Abstract
Pressurized cells with strong walls make up the hydrostatic skeleton of plants. Assembly and expansion of such stressed walls depend on a family of secreted RAPID ALKALINIZATION FACTOR (RALF) peptides, which bind both a membrane receptor complex and wall-localized LEUCINE-RICH REPEAT EXTENSIN (LRXs) in a mutually exclusive way. Here we show that, in root hairs, the RALF22 peptide has a dual structural and signalling role in cell expansion. Together with LRX1, it directs the compaction of charged pectin polymers at the root hair tip into periodic circumferential rings. Free RALF22 induces the formation of a complex with LORELEI-LIKE-GPI-ANCHORED PROTEIN 1 and FERONIA, triggering adaptive cellular responses. These findings show how a peptide simultaneously functions as a structural component organizing cell wall architecture and as a feedback signalling molecule that regulates this process depending on its interaction partners. This mechanism may also underlie wall assembly and expansion in other plant cell types.







Similar content being viewed by others
Data availability
Source data have been made available for this paper. Raw imaging data and accompanying information will be provided upon request due to the size and complexity of the datasets. Root-specific transcription profiles were retrieved from The Arabidpsis eFP Browser 2.0 (https://bar.utoronto.ca/efp2/Arabidopsis/Arabidopsis_eFPBrowser2.html). Single-cell transcriptome data were collected from the Root scRNA-Seq Atlas database (https://www.zmbp-resources.uni-tuebingen.de/timmermans/plant-single-cell-browser-root-atlas/). The crystal structures of RALF4 (PDB: 6TME), LRX2 (PDB: 6QXP) and LLG2 (PDB: 6A5E) are available via the PDB. Source data are provided with this paper.
References
Kjellén, L. & Lindahl, U. Specificity of glycosaminoglycan–protein interactions. Curr. Opin. Struct. Biol. 50, 101–108 (2018).
Haas, K. T., Wightman, R., Peaucelle, A. & Höfte, H. The role of pectin phase separation in plant cell wall assembly and growth. Cell Surf. 7, 100054 (2021).
Coen, E. & Cosgrove, D. J. The mechanics of plant morphogenesis. Science 379, eade8055 (2023).
Wilson, L. A., Deligey, F., Wang, T. & Cosgrove, D. J. Saccharide analysis of onion outer epidermal walls. Biotechnol. Biofuels 14, 1–14 (2021).
Levesque-Tremblay, G., Pelloux, J., Braybrook, S. A. & Müller, K. Tuning of pectin methylesterification: consequences for cell wall biomechanics and development. Planta 242, 791–811 (2015).
Vincent, R. R. & Williams, M. A. K. Microrheological investigations give insights into the microstructure and functionality of pectin gels. Carbohydr. Res. 344, 1863–1871 (2009).
Derbyshire, P., McCann, M. C. & Roberts, K. Restricted cell elongation in Arabidopsis hypocotyls is associated with a reduced average pectin esterification level. BMC Plant Biol. 7, 1–12 (2007).
Boyer, J. S. Enzyme-less growth in Chara and terrestrial plants. Front. Plant Sci. 7, 1–15 (2016).
Rojas, E. R., Hotton, S. & Dumais, J. Chemically mediated mechanical expansion of the pollen tube cell wall. Biophys. J. 101, 1844–1853 (2011).
Dumais, J. Mechanics and hydraulics of pollen tube growth. N. Phytol. 232, 1549–1565 (2021).
Haas, K. T., Wightman, R., Meyerowitz, E. M. & Peaucelle, A. Pectin homogalacturonan nanofilament expansion drives morphogenesis in plant epidermal cells. Science 367, 1003–1007 (2020).
Ma, Y. et al. Endoreplication mediates cell size control via mechanochemical signaling from cell wall. Sci. Adv. 8, 1–11 (2022).
Bidhendi, A. J. & Geitmann, A. Relating the mechanics of the primary plant cell wall to morphogenesis. J. Exp. Bot. 67, 449–461 (2016).
Geitmann, A. & Ortega, J. K. E. Mechanics and modeling of plant cell growth. Trends Plant Sci. 14, 467–478 (2009).
Palin, R. & Geitmann, A. The role of pectin in plant morphogenesis. BioSystems 109, 397–402 (2012).
Richter, R. P., Baranova, N. S., Day, A. J. & Kwok, J. C. Glycosaminoglycans in extracellular matrix organisation: are concepts from soft matter physics key to understanding the formation of perineuronal nets? Curr. Opin. Struct. Biol. 50, 65–74 (2018).
Valentin, R. et al. Elaboration of extensin–pectin thin film model of primary plant cell wall. Langmuir 26, 9891–9898 (2010).
Fry, S. C. Isodityrosine, a new cross-linking amino acid from plant cell wall glycoprotein. Biochem. J. 204, 449–455 (1982).
Held, M. A. et al. Di-isodityrosine is the intermolecular cross-link of isodityrosine-rich extensin analogs cross-linked in vitro. J. Biol. Chem. 279, 55474–55482 (2004).
Moussu, S. & Ingram, G. The EXTENSIN enigma. Cell Surf. 9, 100094 (2023).
Pearce, G., Moura, D. S., Stratmann, J. & Ryan, C. A. RALF, a 5-kDa ubiquitous polypeptide in plants, arrests root growth and development. Proc. Natl Acad. Sci. USA 98, 12843–12847 (2001).
Abarca, A., Franck, C. M. & Zipfel, C. Family-wide evaluation of rapid alkalinization factor peptides. Plant Physiol. 187, 996–1010 (2021).
Xiao, Y. et al. Mechanisms of RALF peptide perception by a heterotypic receptor complex. Nature 572, 270–274 (2019).
Ge, Z. et al. Arabidopsis pollen tube integrity and sperm release are regulated by RALF-mediated signaling. Science 358, 1596–1600 (2017).
Ge, Z. et al. LLG2/3 are co-receptors in BUPS/ANX-RALF signaling to regulate Arabidopsis pollen tube integrity. Curr. Biol. 29, 3256–3265.e5 (2019).
Gao, Q. et al. RALF signaling pathway activates MLO calcium channels to maintain pollen tube integrity. Cell Res. 33, 71–79 (2023).
Mecchia, M. A. et al. RALF4/19 peptides interact with LRX proteins to control pollen tube growth in Arabidopsis. Science 358, 1600–1603 (2017).
Solis-Miranda, J. & Quinto, C. The CrRLK1L subfamily: one of the keys to versatility in plants. Plant Physiol. Biochem. 166, 88–102 (2021).
Moussu, S. et al. Structural basis for recognition of RALF peptides by LRX proteins during pollen tube growth. PNAS 117, 7494–7503 (2020).
Herger, A. et al. Overlapping functions and protein–protein interactions of LRR-extensins in Arabidopsis. PLoS Genet. 16, 1–24 (2020).
Zhao, C. et al. Leucine-rich repeat extensin proteins regulate plant salt tolerance in Arabidopsis. Proc. Natl Acad. Sci. USA 115, 13123–13128 (2018).
Baumberger, N. et al. Whole-genome comparison of leucine-rich repeat extensins in Arabidopsis and rice. A conserved family of cell wall proteins form a vegetative and a reproductive clade. Plant Physiol. 131, 1313–1326 (2003).
Schoenaers, S., Balcerowicz, D. & Vissenberg, K. in Pollen Tip Growth: from Biophysical Aspects to Systems Biology 167–243 (Springer, 2017).
Hocq, L., Pelloux, J. & Lefebvre, V. Connecting homogalacturonan-type pectin remodeling to acid growth. Trends Plant Sci. 22, 20–29 (2017).
Pacifici, E., Di Mambro, R., Dello Ioio, R., Costantino, P. & Sabatini, S. Acidic cell elongation drives cell differentiation in the Arabidopsis root. EMBO J. 37, 1–9 (2018).
Denyer, T. et al. Spatiotemporal developmental trajectories in the Arabidopsis root revealed using high-throughput single-cell RNA sequencing. Dev. Cell 48, 840–852.e5 (2019).
Stegmann, M. et al. The receptor kinase FER is a RALF-regulated scaffold controlling plant immune signaling. Science 355, 287–289 (2017).
Gonneau, M. et al. Receptor kinase THESEUS1 is a rapid alkalinization factor 34 receptor in Arabidopsis. Curr. Biol. https://doi.org/10.1016/j.cub.2018.05.075 (2018).
Srivastava, R., Liu, J. X., Guo, H., Yin, Y. & Howell, S. H. Regulation and processing of a plant peptide hormone, AtRALF23, in Arabidopsis. Plant J. 59, 930–939 (2009).
Schoenaers, S. et al. The auxin-regulated CrRLK1L kinase ERULUS controls cell wall composition during root hair tip growth. Curr. Biol. 28, 722–732.e6 (2018).
Duan, Q., Kita, D., Li, C., Cheung, A. Y. & Wu, H. M. FERONIA receptor-like kinase regulates RHO GTPase signaling of root hair development. Proc. Natl Acad. Sci. USA 107, 17821–17826 (2010).
Shen, Q., Bourdais, G., Pan, H., Robatzek, S. & Tang, D. Arabidopsis glycosylphosphatidylinositol-anchored protein LLG1 associates with and modulates FLS2 to regulate innate immunity. Proc. Natl Acad. Sci. USA 114, 5749–5754 (2017).
Toyota, M. et al. Glutamate triggers long-distance, calcium-based plant defense signaling. Science 361, 1112–1115 (2018).
Voetmann, L. M., Rolin, B., Kirk, R. K., Pyke, C. & Hansen, A. K. The intestinal permeability marker FITC-dextran 4kDa should be dosed according to lean body mass in obese mice. Nutr. Diabetes 13, 4–8 (2023).
Chen, H. et al. Mouse strain– and charge-dependent vessel permeability of nanoparticles at the lower size limit. Front. Chem. 10, 1–9 (2022).
Mravec, J. et al. An oligogalacturonide-derived molecular probe demonstrates the dynamics of calcium-mediated pectin complexation in cell walls of tip-growing structures. Plant J. 91, 534–546 (2017).
Jaafar, Z. et al. Meaning of xylan acetylation on xylan–cellulose interactions: a quartz crystal microbalance with dissipation (QCM-D) and molecular dynamic study. Carbohydr. Polym. 226, 115315 (2019).
Tanaka, T. Gels. Sci. Am. https://doi.org/10.1038/scientificamerican0181-124 (1981).
Shibayama, M. & Tanaka, T. Volume phase transition and related phenomena of polymer gels. Adv. Polym. Sci. 109, 1–60 (1993).
Baumberger, N., Steiner, M., Ryser, U., Keller, B. & Ringli, C. Synergistic interaction of the two paralogous Arabidopsis genes LRX1 and LRX2 in cell wall formation during root hair development. Plant J. 35, 71–81 (2003).
Baumberger, N., Ringli, C. & Keller, B. The chimeric leucine-rich repeat/extensin cell wall protein LRX1 is required for root hair morphogenesis in Arabidopsis thaliana. Genes Dev. 15, 1128–1139 (2001).
Verhertbruggen, Y., Marcus, S. E., Haeger, A., Ordaz-Ortiz, J. J. & Knox, J. P. An extended set of monoclonal antibodies to pectic homogalacturonan. Carbohydr. Res. 344, 1858–1862 (2009).
Liners, F., Letesson, J. J., Didembourg, C. & Van Cutsem, P. Monoclonal antibodies against pectin: recognition of a conformation induced by calcium. Plant Physiol. 91, 1419–1424 (1989).
Willats, W. G. T., Gilmartin, P. M., Mikkelsen, J. D. & Knox, J. P. Cell wall antibodies without immunization: generation and use of de-esterified homogalacturonan block-specific antibodies from a naive phage display library. Plant J. 18, 57–65 (1999).
Moussu, S. et al. Plant cell wall patterning and expansion mediated by protein–peptide–polysaccharide interaction. Science 382, 719–725 (2023).
Langford-Smith, A., Keenan, T. D. L., Clark, S. J., Bishop, P. N. & Day, A. J. The role of complement in age-related macular degeneration: heparan sulphate, a ZIP code for complement factor H? J. Innate Immun. 6, 407–416 (2014).
Voxeur, A. et al. Oligogalacturonide production upon Arabidopsis thaliana–Botrytis cinerea interaction. Proc. Natl Acad. Sci. USA 116, 19743–19752 (2019).
Willats, W. G. T. et al. A xylogalacturonan epitope is specifically associated with plant cell detachment. Planta 218, 673–681 (2004).
Tanaka, T. et al. Phase transitions of gels. Annu. Rev. Mater. Sci. https://doi.org/10.1146/ANNUREV.MS.22.080192.001331 (1992).
Cannon, M. C. et al. Self-assembly of the plant cell wall requires an extensin scaffold. Proc. Natl Acad. Sci. USA 105, 2226–2231 (2008).
Zhu, S. et al. The RALF1–FERONIA complex phosphorylates eIF4E1 to promote protein synthesis and polar root hair growth. Mol. Plant 13, 698–716 (2020).
Li, L., Chen, H., Alotaibi, S. S. & Friml, J. RALF1 peptide triggers biphasic root growth inhibition upstream of auxin biosynthesis. Proc. Natl Acad. Sci. USA 119, e2121058119 (2022).
Shih, H. W., Miller, N. D., Dai, C., Spalding, E. P. & Monshausen, G. B. The receptor-like kinase FERONIA is required for mechanical signal transduction in Arabidopsis seedlings. Curr. Biol. 24, 1887–1892 (2014).
Tang, W. et al. Mechano-transduction via the pectin–FERONIA complex activates ROP6 GTPase signaling in Arabidopsis pavement cell morphogenesis. Curr. Biol. 32, 508–517.e3 (2022).
Feng, W. et al. The FERONIA receptor kinase maintains cell-wall integrity during salt stress through Ca2+ signaling. Curr. Biol. 28, 666–675.e5 (2018).
Brost, C. et al. Multiple cyclic nucleotide-gated channels coordinate calcium oscillations and polar growth of root hairs. Plant J. 99, 910–923 (2019).
Park, S., Szumlanski, A. L., Gu, F., Guo, F. & Nielsen, E. A role for CSLD3 during cell-wall synthesis in apical plasma membranes of tip-growing root-hair cells. Nat. Cell Biol. 13, 973–980 (2011).
Cavalier, D. M. et al. Disrupting two Arabidopsis thaliana xylosyltransferase genes results in plants deficient in xyloglucan, a major primary cell wall component. Plant Cell 20, 1519–1537 (2008).
Lin, C., Choi, H.-S. & Cho, H.-T. Root hair-specific EXPANSIN A7 is required for root hair elongation in Arabidopsis. Mol. Cells 31, 393–397 (2011).
Zhao, C. et al. The LRXs-RALFs-FER module controls plant growth and salt stress responses by modulating multiple plant hormones. Natl Sci. Rev. 8, nwaa149 (2021).
Song, L. et al. The RALF1–FERONIA complex interacts with and activates TOR signaling in response to low nutrients. Mol. Plant 15, 1120–1136 (2022).
Winter, D. et al. An ‘electronic fluorescent pictograph’ browser for exploring and analyzing large-scale biological data sets. PLoS ONE 2, 1–12 (2007).
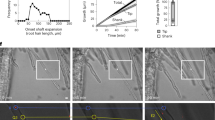
Herburger, K., Schoenaers, S., Vissenberg, K. & Mravec, J. Shank-localized cell wall growth contributes to Arabidopsis root hair elongation. Nat. Plants 8, 1222–1232 (2022).
Shimada, T. L., Shimada, T. & Hara-Nishimura, I. A rapid and non-destructive screenable marker, FAST, for identifying transformed seeds of Arabidopsis thaliana: technical advance. Plant J. 61, 519–528 (2010).
Clough, S. J. & Bent, A. F. Floral dip: a simplified method for Agrobacterium-mediated transformation of Arabidopsis thaliana. Plant J. 16, 735–743 (1998).
Costes, S. V. et al. Automatic and quantitative measurement of protein–protein colocalization in live cells. Biophys. J. 86, 3993–4003 (2004).
Futatsumori-Sugai, M. & Tsumoto, K. Signal peptide design for improving recombinant protein secretion in the Baculovirus expression vector system. Biochem. Biophys. Res. Commun. 391, 931–935 (2010).
Hashimoto, Y., Zhang, S. & Blissard, G. W. Ao38, a new cell line from eggs of the black witch moth, Ascalapha odorata (Lepidoptera: Noctuidae), is permissive for AcMNPV infection and produces high levels of recombinant proteins. BMC Biotech. 10, 50 (2010).
Hashimoto, Y., Zhang, S., Zhang, S., Chen, Y. R. & Blissard, G. W. Correction: BTI-Tnao38, a new cell line derived from Trichoplusia ni, is permissive for AcMNPV infection and produces high levels of recombinant proteins. BMC Biotech. 12, 3–6 (2012).
Kyomugasho, C., Christiaens, S., Shpigelman, A., Van Loey, A. M. & Hendrickx, M. E. FT-IR spectroscopy, a reliable method for routine analysis of the degree of methylesterification of pectin in different fruit- and vegetable-based matrices. Food Chem. 176, 82–90 (2015).
Englyst, H., Wiggins, H. S. & Cummings, J. H. Determination of the non-starch polysaccharides in plant foods by gas–liquid chromatography of constituent sugars as alditol acetates. Analyst 107, 307–318 (1982).
Thibault, J.-F. Automatisation du dosage des substances pectiques par la méthode au méta-hidroxydiphenyl. Lebensm. Wiss. Technol. 12, 247–251 (1979).
Huynh, K. & Partch, C. L. Analysis of protein stability and ligand interactions by thermal shift assay. Curr. Protoc. Protein Sci. 79, 28.9.1–28.9.14 (2015).
Gonneau, M. et al. in Plant Peptide Hormones and Growth Factors 279–293 (Springer Nature, 2024).
Weng, G. et al. HawkDock: a web server to predict and analyze the protein–protein complex based on computational docking and MM/GBSA. Nucleic Acids Res. 47, W322–W330 (2019).
Raveh, B., London, N. & Schueler-Furman, O. Subangstrom modeling of complexes between flexible peptides and globular proteins. Proteins Struct. Funct. Bioinform. 78, 2029–2040 (2010).
London, N., Raveh, B., Cohen, E., Fathi, G. & Schueler-Furman, O. Rosetta FlexPepDock web server—high-resolution modeling of peptide–protein interactions. Nucleic Acids Res. 39, 249–253 (2011).
Team, R. C. R: a language and environment for statistical computing. R Foundation for Statistical Computing https://www.r-project.org/ (2021).
Acknowledgements
This work was funded by the Research Foundation Flanders (Fonds Wetenschappelijk Onderzoek (FWO); grants 1225120N and G013023N), the University of Antwerp (UA; BOF-KP; DOCPRO4) and LASERLAB-EUROPE (grant 654148) to S.S. and K.V.; the Agence Nationale pour la Recherche (ANR), project ‘HOMEOWALL’ to H.H.; and the University of Lausanne, the European Research Council grant agreement no. 716358 and the Swiss National Science Foundation grant 310030_204526 to J.S. We thank A. Peaucelle and K. Haas for the fruitful discussions and J. Mravec, C. Ringli and S. Gilroy for sharing OG7-13647, lrx1-1/2-1 and lrx1-1 × pLRX1::cmyc-LRX1 seeds and Col-0 × 35S::GCaMP3 seeds, respectively. Our gratitude also goes to CPKelco (Denmark) for donating industrial low and high methoxyl pectin samples. The Institut Jean-Pierre Bourgin benefits from the support of Saclay Plant Sciences-SPS (ANR-17-EUR-0007). This work has benefited from the support of Institut Jean-Pierre Bourgin’s Plant Observatory technological platforms. Access to fluorescence microscopy infrastructure was provided by the Antwerp Centre for Advanced Microscopy. The purchase of the PerkinElmer UltraVIEW Vox and Leica SP8 confocal microscopes was supported by FWO-HERCULES infrastructure grants AUHA-09-001, AUHA-15-12. The Nikon Sora microscope was purchased with the support of an FWO mid-size infrastructure (I003420N) and FWO International Research Infrastructure grant (I000123N).
Author information
Authors and Affiliations
Contributions
S.S., K.V., H.H. and J.S. conceived the project. S.M. aided in conceptualizing the research based on his work on LRX8–RALF4. H.K.L. and J.S. designed, produced and characterized all recombinant proteins by SEC and SDS analysis. C.B. provided technical assistance for protein production. H.K.L. performed the TSAs. S.S. generated all crosses and performed the cloning of the RALF22-related constructs. E.A. generated and characterized the ralf22-1 line. E.A. and S.S. identified and characterized the ralf22-2 line. S.S. and E.F. optimized microfluidics for imaging. S.S. performed the phenotyping, staining and imaging of live cell and fixed-tissue samples. S.S. performed image analysis, bioinformatics and statistics. N.C. assisted with live cell imaging, cloning and sample preparations. D.B. assisted with cloning, phenotyping and sample preparation. S.S. and M.G. performed MST analysis. H.H. and B.C. conceived the QCM-D analysis. T.L. and H.H. performed the QCM-D analysis. T.L. characterized the pectin solutions. E.B. and C.M. provided technical and conceptual assistance with the in vitro QCM-D and pectin work. H.A. aided in conceptualizing the work and provided assistance with sample preparation for phenotyping. A.B., A.C., D.S.C.D and J.A.F. aided in the visualization and analysis of oscillatory parameters in live cell imaging. J.S. generated 3D protein structures and performed peptide-docking modelling. S.S., H.H. and K.V. wrote the paper.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Plants thanks Christoph Ringli and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Figs. 1–6.
Supplementary Tables
Supplementary Tables 1 and 2 (overview of statistics, product overview).
Supplementary Video 1
Time lapse video of Col-0 and ralf22-2 roots showing the short and bursting ralf22-2 RH phenotype. RALF22 loss-of-function RHs display aberrant growth and frequent bursting. Representative 2 h time lapse acquisitions of growing Col-0 and ralf22-2 (−/−) RHs.
Supplementary Video 2
Live cell imaging of changes in the growth rate, [Ca2+]cyt, pHex and cell wall physico-chemistry of Col-0 and ralf22-2 RHs being treated with 5 µM RALF22. Exogenous RALF22 supplementation induces a FER-dependent growth arrest/signalling response and a FER-independent change in cell wall physico-chemistry. Representative time lapse acquisitions (10 min) of longitudinal optical sections of growing Col-0 and fer-4 RHs expressing the [Ca2+]cyt sensor GCaMP3, in the presence of the dextran-coupled dyes FITC (110 kDa dextran, pH sensitive) and TRITC (20 kDa dextran, pH insensitive). RHs were imaged in a microfluidics chip for 4 min before the addition of 5 µM RALF22. The RH’s response was followed for another 6 min (scale bar, 5 µm).
Supplementary Video 3
Live cell imaging of ralf22-2 × pRALF22::mCherry-RALF22mature RHs showing RALF22 microdomain formation in the RH CW. RALF22 is secreted to the RH CW where it forms immobile periodic microdomains. a, Representative time lapse acquisition (10 min) of a longitudinal optical section of a growing ralf22-2 × pRALF22::mCherry-RALF22mature RH showing mCherry–RALF22mature localization to the entire RH CW. Corresponding kymograph showing the immobility of secreted mCherry–RALF22mature microdomains (shown as vertical red lines) throughout the acquisition (scale bar, 5 µm). b, RALF22 microdomains form in the growing RH dome. Consecutive time lapse frames of the growing tip have been aligned to allow visual tracking of mCherry–RALF22mature microdomains as they move from tip to shank while the tip grows forward. The concave shape of the growing dome was straightened to generate a kymograph. Red lines in the kymograph depict mCherry–RALF22mature microdomains, which originate at the very apex and remain immobile in the cell wall as they move towards the shank (scale bar, 5 µm).
Supplementary Video 4
FRAP of ralf22-2 × pRALF22::mCherry-RALF22mature RHs showing that RALF22 is secreted at the growing tip. RALF22 is secreted at the growing RH tip. Representative time lapse acquisition (10 min) of a longitudinal optical section of a growing ralf22-2 × pRALF22::mCherry-RALF22mature RH. After 4 min of growth, mCherry–RALF22mature fluorescence was bleached in a ROI in the tip and shank. FRAP was followed for an additional 6 min. Rapid mCherry–RALF22mature fluorescence recovery was observed in the tip, but not in subapical regions (scale bar, 5 µm).
Supplementary Video 5
Time lapse video of Col-0, ralf22-2 and lrx1-1/2-1 roots showing that the lrx1-1/2-1 phenotype is indistinguishable from ralf22-2. The lrx1-1/2-1 RH phenotype is indistinguishable from ralf22-2. Representative 2 h time lapse acquisitions of growing Col-0, ralf22-2 and lrx1-1/2-1 RHs.
Supplementary Video 6
Live cell imaging of ralf22-2 and lrx1-1/2-1 RHs expressing mCherry-RALF22mature, showing that LRX1/2 is required for secretion of RALF22 to the CW. LRX1/2 is required for RALF22 secretion. Representative time lapse acquisitions (10 min) of a longitudinal optical section of growing ralf22-2 × pRALF22::mCherry-RALF22mature and lrx1-1/2-1 × pRALF22::mCherry-RALF22mature RHs illustrating the accumulation of mCherry–RALF22mature in intracellular compartments in lrx1-1/2-1 RHs.
Supplementary Data 1
Statistical source data for Supplementary Fig. 5.
Source data
Source Data Fig. 1
Statistical source data.
Source Data Fig. 2
Statistical source data.
Source Data Fig. 3
Statistical source data.
Source Data Fig. 4
Statistical source data.
Source Data Fig. 5
Statistical source data.
Source Data Fig. 6
Statistical source data.
Source Data Fig. 5e
Unprocessed gels.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Schoenaers, S., Lee, H.K., Gonneau, M. et al. Rapid alkalinization factor 22 has a structural and signalling role in root hair cell wall assembly. Nat. Plants 10, 494–511 (2024). https://doi.org/10.1038/s41477-024-01637-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41477-024-01637-8
- Springer Nature Limited