE-B splice variants encoding either two or three helices in the HLH domain, along with other COE-A and COE-B splice variants affecting the DBD and C-terminal transactivation regions. In situ hybridisation revealed expression in the lamprey nervous system including the brain, spinal cord and cranial sensory ganglia. We also detected expression of both genes in mesenchyme in the pharyngeal arches and underlying the notochord. This allows us to establish the primitive vertebrate expression pattern for COE genes and compare this to that of invertebrate chordates and other animals to develop a model for COE gene evolution in chordates.







Similar content being viewed by others
References
Akerblad P, Lind U, Liberg D, Bamberg K, Sigvardsson M (2002) Early B-cell factor (O/E-1) is a promoter of adipogenesis and involved in control of genes important for terminal adipocyte differentiation. Mol Cell Biol 22:8015–8025
Akerblad P et al (2005) Gene expression analysis suggests that EBF-1 and PPARgamma2 induce adipogenesis of NIH-3T3 cells with similar efficiency and kinetics. Physiol Genomics 23:206–216. https://doi.org/10.1152/physiolgenomics.00015.2005
Bally-Cuif L, Dubois L, Vincent A (1998) Molecular cloning of Zcoe2, the zebrafish homolog of Xenopus Xcoe2 and mouse EBF-2, and its expression during primary neurogenesis. Mech Dev 77:85–90
Bielle F et al (2011) Slit2 activity in the migration of guidepost neurons shapes thalamic projections during development and evolution. Neuron 69:1085–1098. https://doi.org/10.1016/j.neuron.2011.02.026
Corradi A et al (2003) Hypogonadotropic hypogonadism and peripheral neuropathy in Ebf2-null mice. Development 130:401–410
Crozatier M, Vincent A (1999) Requirement for the Drosophila COE transcription factor Collier in formation of an embryonic muscle: transcriptional response to notch signalling. Development 126:1495–1504
Crozatier M, Valle D, Dubois L, Ibnsouda S, Vincent A (1996) Collier, a novel regulator of Drosophila head development, is expressed in a single mitotic domain. Curr Biol 6:707–718
Daburon V, Mella S, Plouhinec JL, Mazan S, Crozatier M, Vincent A (2008) The metazoan history of the COE transcription factors. Selection of a variant HLH motif by mandatory inclusion of a duplicated exon in vertebrates. BMC Evol Biol 8:131. https://doi.org/10.1186/1471-2148-8-131
Davis JA, Reed RR (1996) Role of Olf-1 and Pax-6 transcription factors in neurodevelopment. J Neurosci 16:5082–5094
Demilly A, Simionato E, Ohayon D, Kerner P, Garces A, Vervoort M (2011) Coe genes are expressed in differentiating neurons in the central nervous system of protostomes. PLoS One 6:e21213. https://doi.org/10.1371/journal.pone.0021213
Dubois L, Vincent A (2001) The COE—Collier/Olf1/EBF—transcription factors: structural conservation and diversity of developmental functions. Mech Dev 108:3–12
Dubois L, Bally-Cuif L, Crozatier M, Moreau J, Paquereau L, Vincent A (1998) XCoe2, a transcription factor of the Col/Olf-1/EBF family involved in the specification of primary neurons in Xenopus. Curr Biol 8:199–209
El-Magd MA, Saleh AA, El-Aziz RM, Salama MF (2014a) The effect of RA on the chick Ebf1-3 genes expression in somites and pharyngeal arches. Dev Genes Evol 224:245–253. https://doi.org/10.1007/s00427-014-0483-y
El-Magd MA, Saleh AA, Farrag F, Abd El-Aziz RM, Ali HA, Salama MF (2014b) Regulation of chick Ebf1–3 gene expression in the pharyngeal arches, cranial sensory ganglia and placodes. Cells Tissues Organs 199:278–293. https://doi.org/10.1159/000369880
Fields S, Ternyak K, Gao H, Ostraat R, Akerlund J, Hagman J (2008) The ‘zinc knuckle’ motif of Early B cell Factor is required for transcriptional activation of B cell-specific genes. Mol Immunol 45:3786–3796. https://doi.org/10.1016/j.molimm.2008.05.018
Furlong RF, Holland PW (2002) Were vertebrates octoploid? Philos Trans R Soc Lond Ser B Biol Sci 357:531–544. https://doi.org/10.1098/rstb.2001.1035
Garcia-Dominguez M, Poquet C, Garel S, Charnay P (2003) Ebf gene function is required for coupling neuronal differentiation and cell cycle exit. Development 130:6013–6025. https://doi.org/10.1242/dev.00840
Garel S, Marin F, Mattei MG, Vesque C, Vincent A, Charnay P (1997) Family of Ebf/Olf-1-related genes potentially involved in neuronal differentiation and regional specification in the central nervous system. Dev Dyn 210:191–205. https://doi.org/10.1002/(SICI)1097-0177(199711)210:3<191::AID-AJA1>3.0.CO;2-B
Garel S, Garcia-Dominguez M, Charnay P (2000) Control of the migratory pathway of facial branchiomotor neurones. Development 127:5297–5307
Grabherr MG et al (2011) Full-length transcriptome assembly from RNA-Seq data without a reference genome. Nat Biotechnol 29:644–U130. https://doi.org/10.1038/nbt.1883
Green YS, Vetter ML (2011) EBF proteins participate in transcriptional regulation of Xenopus muscle development. Dev Biol 358:240–250. https://doi.org/10.1016/j.ydbio.2011.07.034
Hagman J, Gutch MJ, Lin H, Grosschedl R (1995) EBF contains a novel zinc coordination motif and multiple dimerization and transcriptional activation domains. EMBO J 14:2907–2916
Hall TA (1999) BioEdit: a user-friendly biological sequence alignment editor and analysis program for Windows 95/98/NT. Nucleic Acids Symp Ser 41:95–98
Hesslein DG et al (2009) Ebf1-dependent control of the osteoblast and adipocyte lineages. Bone 44:537–546. https://doi.org/10.1016/j.bone.2008.11.021
Imai KS, Hino K, Yagi K, Satoh N, Satou Y (2004) Gene expression profiles of transcription factors and signaling molecules in the ascidian embryo: towards a comprehensive understanding of gene networks. Development 131:4047–4058. https://doi.org/10.1242/dev.01270
Jackson DJ et al (2010) Developmental expression of COE across the Metazoa supports a conserved role in neuronal cell-type specification and mesodermal development. Dev Genes Evol 220:221–234. https://doi.org/10.1007/s00427-010-0343-3
Jimenez MA, Akerblad P, Sigvardsson M, Rosen ED (2007) Critical role for Ebf1 and Ebf2 in the adipogenic transcriptional cascade. Mol Cell Biol 27:743–757. https://doi.org/10.1128/MCB.01557-06
Jin S et al (2014) Ebf factors and MyoD cooperate to regulate muscle relaxation via Atp2a1. Nat Commun 5:3793. https://doi.org/10.1038/ncomms4793
Katoh K, Toh H (2008) Recent developments in the MAFFT multiple sequence alignment program. Brief Bioinform 9:286–298. https://doi.org/10.1093/bib/bbn013
Katoh K, Misawa K, Kuma K, Miyata T (2002) MAFFT: a novel method for rapid multiple sequence alignment based on fast Fourier transform. Nucleic Acids Res 30:3059–3066
Kieslinger M et al (2005) EBF2 regulates osteoblast-dependent differentiation of osteoclasts. Dev Cell 9:757–767. https://doi.org/10.1016/j.devcel.2005.10.009
Kozmik Z, Holland ND, Kalousova A, Paces J, Schubert M, Holland LZ (1999) Characterization of an amphioxus paired box gene, AmphiPax2/5/8: developmental expression patterns in optic support cells, nephridium, thyroid-like structures and pharyngeal gill slits, but not in the midbrain-hindbrain boundary region. Development 126:1295–1304
Kuraku S, Meyer A, Kuratani S (2009) Timing of genome duplications relative to the origin of the vertebrates: did cyclostomes diverge before or after? Mol Biol Evol 26:47–59. https://doi.org/10.1093/molbev/msn222
Lara-Ramirez R, Patthey C, Shimeld SM (2015) Characterization of two neurogenin genes from the brook lamprey lampetra planeri and their expression in the lamprey nervous system. Dev Dyn. https://doi.org/10.1002/dvdy.24273
Li S, Yin M, Liu S, Chen Y, Yin Y, Liu T, Zhou J (2010) Expression of ventral diencephalon-enriched genes in zebrafish. Dev Dyn 239:3368–3379. https://doi.org/10.1002/dvdy.22467
Lin H, Grosschedl R (1995) Failure of B-cell differentiation in mice lacking the transcription factor EBF. Nature 376:263–267. https://doi.org/10.1038/376263a0
Lopez-Bendito G et al (2006) Tangential neuronal migration controls axon guidance: a role for neuregulin-1 in thalamocortical axon navigation. Cell 125:127–142. https://doi.org/10.1016/j.cell.2006.01.042
Louis A, Nguyen NT, Muffato M, Roest Crollius H (2015) Genomicus update 2015: KaryoView and MatrixView provide a genome-wide perspective to multispecies comparative genomics. Nucleic Acids Res 43:D682–D689. https://doi.org/10.1093/nar/gku1112
Malgaretti N et al (1997) Mmot1, a new helix-loop-helix transcription factor gene displaying a sharp expression boundary in the embryonic mouse brain. J Biol Chem 272:17632–17639
Mazet F, Masood S, Luke GN, Holland ND, Shimeld SM (2004) Expression of AmphiCoe, an amphioxus COE/EBF gene, in the developing central nervous system and epidermal sensory neurons. Genesis 38:58–65. https://doi.org/10.1002/gene.20006
Mazet F, Hutt JA, Milloz J, Millard J, Graham A, Shimeld SM (2005) Molecular evidence from Ciona intestinalis for the evolutionary origin of vertebrate sensory placodes. Dev Biol 282:494–508. https://doi.org/10.1016/j.ydbio.2005.02.021
Mehta TK et al (2013) Evidence for at least six Hox clusters in the Japanese lamprey (Lethenteron japonicum). Proc Natl Acad Sci U S A 110:16044–16049. https://doi.org/10.1073/pnas.1315760110
Murakami Y, Ogasawara M, Sugahara F, Hirano S, Satoh N, Kuratani S (2001) Identification and expression of the lamprey Pax6 gene: evolutionary origin of the segmented brain of vertebrates. Development 128:3521–3531
Murakami Y, Ogasawara M, Satoh N, Sugahara F, Myojin M, Hirano S, Kuratani S (2002) Compartments in the lamprey embryonic brain as revealed by regulatory gene expression and the distribution of reticulospinal neurons. Brain Res Bull 57:271–275
Murakami Y, Pasqualetti M, Takio Y, Hirano S, Rijli FM, Kuratani S (2004) Segmental development of reticulospinal and branchiomotor neurons in lamprey: insights into the evolution of the vertebrate hindbrain. Development 131:983–995. https://doi.org/10.1242/dev.00986
Nakatani Y, Takeda H, Kohara Y, Morishita S (2007) Reconstruction of the vertebrate ancestral genome reveals dynamic genome reorganization in early vertebrates. Genome Res 17:1254–1265. https://doi.org/10.1101/gr.6316407
Osorio J, Mazan S, Retaux S (2005) Organisation of the lamprey (Lampetra fluviatilis) embryonic brain: insights from LIM-homeodomain, Pax and hedgehog genes. Dev Biol 288:100–112. https://doi.org/10.1016/j.ydbio.2005.08.042
Pang K, Matus DQ, Martindale MQ (2004) The ancestral role of COE genes may have been in chemoreception: evidence from the development of the sea anemone, Nematostella vectensis (Phylum Cnidaria; Class Anthozoa). Dev Genes Evol 214:134–138. https://doi.org/10.1007/s00427-004-0383-7
Parker HJ, Bronner ME, Krumlauf R (2016) The vertebrate Hox gene regulatory network for hindbrain segmentation: evolution and diversification: coupling of a Hox gene regulatory network to hindbrain segmentation is an ancient trait originating at the base of vertebrates. BioEssays 38:526–538. https://doi.org/10.1002/bies.201600010
Pozzoli O, Bosetti A, Croci L, Consalez GG, Vetter ML (2001) Xebf3 is a regulator of neuronal differentiation during primary neurogenesis in Xenopus. Dev Biol 233:495–512. https://doi.org/10.1006/dbio.2001.0230
Putnam NH et al (2008) The amphioxus genome and the evolution of the chordate karyotype. Nature 453:1064–1071. https://doi.org/10.1038/nature06967
Rajakumari S et al (2013) EBF2 determines and maintains brown adipocyte identity. Cell Metab 17:562–574. https://doi.org/10.1016/j.cmet.2013.01.015
Razy-Krajka F, Lam K, Wang W, Stolfi A, Joly M, Bonneau R, Christiaen L (2014) Collier/OLF/EBF-dependent transcriptional dynamics control pharyngeal muscle specification from primed cardiopharyngeal progenitors. Dev Cell 29:263–276. https://doi.org/10.1016/j.devcel.2014.04.001
Ronquist F, Huelsenbeck JP (2003) MrBayes 3: Bayesian phylogenetic inference under mixed models. Bioinformatics 19:1572–1574
Schubert M, Holland ND, Escriva H, Holland LZ, Laudet V (2004) Retinoic acid influences anteroposterior positioning of epidermal sensory neurons and their gene expression in a developing chordate (amphioxus). Proc Natl Acad Sci U S A 101:10320–10325. https://doi.org/10.1073/pnas.0403216101
Shimeld SM, Donoghue PC (2012) Evolutionary crossroads in developmental biology: cyclostomes (lamprey and hagfish). Development 139:2091–2099. https://doi.org/10.1242/dev.074716
Singh PP, Arora J, Isambert H (2015) Identification of ohnolog genes originating from whole genome duplication in early vertebrates, based on synteny comparison across multiple genomes. PLoS Comput Biol 11:e1004394. https://doi.org/10.1371/journal.pcbi.1004394
Siponen MI et al (2010) Structural determination of functional domains in early B-cell factor (EBF) family of transcription factors reveals similarities to Rel DNA-binding proteins and a novel dimerization motif. J Biol Chem 285:25875–25879. https://doi.org/10.1074/jbc.C110.150482
Smith JJ, Keinath MC (2015) The sea lamprey meiotic map improves resolution of ancient vertebrate genome duplications. Genome Res 25:1081–1090. https://doi.org/10.1101/gr.184135.114
Smith JJ et al (2013) Sequencing of the sea lamprey (Petromyzon marinus) genome provides insights into vertebrate evolution. Nat Genet 45:415–421, 421e411–412. https://doi.org/10.1038/ng.2568
Sugahara F, Aota S, Kuraku S, Murakami Y, Takio-Ogawa Y, Hirano S, Kuratani S (2011) Involvement of Hedgehog and FGF signalling in the lamprey telencephalon: evolution of regionalization and dorsoventral patterning of the vertebrate forebrain. Development 138:1217–1226. https://doi.org/10.1242/dev.059360
Sugahara F et al (2016) Evidence from cyclostomes for complex regionalization of the ancestral vertebrate brain. Nature 531:97–100. https://doi.org/10.1038/nature16518
Tahara Y (1988) Normal stages of development in the lamprey, Lampetra-reissneri (Dybowski). Zoological Science 5:109–118
Tamura K, Peterson D, Peterson N, Stecher G, Nei M, Kumar S (2011) MEGA5: molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods. Molecular Biology and Evolution 28:2731–2739. https://doi.org/10.1093/molbev/msr121
Treiber N, Treiber T, Zocher G, Grosschedl R (2010) Structure of an Ebf1:DNA complex reveals unusual DNA recognition and structural homology with Rel proteins. Genes Dev 24:2270–2275. https://doi.org/10.1101/gad.1976610
Tulenko FJ et al (2013) Body wall development in lamprey and a new perspective on the origin of vertebrate paired fins. Proc Natl Acad Sci U S A 110:11899–11904. https://doi.org/10.1073/pnas.1304210110
Wang MM, Reed RR (1993) Molecular cloning of the olfactory neuronal transcription factor Olf-1 by genetic selection in yeast. Nature 364:121–126. https://doi.org/10.1038/364121a0
Wang SS, Tsai RY, Reed RR (1997) The characterization of the Olf-1/EBF-like HLH transcription factor family: implications in olfactory gene regulation and neuronal development. J Neurosci 17:4149–4158
Wang SS, Betz AG, Reed RR (2002) Cloning of a novel Olf-1/EBF-like gene, O/E-4, by degenerate oligo-based direct selection. Mol Cell Neurosci 20:404–414
Whelan S, Goldman N (2001) A general empirical model of protein evolution derived from multiple protein families using a maximum-likelihood approach. Mol Biol Evol 18:691–699
Acknowledgements
We thank the Forestry Commission of England for permission to collect lamprey embryos and Dr. Jo Begbie for access to histology facilities. RL-R was supported by the Mexican National Council for Science and Technology (CONACYT). CP was supported by a Royal Society Newton International Fellowship and by an EMBO Long-Term Fellowship.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by Karen E. Sears
Electronic supplementary material
Figure S1
(PDF 125 kb)
Figure S2
(PDF 112 kb)
Figure S3
(PDF 83 kb)
Supplementary file 1
(FAS 8 kb)
Supplementary file 2
(FAS 7 kb)
Rights and permissions
About this article
Cite this article
Lara-Ramírez, R., Poncelet, G., Patthey, C. et al. The structure, splicing, synteny and expression of lamprey COE genes and the evolution of the COE gene family in chordates. Dev Genes Evol 227, 319–338 (2017). https://doi.org/10.1007/s00427-017-0591-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00427-017-0591-6