Abstract
The plant-specific expansin proteins constitute an ancient and major gene family known to have roles in regulating diverse biological processes in plants. Although the functions of many expansin genes have been identified in wheat and other species, little is known about the evolution and genomic locations of the expansin genes in wheat (Triticum aestivum). In this study, a total of 87 expansin genes were identified in the wheat genome, including 52 EXPAs, 42 EXPBs and 4 EXLAs. The EXLB gene was not found in the wheat genome. Phylogenetic tree and comparative analysis revealed amplification of the EXPBs in rice, maize and wheat. The predicted wheat expansins were distributed across 14 of 21 chromosomes with different densities, 3 tightly co-located clusters and 15 paralogous pairs, indicating that tandem duplication and segmental duplication events also played roles in the evolution of expansins in wheat. In addition, the gene structures and conserved protein domains of wheat expansins suggest high levels of conservation within the phylogenetic subgroups. Analysis of a published microarray database showed that most wheat expansin genes exhibit different expression levels in different tissues and developmental stages. To our knowledge, this is the first report of a genome-wide analysis of the wheat expansin gene family, which should provide valuable information for further elucidating the classification and putative functions of the entire gene family.





Similar content being viewed by others
Abbreviations
- BLAST:
-
The Basic Local Alignment Search Tool
- HMM:
-
Hidden Markov Model
- MEGA:
-
Molecular Evolutionary Genetics Analysis
- MUSCEL:
-
Multiple Sequence Comparison by Log-Expectation
- NJ:
-
The neighbor-joining
- SMART:
-
Simple Modular Architecture Research Tool
- GSDS:
-
Gene Structure Display Server
- NCBI:
-
National Center for Biotechnology Information
- ORF:
-
Open Reading Frame
- UTR:
-
Untranslated Regions
- Aa:
-
Amino acid
- DPBB:
-
Double-psi beta-barrel
References
Abuqamar S, Ajeb S, Sham A, Enan MR, Iratni R (2013) A mutation in the expansin-like A2 gene enhances resistance to necrotrophic fungi and hypersensitivity to abiotic stress in Arabidopsis thaliana. Mol Plant Pathol 14:813–827
Altschul SF, Gish W, Miller W, Myers EW, Lipman DJ (1990) Basic local alignment search tool. J Mol Biol 215:403–410
Bae JM, Kwak MS, Noh SA, Oh MJ, Kim YS, Shin JS (2014) Overexpression of sweetpotato expansin cDNA (IbEXP1) increases seed yield in Arabidopsis. Transgenic Res 23:657–667
Brotman Y, Briff E, Viterbo A, Chet I (2008) Role of swollenin, an expansin-like protein from Trichoderma, in plant root colonization. Plant Physiol 147:779–789
Brummell DA, Harpster MH, Civello PM, Palys JM, Bennett AB, Dunsmuir P (1999) Modification of expansin protein abundance in tomato fruit alters softening and cell wall polymer metabolism during ripening. Plant Cell 11:2203–2216
Carey RE, Cosgrove DJ (2007) Portrait of the expansin superfamily in physcomitrella patens: comparisons with angiosperm expansins. Ann Bot 99:1131–1141
Chen F, Dahal P, Bradford KJ (2001) Two tomato expansin genes show divergent expression and localization in embryos during seed development and germination. Plant Physiol 127:928–936
Cho HT, Cosgrove DJ (2000) Altered expression of expansin modulates leaf growth and pedicel abscission in Arabidopsis thaliana. Proc Natl Acad Sci USA 97:9783–9788
Cho HT, Cosgrove DJ (2002) Regulation of root hair initiation and expansin gene expression in Arabidopsis. Plant Cell 14:3237–3253
Choi D, Lee Y, Cho HT, Kende H (2003) Regulation of expansin gene expression affects growth and development in transgenic rice plants. Plant Cell 15:1386–1398
Cosgrove DJ (1997) Relaxation in a high-stress environment: the molecular bases of extensible cell walls and cell enlargement. Plant Cell 9:1031–1041
Cosgrove DJ (2000) Loosening of plant cell walls by expansins. Nature 407:321–326
Cosgrove DJ (2005) Growth of the plant cell wall. Nat Rev Mol Cell Biol 6:850–861
Cox MC, Benschop JJ, Vreeburg RA, Wagemaker CA, Moritz T, Peeters AJ, Voesenek LA (2004) The roles of ethylene, auxin, abscisic acid, and gibberellin in the hyponastic growth of submerged rumex palustris petioles. Plant Physiol 136:2948–2960
Dal Santo S, Vannozzi A, Tornielli GB, Fasoli M, Venturini L, Pezzotti M, Zenoni S (2013) Genome-wide analysis of the expansin gene superfamily reveals grapevine-specific structural and functional characteristics. PLoS One 8:62206
Dong Q, Schlueter SD, Brendel V (2004) PlantGDB, plant genome database and analysis tools. Nucleic Acids Res 32:354–359
Edgar RC (2004) MUSCLE: multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res 32:1792–1797
Grennan AK (2006) Genevestigator facilitating web-based gene-expression analysis. Plant Physiol 141:1164–1166
Guo AY, Zhu QH, Chen X, Luo JC (2007) GSDS: a gene structure display server. Yi Chuan 29:1023–1026
Guo W, Zhao J, Li X, Qin L, Yan X, Liao H (2011) A soybean beta-expansin gene GmEXPB2 intrinsically involved in root system architecture responses to abiotic stresses. Plant J 66:541–552
Haider N (2013) The origin of the B-genome of bread wheat (Triticum aestivum L). Genetika 49:303–314
Han Y, Li A, Li F, Zhao M, Wang W (2012) Characterization of a wheat (Triticum aestivum L.) expansin gene, TaEXPB23, involved in the abiotic stress response and phytohormone regulation. Plant Physiol Biochem 54:49–58
Harrison EP, McQueen-Mason SJ, Manning K (2001) Expression of six expansin genes in relation to extension activity in developing strawberry fruit. J Exp Bot 52:1437–1446
He X, Zeng J, Cao F, Ahmed IM, Zhang G, Vincze E, Wu F (2015) HvEXPB7, a novel beta-expansin gene revealed by the root hair transcriptome of tibetan wild barley, improves root hair growth under drought stress. J Exp Bot 66:7405–7419
Hemalatha N, Rajesh MK, Narayanan NK (2011) Genome-wide analysis and identification of genes related to expansin gene family in indica rice. Int J Bioinform Res Appl 7:162–167
Jeanmougin F, Thompson JD, Gouy M, Higgins DG, Gibson TJ (1998) Multiple sequence alignment with clustal X. Trends Biochem Sci 23:403–405
Jensen LJ, Gupta R, Blom N, Devos D, Tamames J, Kesmir C, Nielsen H, Staerfeldt HH, Rapacki K, Workman C, Andersen CA, Knudsen S, Krogh A, Valencia A, Brunak S (2002) Prediction of human protein function from post-translational modifications and localization features. J Mol Biol 319:1257–1265
Jensen LJ, Gupta R, Staerfeldt HH, Brunak S (2003) Prediction of human protein function according to gene ontology categories. Bioinformatics 19:635–642
Kende H, Bradford K, Brummell D, Cho HT, Cosgrove D, Fleming A, Gehring C, Lee Y, McQueen-Mason S, Rose J, Voesenek LA (2004) Nomenclature for members of the expansin superfamily of genes and proteins. Plant Mol Biol 55:311–314
Krishnamurthy P, Hong JK, Kim JA, Jeong MJ, Lee YH, Lee SI (2015) Genome-wide analysis of the expansin gene superfamily reveals Brassica rapa-specific evolutionary dynamics upon whole genome triplication. Mol Genet Genom 290:521–530
Krzywinski M, Schein J, Birol I, Connors J, Gascoyne R, Horsman D, Jones SJ, Marra MA (2009) Circos: an information aesthetic for comparative genomics. Genome Res 19:1639–1645
Lamesch P, Berardini TZ, Li D, Swarbreck D, Wilks C, Sasidharan R, Muller R, Dreher K, Alexander DL, Garcia-Hernandez M, Karthikeyan AS, Lee CH, Nelson WD, Ploetz L, Singh S, Wensel A, Huala E (2012) The Arabidopsis information resource (TAIR): improved gene annotation and new tools. Nucleic Acids Res 40:1202–1210
Lee A, Giordano W, Hirsch AM (2008) Cytokinin induces expansin gene expression in melilotus alba desrwild-type and the non-nodulating, non-mycorrhizal (NodMyc) mutant masym3. Plant Signal Behav 3:218–223
Letunic I, Doerks T, Bork P (2012) SMART 7: recent updates to the protein domain annotation resource. Nucleic Acids Res 40:302–305
Li Y, Darley CP, Ongaro V, Fleming A, Schipper O, Baldauf SL, McQueen-Mason SJ (2002) Plant expansins are a complex multigene family with an ancient evolutionary origin. Plant physiol 128:854–864
Li Y, Jones L, McQueen-Mason S (2003) Expansins and cell growth. Curr Opin Plant Biol 6:603–610
Li X, Zhao J, Tan Z, Zeng R, Liao H (2015) GmEXPB2, a cell wall beta-expansin, affects soybean nodulation through modifying root architecture and promoting nodule formation and development. Plant Physiol 169:2640–2653
Librado P, Rozas J (2009) DnaSP v5: a software for comprehensive analysis of DNA polymorphism data. Bioinformatics 25:1451–1452
Lin Z, Ni Z, Zhang Y, Yao Y, Wu H, Sun Q (2005) Isolation and characterization of 18 genes encoding α-and β-expansins in wheat (Triticum aestivum L.). Mol Genet Genom 274:548–556
Lin C, Choi HS, Cho HT (2011) Root hair-specific EXPANSIN A7 is required for root hair elongation in Arabidopsis. Mol Cells 31:393–397
Lu P, Kang M, Jiang X, Dai F, Gao J, Zhang C (2013) RhEXPA4, a rose expansin gene, modulates leaf growth and confers drought and salt tolerance to Arabidopsis. Planta 237:1547–1559
Mayer KFX (2014) A chromosome-based draft sequence of the hexaploid bread wheat (Triticum aestivum) genome. Science 345:1251788
McQueen-Mason S, Cosgrove DJ (1994) Disruption of hydrogen bonding between plant cell wall polymers by proteins that induce wall extension. Proc Natl Acad Sci USA 91:6574–6578
McQueen-Mason S, Durachko DM, Cosgrove DJ (1992) Two endogenous proteins that induce cell wall extension in plants. Plant Cell 4:1425–1433
Palapol Y, Kunyamee S, Thongkhum M, Ketsa S, Ferguson IB, van Doorn WG (2015) Expression of expansin genes in the pulp and the dehiscence zone of ripening durian (durio zibethinus) fruit. J Plant Physiol 182:33–39
Park CH, Kim TW, Son SH, Hwang JY, Lee SC, Chang SC, Kim SH, Kim SW, Kim SK (2010) Brassinosteroids control AtEXPA5 gene expression in Arabidopsis thaliana. Phytochemistry 71:380–387
Poole RL (2007) The TAIR database. Methods Mol Biol 406:179–212
Punta M, Coggill PC, Eberhardt RY, Mistry J, Tate J, Boursnell C, Pang N, Forslund K, Ceric G, Clements J, Heger A, Holm L, Sonnhammer EL, Eddy SR, Bateman A, Finn RD (2012) The Pfam protein families database. Nucleic Acids Res 40:290–301
Rozas J (2009) DNA sequence polymorphism analysis using DnaSP. Methods Mol Biol 537:337–350
Sampedro J, Cosgrove DJ (2005) The expansin superfamily. Genome Biol 6:242
Sampedro J, Carey RE, Cosgrove DJ (2006) Genome histories clarify evolution of the expansin superfamily: new insights from the poplar genome and pine ESTs. J Plant Res 119:11–21
Sarkar P, Bosneaga E, Auer M (2009) Plant cell walls throughout evolution: towards a molecular understanding of their design principles. J Exp Bot 60:3615–3635
Tamura K, Peterson D, Peterson N, Stecher G, Nei M, Kumar S (2011) MEGA5: molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods. Mol Biol Evol 28:2731–2739
Vreeburg RA, Benschop JJ, Peeters AJ, Colmer TD, Ammerlaan AH, Staal M, Elzenga TM, Staals RH, Darley CP, McQueen-Mason SJ, Voesenek LA (2005) Ethylene regulates fast apoplastic acidification and expansin a transcription during submergence-induced petiole elongation in rumex palustris. Plant J 43:597–610
Wang G, Gao Y, Wang J, Yang L, Song R, Li X, Shi J (2011) Overexpression of two cambium-abundant Chinese fir (Cunninghamia lanceolata) alpha-expansin genes ClEXPA1 and ClEXPA2 affect growth and development in transgenic tobacco and increase the amount of cellulose in stem cell walls. Plant Biotechnol J 9:486–502
Wu X, Song C, Wang B, Cheng J (2002) Hidden markov model used in protein sequence analysis. J Biomed Eng 19:455–458
Yennawar NH, Li LC, Dudzinski DM, Tabuchi A, Cosgrove DJ (2006) Crystal structure and activities of EXPB1 (Zea m 1), a β-expansin and group-1 pollen allergen from maize. Proc Natl Acad Sci USA 103:14664–14671
Zhang XQ, Wei PC, Xiong YM, Yang Y, Chen J, Wang XC (2011) Overexpression of the Arabidopsis alpha-expansin gene AtEXPA1 accelerates stomatal opening by decreasing the volumetric elastic modulus. Plant Cell Rep 30:27–36
Zhang S, Xu R, Gao Z, Chen C, Jiang Z, Shu H (2014a) A genome-wide analysis of the expansin genes in malus x domestica. Mol Genet Genom 289:225–236
Zhang W, Yan H, Chen W, Liu J, Jiang C, Jiang H, Zhu S, Cheng B (2014b) Genome-wide identification and characterization of maize expansin genes expressed in endosperm. Mol Genet Genom 289:1061–1074
Zhu Y, Wu N, Song W, Yin G, Qin Y, Yan Y, Hu Y (2014) Soybean (Glycine max) expansin gene superfamily origins: segmental and tandem duplication events followed by divergent selection among subfamilies. BMC Plant Biol 14:93–111
Acknowledgments
This work was supported by Youth Scientific Research Foundation of Shandong Academy of Agricultural Sciences (Grant No. 2014QNM09 to Nana Li), Natural Science Foundation of Shandong Province (Grant No. ZR2014YL017 to Nana Li) and Shandong Province Science and Technology Development Program (Grant No. 2012G0021031 to Nana Li).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
Nana Li, Yanyan Pu, Yongchao Gong, Yanli Yu and Hanfeng Ding declares no conflict of interest.
Ethical approval
The article does not contain any studies with human participants or animals performed by any of the authors.
Electronic supplementary material
Below is the link to the electronic supplementary material.
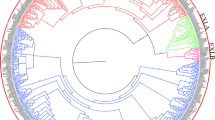
Figure S1
Phylogenetic tree of AtEXPs, OsEXPs, ZmEXPs and TaEXPs was constructed by maximum likelihood method (JPEG 4947 kb)
Table S1
The information of the Expansin genes in wheat (DOC 182 kb)
Table S2
Ka/Ks analysis and estimated divergence times for duplicated TaEXP paralogs (DOC 46 kb)
Rights and permissions
About this article
Cite this article
Li, N., Pu, Y., Gong, Y. et al. Genomic location and expression analysis of expansin gene family reveals the evolutionary and functional significance in Triticum aestivum . Genes Genom 38, 1021–1030 (2016). https://doi.org/10.1007/s13258-016-0446-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13258-016-0446-y