Abstract
This review study presents the state-of-the-art machine and deep learning-based COVID-19 detection approaches utilizing the chest X-rays or computed tomography (CT) scans. This study aims to systematically scrutinize as well as to discourse challenges and limitations of the existing state-of-the-art research published in this domain from March 2020 to August 2021. This study also presents a comparative analysis of the performance of four majorly used deep transfer learning (DTL) models like VGG16, VGG19, ResNet50, and DenseNet over the COVID-19 local CT scans dataset and global chest X-ray dataset. A brief illustration of the majorly used chest X-ray and CT scan datasets of COVID-19 patients utilized in state-of-the-art COVID-19 detection approaches are also presented for future research. The research databases like IEEE Xplore, PubMed, and Web of Science are searched exhaustively for carrying out this survey. For the comparison analysis, four deep transfer learning models like VGG16, VGG19, ResNet50, and DenseNet are initially fine-tuned and trained using the augmented local CT scans and global chest X-ray dataset in order to observe their performance. This review study summarizes major findings like AI technique employed, type of classification performed, used datasets, results in terms of accuracy, specificity, sensitivity, F1 score, etc., along with the limitations, and future work for COVID-19 detection in tabular manner for conciseness. The performance analysis of the four majorly used deep transfer learning models affirms that Visual Geometry Group 19 (VGG19) model delivered the best performance over both COVID-19 local CT scans dataset and global chest X-ray dataset.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The word “novel” is used often with coronavirus to mean that it is a new strain in the family of perilous viruses [1]. According to WHO, coronavirus belongs to a large family of viruses ranging from common cold to unsafe diseases (www.who.int). Such diseases can infect both humans and animals. The coronavirus COVID-19 strain started spreading in Wuhan, China, in December 2019, since then it has become a serious health problem in the world. The coronavirus COVID-19 strain has its place in two different coronaviruses called Middle East Respiratory Syndrome (MERS) and Severe Acute Respiratory Syndrome (SARS). Respiratory complications like pneumonia, kidney disorder, and liquid formation in the lungs are among the symptoms of coronavirus infection. Coronavirus (CoV) is the dangerous one due to its serial interval (5 to 7.5) and reproductive rate (2 to 3) [2]. The CoV has its roots in the single-stranded RNA viruses (+ ssRNA) family, mostly seen in animals [3, 4]. These viruses have no species barriers and can cause epidemics like MERS and SARS which were seen in the last two decades. The SARS-CoV began in China, blew out to twenty-four countries and caused 8000 cases and 800 deaths. The MERS-CoV started in Saudi Arabia and reported 2500 cases and 8700 deaths. About 2% of the population are healthy carriers of CoV and these viruses are accountable for approximately 5 to 10% of acute respiratory infections [5]. The virus behind COVID-19 pandemic is called Severe Acute Respiratory Syndrome Coronavirus-2 (SARS-CoV-2) [6]. The CoV details are given in Table 1.
COVID-19 is a new species discovered in 2019 that has not been previously identified in humans. Bats have been recognized as natural reservoirs and vectors of a variety of viruses including coronaviruses have crossed species barriers to infect humans as well as different kinds of animals, including avian, rodents, and chiropters [7]. The CoV is so named due to their solar corona (crown-like) appearance when observed under an electron microscope. COVID-19 is an acute resolved disease, but it can also be deadly as depicted in Fig. 1, based on the data from the WHO. Severe disease onset might result in death due to massive alveolar damage and progressive respiratory failure [8]. Respiratory droplets of size greater than 5–10 µm acts as a mode of airborne transmission (https://apps.who.int/iris/handle/10665/331601). COVID-19 carries a higher growth factor than SARS and MERS due to the fact that interaction without safety measures can be extremely contagious and it causes lighter symptoms in most cases. The top ten COVID-19 infected countries statistics in terms of infected cases and deaths are presented with the help of Fig. 1.
Faster spreading rate is a major concern for COVID-19 pandemic and thus detecting who has the COVID-19 virus infection at an early stage is critical [9] to curtail its spread. Viral nucleic acid detection using real-time polymerase chain reaction (RT-PCR) is the accepted standard diagnostic method [10, 11]. However, this test has suboptimal sensitivity and specificity and many hyper-endemic regions and countries are not able to provide sufficient RT-PCR testing for tens of thousands of suspected subjects in a short period of time. Other concerns about RT-PCR are its painfulness, lack of swabs, need of reagents, delays in producing results and substantial false-negative rate. Considering these concerns, other approaches to diagnosis are worthy of investigation [12]. All such approaches should be accurate, fast, and effective tools for detecting COVID-19 infection to provide the prerequisite for rigorous detection, contact tracing and isolation of infected subjects at primary stage of infection. Artificial intelligence techniques are now exceptionally found to be beneficial for training, forecasting, and evaluation purposes. Neural networks are widely employed for developing prediction models. But neural networks still have limitations like slow convergence and learning capability [13]. ALzubi et al. [14] demonstrated the fact that deep learning is a beneficial technique to improve the diagnostic pace, since it can be used for making predictions and clinical decisions in medical system. These researches also stated that linking medical image and diagnostic parameters is an efficient scheme that will assist doctors to perform patients’ diagnosis utilizing big data. To assist doctors to evaluate the COVID-19 disease and to optimize prevention and control measures as earliest as possible, medical imaging can be considered a vital technique to diagnose COVID-19 infections using radiological images such as X-rays or computed tomography (CT) scans. It has been established that anomalies can be found in COVID-19 patients in chest CT scans in the shape of Ground-Glass Opacities (GGO) [15]. Much research has demonstrated that a system using chest CT scans can be created for diagnosing and quantifying COVID-19 cases [16]. To detect COVID-19, X-ray images can also be utilized instead of CT scans. Hence, medical images like chest X-rays (CXR) and CT images can be studied to give comparatively instant diagnostic information by identifying possible patterns that may lead to the automatic diagnosis of the disease. Chest X-ray is the universally used imaging modality in the diagnostic checkup of patients with thoracic abnormalities, due to its fast imaging speed, low radiation, and low cost [17], universal availability in both emergency and hospital settings, where interpretation is often done without expert radiologists.
Unlike laboratory tests that involve probing the patient’s respiratory system, X-rays can be taken without the increased risk of aerosolizing the pathogen. The X-rays may also facilitate the triage of patients into highest risk, high risk, and lower risk of further complications besides indicating the severity of disease at one or more time points. Unlike computed tomography (CT) scans, chest X-rays cannot provide 3D anatomy but can differentiate pneumonia even though it is probably understood as the most challenging plain film to interpret correctly [18]. Accurate interpretation is vital for patient management in the severe situation, and to help identify clustering occurrences of COVID-19. CT, being a noninvasive imaging approach, can portray certain characteristic manifestations in the lung which are associated with COVID-19 [19, 20]. CT can be used as an effective way for early diagnosis of COVID-19 but CT may determine similar imaging features between COVID-19 and other types of pneumonia, thus making it difficult to differentiate between them. CT imaging is significantly more time consuming than X-ray imaging, and also involves complex sanitization procedures between switching patients. Moreover, sufficient high-quality CT scanners may not be commonly available, making it difficult for a timely viral pneumonia screening. The role of medical imaging is vital for the fast diagnosis of COVID-19 [18]. The first image based approach used in Spain (https://healthcare-in-europe.com/en/news/imaging-the-coronavirus-disease-covid-19.html). Hence, the combination of AI and chest imaging can facilitate the detection of complications of COVID-19 [21].
Recent research work shows that computer vision [22], machine learning [23,24,25], and deep learning [26, 27] can be used for automatic diagnosis of different ailments in the human body [28, 29]. The deep learning method is used as a feature extractor that enhances classification accuracies [30]. Although radiography can be quickly performed and generally available due to commonality of chest radiology imaging systems in hospitals, the interpretation of radiography images by radiologists is still a major concern due to the human capacity in detecting the subtle visual features present in the images. Deep learning can discover patterns in chest X-rays that can be missed by radiologists [31,32,33,34]. Deep learning, which has been used to detect tuberculosis in chest X-rays, could also be used for identifying lung abnormalities related to COVID-19 [35] due to its high capability of feature extraction [36,37,38]. This will help clinicians in deciding the order of treatment of high-risk COVID-19 patients. Deep learning was used to detect and segregate bacterial and viral pneumonia on pediatric chest radiographs [39, 40]. Efforts have also been made to detect various imaging features of chest CT scans [41, 42].
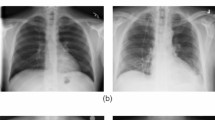
Deep learning (DL) is a branch of machine learning (ML) which is inspired by the way the human brain works and utilized for feature extraction as well as classification of images. Main strength of DL is that it is an unsupervised learning i.e., it can learn from unlabeled data. DL has been vastly used in industries, self-driven cars, face recognition, object detection, image classification, etc. [43] due to characteristics like unlabeled data utilization, working without feature engineering, prediction with high accuracy and precision. Convolutional neural network (CNN) is a DL algorithm that has been used extensively in solving problems like document analysis, different types of image classification, pose detection, and recognizing various actions [44]. Medical imaging is one of the areas where CNN has been showing encouraging results [45], and thus, convolutional neural networks (CNNs) have been doing well in detecting several diseases like coronary artery disease, malaria, Alzheimer’s disease, different dental diseases, and Parkinson’s disease. Likewise, CNN has considerable prospects for differentiating COVID-19 from non-COVID-19 infections with medical images like chest X-rays and CTs using public databases of chest X-rays and CTs. The chest X-rays and CT scans of COVID-19 positive cases and normal are presented with the help of Fig. 2.
The real motivation behind writing this review paper is to illustrate the latest trends and development in the domain of COVID-19 detection and classification approaches based on deep learning. Apart from this, an analysis and comparison is also done utilizing the five majorly used deep transfer learning models as per the literature review for the COVID-19 detection in the analysis and evaluation section. All these five models are trained and evaluated on the locally developed COVID-19 CT scan dataset and two global chest X-ray image dataset in order to observe their performance.
This state-of-the-art review paper is organized into a total of five sections. Initially, the “Introduction” section is all about illustrating the general introduction to the COVID-19 disease and its impact over the world in the present scenario. The “Research Methodology” section deals with the research methodology adopted to conduct this review study. The “Literature Review” section deals with the brief review and comparison of major automated deep learning and machine learning-based COVID-19 detection approaches proposed by the various researchers since March 2020. The “Majorly Used COVID-19 Chest X Ray, CT Scan, and Ultrasound Image Dataset Description” section elaborates the various COVID-19 chest X-rays and CT scan datasets available online for research. The “Analysis and Evaluation” section presents an analysis and evaluation among the four majorly used deep transfer learning models over the COVID-19 local CT scan and global chest X-ray datasets.
Research Methodology
The presented review study aims to assess the existing research done in the domain of deep learning application for the detection of COVID-19 utilizing the chest X-rays and CT scan images. Various databases e.g. IEEE Xplore, PubMed, Web of Science etc. are searched exhaustively with the specific search items. The research studies included in this review study are based on the following selection criteria:
-
Only deep learning-based approaches for the COVID-19 binary or multiclass classification are included.
-
The considered research studies were limited to the period from March 2020 to August 2021.
-
The research studies utilizing either the chest X-ray or CT scan imaging modalities are included. Other medical imaging modalities are excluded.
-
Only classification or detection approaches are included, whereas prediction approaches utilizing big data are not excluded from this study.
-
Research studies which mentioned the future direction or at least offered some narrative to improve the existing work.
After the elimination of duplicate and redundant works, more than 50 unique studies were considered in this review study. Table 2 below summarizes the search items employed for the searching of research studies for the COVID-19 classification and detection.
Literature Review
Since March 2020, a substantial amount of research has been carried out in the domain of COVID-19 detection based on deep learning. These deep learning models are trained and tested either using chest X-ray images or CT scan images or sometimes both. This fact is very well proved by the Figs. 3 and 4 representing the data collected from major research databases like PubMed and Web of Science. These two graphs simply illustrate the number of COVID-19 detection research studies done using either CT scan or Chest X-ray datasets and established on deep learning, deep transfer learning. Figures 5 and 6 demonstrate the general COVID-19 detection approaches based on machine learning, deep learning, and deep transfer learning whereas Fig. 7 depicts deep learning in conjunction with traditional machine learning classifiers, also known as the hybrid models.
Deep Learning and Deep Transfer Learning-Based Approaches
Ucar et al. [46] proposed a fine tuned deep learning model based on SqueezeNet and Bayesian optimization for the screening of COVID-19 patients. This Deep Bayes-SqueezeNet learning model takes chest X-ray images in order to diagnose COVID-19 disease. The proposed SqueezeNet is composed of 15 layers; 5 different layers as 2 convolution layers, 3 max pooling layers, 8 fire layers, 1 global average pooling layer, and 1 output layer softmax. This proposed model offered an accuracy of 100%, 98.04%,and 96.73% for the COVID-19, normal, and pneumonia cases. Hammoudi et al. [47] proposed a deep transfer learning-based model which is established on InceptionResNetV2 for the screening and diagnosis of COVID-19 patients. Their DenseNet169 model delivered approximately 96% average accuracy for the correct classification of pneumonia cases using the chest X-ray imaging modality. Rajaraman et al. [48] proposed an iteratively pruned deep learning model for the detection of COVID-19 using the chest X-ray images and ImageNet models. The results give the accuracy of 99.01% and area under the curve = 99.72%. Also, the CXR images are taken for clear lung, bacterial pneumonia infections, and COVID-19 pneumonia infection manifesting at peripheral opacities in the left lung. Hall et al. [49] proposed a pre-trained CNN based on ResNet50 for screening of COVID-19 and pneumonia patients with tenfold cross validation. Their model achieved an overall accuracy of 89.2% and area under the curve was 95%. Their work focuses on CXRs which are simpler and cheaper to obtain but provide less information than CT. Rahimzadeh et al. [50] proposed a deep learning model based on Xception and ResNet50V2 for screening of COVID-19 patients. The proposed model performs the multiclass classification as normal cases, pneumonia, and COVID-19 cases. In their study both the Xception and ResNet50V2 are used for extracting deep features and then the softmax classifier performs the multiclass classification. Zhang et al. [51] proposed a deep anomaly detection model for reliable and fast screening in order to identify COVID-19 from non-COVID-19 cases. This model is composed of three components, namely, a backbone network, a classification head, and an anomaly detection head. An 18-layer residual convolutional neural network pre-trained on the ImageNet dataset is used as the backbone network. Hemdan et al. [52] conducted a comparison study using the VGG19, DenseNet121, InceptionV3, ResNetV2, Inception-ResNet-V2, Xception, and MobileNetV2 DTL models for detection of COVID-19. The VGG19 and Dense CNN models showed good performances compared to other DTL models in their research study. Wang et al. [53] proposed a DL system consisting of three stages i.e. automatic lung segmentation, non-lung area suppression, and COVID-19 diagnostic and prognostic analysis. In this system, two DL networks were used initially, a DenseNet121-FPN for lung segmentation in chest CT image, and the proposed novel COVID-19Net for COVID-19 diagnostic and prognostic analysis. This COVID-19Net model used a DenseNet-like structure, consisting of four dense blocks, where each dense block had multiple stacks of convolution, batch normalization, and ReLU activation layers. Each dense block uses a dense connection to contemplate multi-level image information. Zheng et al. [54] developed a weakly-supervised deep learning-based software system using 3D CT volumes to detect COVID-19. In their system, the lung region was segmented using a pre-trained UNet and then the segmented 3D lung region was fed into a 3D deep neural network to predict the probability of COVID-19 infection. Apostolopoulos et al. [55] proposed an automated detection system based on MobileNet V2. Different strategies were utilized in their study, such as transfer learning with off-the-shelf-feature extraction, transfer learning with fine tuning, and training from scratch. The training and evaluation procedure was performed with tenfold-cross-validation. Fu et al. [56] proposed a deep learning-based diagnostic tool using the ResNet50 architecture to perform multiclass classification into seven classes. The multiclass classification occurs for COVID-19, non-COVID-19 viral pneumonia, bacterial pneumonia, pulmonary tuberculosis, or and normal lung cases. Ardakani et al. [57] presented a comparison study using the DTL models like ResNet-101, AlexNet, VGG-16, VGG-19, SqueezeNet, GoogleNet, MobileNet-V2, ResNet-18, ResNet-50, and Xception for detection of COVID-19 and Non-COVID-19. The best results were delivered by the ResNet-101 and Xception models. Their study compared the performance of radiologists in real-time with the performance of these ten DTL models for COVID-19 detection.
Rehman et al. [58] proposed an automated method for the diagnosis of COVID-19 positive cases. Their model tends to perform the multiclass classification differentiating a COVID-19 from viral, bacterial, and normal cases. Their research study compared the seven DTL pre-trained architectures of CNN which were: (1) AlexNet is composed of 5 convolutional layers and 3 fully connected layers. (2) VGG is composed of 16 convolutional layers and 3 fully connected layers. (3) SqueezeNet contains five modules and an expanded layer. (4) GoogleNet, composed of 9 inception models, 4 max-pooling layers, 2 convolutional layers, an average pooling layer, 2 normalization layers, 1 fully connected layer, and a linear layer. (5) Three variants of ResNet were used. ResNet18, composed of 5 convolution blocks, each containing 2 residual blocks. Each residual block contains 2 convolution layers. ResNet50 contains 5 residual blocks, each with a convolution and identity block. The convolution and identity blocks have 3 convolution layers. ResNet101 contains 3 convolutional, 3 residual blocks, and an identity block, (6) DenseNet contains 1 × 1 convolutional filters and max-pooling layers. (7) MobileNetv2 contains CNN layer, inverted residual, and linear bottleneck layer. Khalifa et al. [59] proposed a method for the detection of COVID-19 cases based on GAN (Generative Adversarial Network) with fine-tuned DTL models. They employed four types of DTL models, which were: (1) AlexNet, (2) SqueezeNet, (3) GoogleNet, (4) RestNet18 with 8, 18, 22, 18 layers respectively and these models are chosen due to the less number of layers, so that the complexity, consumed memory, and time can be reduced. Loey et al. [60] proposed a model based on Generative Adversarial Network (GAN) and Deep Transfer Learning model to analyze various deep transfer learning models such as AlexNet, GoogleNet, and RestNet18 to detect COVID-19 disease. In their model GoogleNet achieved 100% testing accuracy and 99.9% validation accuracy. Then Shan et al. [61] proposed an accurate deep learning-based model for automatic segmentation and quantification of infection regions of COVID-19 from chest CT scans. The proposed model utilized VB-Net segmentation to clearly segment and quantify the infection area. The proposed model offers an accuracy of 91.6% ± 10%. Hu et al. [62], proposed an approach based on the customized CNN architecture for the detection of COVID-19 cases as well as for the quantification of infection region from the chest CT scans. The customized CNN architecture with five convolutional layers performs a multiclass classification as COVID-19, CAP (community-acquired pneumonia) and non-pneumonic to detect COVID-19 disease accurately. Wang et al. [63] proposed a model for the detection of COVID-19 disease using chest X-ray images. They used Deep learning tools like VGG-19, ResNet-50, and COVID-Net with the accuracy of 83.0%, 90.6%, and 93.3% respectively. Li et al. [64] proposed a model for the detection of COVID-19 using 3D deep learning framework based on CovNet. The CovNet framework consists of ResNet50 as the backbone using CT scan images as an input. They performed three-way classification between COVID-19, CAP (community-acquired pneumonia), and non-pneumonia patients. Minaee et al. [65] proposed a model for the detection of the novel coronavirus using chest X-ray images. Four DTL models, namely ResNet18, ResNet50, SqueezeNet, and DenseNet-121 performance, were compared on a very large size dataset in their study. Among these, the SqueezNet model offered the best result of 98% sensitivity and 92.9% specificity. Basu et al. [66] proposed a model for the detection of COVID-19 disease with the help of Domain Extension Transfer Learning (DETL) and Gradient Class Activation Map (Grad-CAM). Their study used pre-trained DTL models like AlexNet, VGGNet, and ResNet which offered an accuracy of 82.98%, 90.13%, and 85.98% respectively. The proposed model performs multiclass classification between normal, pneumonia, other disease, and COVID-19 cases. Khalifa et al. [67] explored a new dimension in the deep learning and deep transfer learning application for the COVID-19 detection. Their novel research study established on the concept of neutrosophic set along with the application of DTL models. In their study, CXR images available in the grayscale domain are converted into the neutrosophic domain. The neutrosophic domain consists of three types of images: indeterminacy (I) images, true (T) images, and the falsity (F) images. Then, these neutrosophic images are used for the training of DTL models like AlexNet, GoogleNet, and ResNet18, which in turn perform the multiclass classification. The four-way classification into normal, bacterial pneumonia, viral pneumonia, and COVID-19 cases are done. Cohen et al. [68] proposed a model for severity score prediction of COVID-19 pneumonia using the CXR images. Such a tool can gauge the severity of COVID-19 lung infections. Their study used a DTL i.e. DenseNet model from the Torch X-Ray Vision Library. Their proposed model can predict geographic extent score and lung opacity score with 1.14 and 0.78 mean absolute error (MAE) respectively. Ying et al. [69], proposed a Detail Relation Extraction Network (DRE-Net)–based model to detect COVID-19 disease using chest CT scan images. Their proposed model performed the multiclass classification and its performance was compared with other DTL models like VGG16, DenseNet, and ResNet. The proposed DRE-Net model offered accuracy 94%.
Wang et al. [70] devised an alternate method for the diagnosis of COVID-19 cases, which was completely tested in a laboratory. Due to the challenges faced in the quality and availability of such laboratories in the infected areas, alternatives such as devising an artificial intelligence-based testing algorithm were proposed. This algorithm can assist the radiologist to easily differentiate between the COVID-19 positive cases and other viral pneumonias. It studied 453 enrolled CT images and used 217 as trained dataset and rest as validation set. The algorithm produced an accuracy of 82.9% in the internal validation and 73.1% in the external validation. Narin et al. [71] felt the need for an automatic COVID-19 case detection method to reduce the risk of spreading this pandemic disease at a widespread range. Three DTL models as ResNet50, InceptionV3, and Inception-ResNetV2 were proposed for detection of COVID cases using the chest X-ray images of suspected patients. The devised algorithm delivered the highest accuracy 98% with the ResNet50 model, InceptionV3, and Inception-ResNetV2 achieved 97% and 87% accuracy respectively. Jin et al. [72] proposed another deep learning-based AI system to increase the rate of diagnosis of COVID-19 disease for the welfare of society during this COVID-19 pandemic. It will enable timely detection of infected patients and help in controlling the growing rates of COVID-19 cases. The algorithm derived is a result of extensive statistical analysis of CT scan images. The analysis was done on nearly 10,000 CT volumes of community-acquired pneumonia (non-viral), influenza-A/B, non-pneumonia, and COVID-19 suspected. Xu et al. [73] proposed a model to distinguish COVID-19 pneumonia from influenza-A viral pneumonia and healthy cases with pulmonary CT images using deep learning techniques. Their CNN model is accompanied with Noisy-OR Bayesian function to come up with an accuracy of 86.7% in testing of COVID-19 cases. Huang et al. [74] developed a deep learning-based algorithm which was focused on quantitative CT. It allows measuring the severity of COVID-19 and helps in studying the growth rate and opacity percentage of lungs within the patient body. The algorithm classifies patients between mild vs. moderate vs. severe vs. critical. All the results were cross-checked by two radiologists and the follow-up test conducted after the diagnosis of opacity percentage of the lungs. Farooq et al. [75] developed an automated approach for the detection and classification of COVID-19 cases by fine-tuning a pre-trained ResNet50 architecture named COVIDNet. The dataset used in their research study consisted of 5941 CXR images from 2839 different patients. The classification is done as normal, bacterial pneumonia, viral pneumonia, and COVID-19. It comes up with an accuracy of 96.23%. Authors now want the deduced algorithm to examine a large dataset and prove its reliability for the noble purpose. Chen et al. [76] devise a new model for the detection of COVID-19. Their study is based on high-resolution CT scan images of the suspected coronavirus pneumonia patients. The devised model is based on UNet + + for image segmentation and ResNet50 for the classification to deduce the results. The results were cross-checked by three radiologists and it is found that the time taken for testing the already evaluated images by radiologists is very less compared to evaluating new images. Asnaoui et al. [77] inspired by the achievements of the medical image analysis technique and motivated to publish a research based on the deep convolutional neural network (DCNN) architectures such as VGG16, VGG19, CNN, Inception_V3, Xception, Resnet50, Inception_Resnet_V2, DenseNet201, and MobileNet_V2. The results were classified in two parts as normal vs. pneumonia. To obtain the results, a total of 5856 images of chest X-ray and CT images were studied, of which 4273 were of pneumonic patients and the rest 1583 were of normal humans. The highest accuracy was achieved by Resnet50 and MobileNet_V2 architecture with 96.61% and 96.27% respectively. Chowdhury et al. [78] implemented and evaluated the eight different pre-trained models known as MobileNetv2, SqueezeNet, ResNet18, ResNet101, DenseNet201, CheXNet, Inceptionv3, and VGG19. The result was classified as normal vs. COVID-19 pneumonia vs. viral pneumonia. Chest X-ray images of 423 COVID-19 patients, 1485 viral pneumonia patients, and 1579 normal patients were examined. DenseNet201 leads the results with the highest accuracy among other models with 99.70%. Apostolopoulos et al. [79] came up with a comparison study of various DTL models for the detection of COVID-19 cases. The models used were VGG19, MobileNet v2, Inception, Xception, and Inception ResNet v2. The highest three-class accuracy was achieved by VGG19 among all the models used. Moreover, the researchers took two datasets to devise the results of their study. Afshar et al. [80] understood that RT-PCR is a time consuming test, which is not desirable and includes too much physical contact with the COVID-19 patients. So, the authors collected the X-ray images from two different datasets and developed a capsule based framework named COVID-CAPS. The COVID-CAPS framework consists of 4 convolutional layers and 3 capsule layers. The result of the study was a binary classification into either COVID-positive patients or COVID-negative patients. The accuracy of the proposed work comes out to be 95.7% with sensitivity around 90% and specificity of about 95.8%. Butt et al. [81] realized the efficiency of artificial intelligence in diagnosing COVID cases. It is a clear fact that early detection of the infection will lead to a way of reducing mortality rates. It has been noted that radiographic patterns are far more fast and accurate in generating results when compared to RT-PCR detection of COVID-19 victims. Studies showed that the detection of different types of viral pneumonia becomes an easier task when diagnosed with the help of artificial intelligence. A sum of 618 CT images was used to process the result, which comes out to an accuracy of, specificity of, and sensitivity of about 99.6%, 92.2%, and 98.2% respectively. Ozturk et al. [82] proposed DarkNet model which was implemented with 17 convolutional layers to come up with accurate results. They succeeded in their task by attaining an accuracy of 98.08% in binary classification and 87.02% accuracy in multiclass classification.
Shah et al. [83] proposed CTnet-10 deep learning CNN–based model to classify CT scan images into COVID-19 and non-COVID-19. The CTnet-10 has 82.1% accuracy. They also observed that their model is faster compared to the RT-PCR method. They also verified DenseNet-169, VGG-16, ResNet-50, InceptionV3, and VGG-19. Among these, VGG-19 proved to be superior, having 94.52% accuracy. Javaheri et al. [84] have developed a model called CovidCTNet using deep learning algorithms to do a binary classification into either COVID-19 and community-acquired pneumonia (CAP) from CT scans. The accuracy of CovidCTNet was 95%. The important facts about CovidCTNet are that it was designed to work with small and heterogeneous sample sizes irrespective of CT scanning hardware and it was open source. Wide pattern and imaging feature resemblance of COVID-19 and CAP challenged the algorithm training but achieved accuracy of CovidCTNet make it a tool to be adapted for clinical decision. Wang et al. [85] assessed a deep learning algorithm using CT images for screening COVID-19 patients throughout the influenza season. To validate their hypothesis, they used 1065 CT images, out which 740 were COVID-19 negative and 325 were COVID-19 positive. Their algorithm delivered 89.5% accuracy, 0.88 specificity, and 0.87 sensitivity.
Their future work will focus on linking hierarchical features of CT images to features of other factors like genetic, epidemiological, and clinical information for the purpose of multi-modeling analysis. This multi-modeling analysis will expedite enhanced diagnosis. Saad et al. [86] have used deep feature concatenation (DFC) mechanism in two ways. In one-way DFC does the linking of deep features extracted from X-ray and CT images through a CNN. In second-way DFC combines extracted features either from X-ray or CT scan using CNN architecture along with two pre-trained CNNs called ResNet and GoogleNet. Their proposed architecture has 3 deep layers to mitigate large time consumption issues. Their first way has delivered 96.13% accuracy, 94.37% precision, 97.04% recall, and an f_score of 95.69%. Their second way has delivered an accuracy of 98.9%, 93.6% precision; a recall of 98.5% and 98.29% f_score when using CT images but when X-ray images used, this second way has got 99.3% accuracy, 99.79% precision, 98.8% recall, and f_score of 99.3%. Serte et al. [87] proposed an AI system to determine COVID-19 from images of a patients’ 3D CT volume. Their AI system employed Resnet-50 deep learning model in combination with majority voting to classify each 3D CT image into COVID-19 and normal CT image. Their AI system also used the ResNet-18 model together with majority voting to predict COVID-19 on a given patient’s 3D CT image. The created ResNet-50 system attained 0.90 area under curve (AUC) and 96% accuracy compared to 0.67 AUC of 3D-ResNet50. The major asset of their work was fine-tuning and majority voting-based modeling.
Singh et al. [89] developed an ensemble model for automated COVID-19 prediction by ensembling deep transfer learning models like ResNet152V2, VGG16, and densely connected convolutional networks (DCCNs). They have used chest CT scanned images for the development of their model. Their ensemble model can do a 4-class classification, whereas most previous models can only do a binary or 3-class classification. They compared their model with 15 other models and demonstrated that their model outdoes prevailing models with respect to f-measure, AUC, specificity, sensitivity, and accuracy of 1.3274%, 1.8372%, 1.8382%, 1.283%, and 1.2738% respectively. The developed ensemble model attained 99.2% accuracy on the training dataset. Kedia et al. [93] created CovNet-19, an ensemble deep convolutional neural network model using chest X-ray images to detect COVID-19. They performed a 3-class classification i.e. COVID-19, pneumonia, normal with an accuracy of 98.28%, 98.33% precision, 98.33% recall, 97.15% Matthews Correlation Coefficient (MCC) whereas accuracy of 99.71% and 99.26% MCC was delivered for binary class classification into Non-COVID-19 and COVID-19. F1 score was 99% for both 3-class and 2-class classification. Elgendi et al. [97] scrutinized 17 deep learning algorithms to figure out the impact of geometric augmentations for COVID-19 detection. Empirical analysis was done to measure the influence of augmentation with reference to accuracy, dataset variety, methodology of augmentation, and network size. Their results demonstrated that Matthews Correlation Coefficient (MCC) of all examined models improved after the removal of geometrical augmentation. They carried out this empirical analysis using MATLAB 2020a on a workstation having GPU NVIDIAGeForce RTX 2080Ti 11 GB, RAM 64 GB, and Intel ProcessorI9-9900 K @3.6 GHz. Ieracitano et al. [99] proposed a CAD system for differentiating portable CXR images of COVID-19 pneumonia patients from the Non-COVID-19 interstitial pneumonia patients in an accurate manner utilizing a local unbalanced dataset. This CAD system called CovNNet is a fuzzy enhanced deep learning-based framework. In this approach, CovNNet tends to extract the deep relevant features from the images which are the results of the combination of portable CXR images and fuzzy images. This CAD system achieved an encouraging accuracy of more than 80% over the local dataset. All these state-of-the-art approaches for the COVID-19 detection and classification based on the CT scans and chest X-ray images are summarized along with their respective future work with the help of Table 3:
Challenges and Limitations
The challenges and limitations in the deep learning-based COVID-19 detection or classification approaches utilizing CT scan and chest X-ray images are as follows:
-
1.
Regulation: During any pandemic like COVID-19, the concerned authorities have to take a crucial role in framing policies and etiquette-like lockdown, social distancing in case of COVID-19. These regulations and etiquette can stimulate scientists, researchers, citizens, technological companies, and social organizations to curtail obstacles to the prevent spread of COVID-19.
-
2.
Handiness of data: Application of DL in medical imaging requires large volumes of data for training of DL models. But in the case of COVID-19, availability of data is low. Also, checking candidness of data is difficult and requires expertise to interpret. So it will take some time to have befitting data to DL and thus to have widespread application of DL techniques in COVID-19 detection and classification.
-
3.
Data privacy concerns: Privacy concerns are biggest hurdle in collection of data like medical images that is required for applications of AI like DL and ML for COVID-19. Unavailability of sufficient data may result in less accurate and questionable DL models.
Deep Learning in Conjunction with Traditional Machine Learning Classifiers and Machine Learning-Based Approaches
The conventional machine learning-based approaches involves three sub-stages i.e. segmentation, feature extraction followed by the training of machine learning classifiers with the aid of these extracted features from the segmented region. Hence, proper manual selection of all methods employed in these sub-stages is very important. The present research trend in this domain involves usage of particular deep learning architectures, especially for performing the segmentation as well as deep feature extraction in a complete automated manner. This practice of employing the various deep learning networks alongside traditional machine learning classifiers is rendering encouraging results and can be termed as Deep learning in conjunction with traditional machine learning classifiers. Tang et al. [100] proposed a chest CT images model based on Ground-Glass Opacity (GGO) regions and Random Forest (RF) model to assess severity in terms of severe and non-severe on COVID-19 patients which also based on quantitative measures. Using three-fold cross validation, it shows a 93.3% true-positive rate, 74.5% true-negative rate, 87.5% accuracy, and 91% AUC. The major resulting thing in GGO shows that the right lung is more affected to severity than the left lung. Barstugan et al. [101] proposed early phase detection of COVID-19 utilizing abdominal CT images, which are acquired from the hospitals in the Zhejiang region of China, using machine learning methods. There are different materials used for the statistical features of data set, such as visual dataset in different subsets in terms of non-infected and infected. The classification is done by support vector machine (SVM). There are different results of five subsets. Their study achieved 99.68% accuracy in tenfold cross-validation. The machine learning methods should be done on CT abdominal images, X-ray chest images, and blood test results. Sethy et al. [102] proposed an approach using deep learning-based methodology, which gives benefit to practitioners that are researching on coronavirus patients. The models that are recommended are Resnet50 plus SVM, which achieved an accuracy of 95.38% for detecting COVID-19 positive patients. Their results were based on data which is available in the repository of GitHub, Kaggle, and Open-I as per their validated X-Ray images. Karawi et al. [103] proposed an approach based on machine learning techniques for analysis of chest CT scan images of COVID-19 patients. A frequency domain algorithm known as the Fast Fourier Transform (FFT)-Gabor scheme based on SVM model works in real-time and gets results with high accuracy along with low false-negative rate. This approach was trained on a dataset of 470 CT scan images in which 275 were positive cases and 195 were negative. Ozkaya et al. [104] proposed a hybrid model based on the SVM classifier for classification and DTL models like Resnet50, GoogleNet, and VGG-16 for deep feature extraction. Their proposed hybrid method shows high performance on both the datasets used in their research study. Alom et al. [105] proposed the multi-task deep learning model based on Inception Recurrent Residual Neural Network (IRRCNN) for COVID-19 classification and NABLA-N network models for infected lung region segmentation. These models were tested on X-ray, abdominal CT, and full body CT images. The results for X-Ray Images and CT Images had an accuracy of 86.67% and 98.78% respectively for COVID-19. Kumar et al. [106] proposed an intelligent system based on the ResNet152 DTL model for the deep feature extraction and machine learning classifiers like Logistic Regression (LR), k-Nearest Neighbour 26 (kNN 26), Decision Trees (DT), Random Forest (RF), Adaptive Boosting (AdaBoost), Naïve Bayes (NB), and XGBoost(XGB) for binary classification. Best results were delivered by the RF and XGBoost Predictive Classifiers. The above mentioned state-of-the-art approaches for the COVID-19 detection and classification are summarized along with their respective future work with the help of Table 4:
Challenges and Limitations
The challenges and limitations of machine learning and deep learning in conjunction with traditional machine learning classifiers approaches for the COVID-19 detection or classification utilizing CT scan and chest X-ray images are as follows:
-
The accuracy and robustness of most of the traditional machine learning-based approaches depends on utilizing the accurate segmentation method followed by the efficient feature extraction methods. Which makes the proper selection of segmentation and feature extraction methods very important and thus affects the overall proposed approach for the COVID-19 detection.
-
There is a need of properly annotated chest X-ray and CT scan datasets. These annotated datasets can be used to evaluate the segmentation accuracy and hence proved to be important for evaluating the performance of the proposed approach.
-
Most of the traditional machine learning-based approaches lack experimentation with various segmentation methods and feature extraction methods. Such experimentation is mandatory to be able to propose an efficient COVID-19 detection approach.
-
There is a scope for experimenting with various deep transfer learning models for performing the segmentation and deep feature extraction, as well as with various machine learning and ensemble learning classifiers for performing the classification of positive COVID-19 cases.
Majorly Used COVID-19 Chest X-Ray, CT Scan, and Ultrasound Image Dataset Description
In the present scenario, the propellant of modern computing, especially machine learning and deep learning, is training data. This training data is available in the form of datasets consisting of either medical images, histopathological images, biopsies images etc. All the deep learning and deep transfer learning-based approaches are totally depend on these training datasets. The COVID-19 detection approaches based on deep learning also require dataset of CT scan images, chest X-ray, statistics related to a country or a region, etc.; therefore, some of the majorly used open-source CT scan and chest X-ray dataset description are presented in Table 5.
Analysis and Evaluation
This section presents a brief analysis and evaluation of majorly used deep transfer learning (DTL) models like VGG16 [113], VGG19 [114], ResNet50 [115], and DenseNet [116] over the COVID-19 local CT scan dataset and global chest X-ray dataset. These four DTL models were initially fine-tuned and trained using the augmented local CT scan and chest X-ray dataset. The objective of this comparison is to illustrate how these commonly used DTL models perform on the local CT scan and global chest X-ray images COVID-19 datasets. The description of the two datasets used for the analysis is given below.
-
Local COVID-19 CT scan dataset: An axial volumetric chest CT scans of COVID-19 positive patients and normal people are present in this dataset. These volumetric CT scans were obtained utilizing the Optima GE CT 660 machine installed at the MP MRI and CT scan center Jabalpur, Madhya Pradesh, India, under the supervision of head radiologist. The 64-slice version of Optima GE CT 660 is available in this center, making it well-suited for cardiac and coronary angiography applications. This machine uses the Performix 40 tube (6.3 MHU) with a 40 mm V-Res detector. This Optima GE-CT660 acquires axial scans in sets of 2 through 64 contiguous images in one 360° rotation. For each rotation of the gantry, the Optima CT660 collects up to 64 rows of scan data. A total of 2080 CT scans were taken from 86 COVID-19 positive patients (mean age of 49.5 ± 19.1 years; range of 16–88 years, male 56, female 30) and 88 healthy people (mean age of 41.5 ± 16.8 years; range of 12–81 years, male 48, female 40). These cases were collected from July 2020 to January 2021. The main clinical symptoms in these patients were cough and fever. All the CT scan sequence are available in 16-bit grayscale DICOM format with 512*512 pixels resolution, which are converted into the PNG format.
-
Global chest X-ray dataset: As the available COVID-19 datasets are of very limited size, so in order to make a decent size balanced dataset, the chest X-ray images of COVID-19 positive patients and healthy people are taken from the three different publicly available datasets. Around 500 COVID-19 chest X-ray images and 500 normal images were taken from the GitHub repository by Dr. Joseph Cohen [96]. Then 220 COVID-19 positive images and 280 normal images were taken from the COVID-19 Radiography Database (COVID-19 Radiography Database 2020). Around 290 COVID-19 positive images and 280 normal images are taken from the IEEE8023/Covid Chest X-Ray Dataset [107]. This dataset consists of a total of 2070 chest X-ray images, which are further subdivided into the training and testing dataset.
All these CT scan and chest X-ray images are initially preprocessed and then augmented in order to create a large dataset for the training of these DTL models. As these medical images are obtained directly from diverse medical devices and may include artifacts and medical symbols, therefore, all these images are resized and cropped. The size of these CT scan and chest X-ray images is changed as per the input requirement of these DTL models. After pre-processing, an augmentation of the above two datasets are performed for the training of these DTL models in order to avoid the over-fitting. The augmentation strategies used in this section involve affine transformations composed of vertical and horizontal flip (0% ± 10%), scaling (0% ± 20%), shearing (0° ± 10°), and rotation (0° ± 10°).
The algorithm for the analysis and evaluation is as follows:
Input
COVID-19 CT scan and chest X-ray images or Normal CT scan or chest X-ray images.
Output
The trained VGG16, VGG 19, ResNet50, and DenseNet models for the detection of COVID-19 positive cases.
Steps
-
All the chest X-ray and CT scan images are preprocess for the elimination of artifacts, noise, and symbols.
-
Utilizing the affine transformations as an augmentation method consist of rotation (0° ± 10°), shearing (0° ± 10°), vertical and horizontal flip (0% ± 10%), and scaling (0% ± 20%) of both the two datasets are done.
-
Resize these CT scan and chest X-ray images to the size of 224-by-224-by 3 for the training of VGG16, VGG 19, DenseNet121, and ResNet50 DTL models.
-
The fine tuning and training of these four DTL models over the augmented datasets.
-
The VGG 16 and VGG19 models tends to converge at 100 epochs.
-
The DenseNet121 and ResNet50 tends to converge at 200 epochs.
-
Simulation and evaluation of these DTL models over the 20% augmented dataset which is reserved for the validation.
Initially, all the four DTL models are taken with similar settings, and then, in order to get the optimum performance from these DTL models, hyper parameter tuning is done during the training. The various combinations of the learning and dropout rate along with the two different optimizers were tried out in order to get the optimal configuration parameters of these four DTL models offering the best performance. The VGG16, VGG19, DenseNet-121, and ResNet50 DTL models’ optimal configuration parameters are illustrated with the help of Table 6 after performing a number of experiments. All the four deep transfer learning models were trained and evaluated with different learning and dropout rates along with the two different types of optimizers, i.e. Adam and Stochastic gradient descent (SGD) [117] for the weights adjustment. Both the VGG16 and 19 models give best performance with the Adam. Similarly, the DenseNet 121 and ResNet50 models with SGD optimizer tend to give better performance. The number of epochs required to converge also varies from model to model. As the VGG 16, VGG 19, and DenseNet121 tend to converge at 100 epochs, and after that, their accuracies are not at all improving. Similarly, the ResNet 50 tends to converges at 200 epochs. The dropout method [118] is utilized in order to avoid the problem of over-fitting.
The training and validation ratio for the two augmented datasets are 80:20, which means 20% is used for validation and rest 80% for training. The performance of these four DTL models is illustrated with the aid of Tables 7 and 8 using the statistical parameters like accuracy, sensitivity, specificity, precision, and F1 score. The training and validation graphs, as well as the Receiver Operating Characteristic Curve (ROC) of all the four DTL models on the Local CT scan dataset, are presented with help of Figs. 8 and 9, whereas the training and validation graphs and the ROC of all the four DTL models on the global chest X-ray dataset are presented with the help of Figs. 10 and 11.
Now the computational and architectural complexity of these four VGG16, VGG19, DenseNet121, and ResNet50 models along with the average accuracy are also compared with the aid of Table 9. The architectural complexity is normally measured in terms of the number of learnable parameters, whereas the architectural complexity of these four models is expressed in terms of FLOPs (floating-point operations per second).
The classification performance of VGG19 deep transfer learning model on both the augmented datasets of COVID-19 is better in comparison to DenseNet121, VGG16, and ResNet50 DTL models. But considering the computational as well as architectural complexities, it is the VGG16 model offering the optimum computational and architectural complexities with decent classification performance as well.
As VGG16 delivers the best classification performance over both the chest X-ray and CT scan datasets. The Gradient weighted Class Activation Mapping (Grad-CAM) explainability technique [119] is used in order to visually interpret as well as to demonstrate the effectiveness of this DTL model. This Grad-CAM is applied to the last convolutional layer of our VGG19 model in order to verify and explain the output result delivered by the VGG19 as COVID-19 and normal case. Some of the CT scan and chest X-ray test case output results delivered by the VGG19 along with Grad-CAM are presented with the help of Fig. 12.
Conclusion and Future Work
This methodical review presented a comprehensive analysis of the state-of-the-art deep and machine learning-based approaches for COVID-19 detection. A decent number of CT scan and chest X-ray datasets, which are available post-March 2020 were presented also in this study. Recent deep learning approaches utilizing chest X-ray images, CT scans, and ultrasound images certainly offer a low cost, rapid, automatic approach and do not require physical contact by medical staff for COVID-19 detection. This study discussed challenges and limitations also. Recently, some COVID-19 ultrasound scan datasets also became available for research. A good number of deep learning architectures are still left to be trained and tested on these datasets, which might offer more accurate results. Hence, further research studies should be conducted with objectives to test deep learning models on large size datasets of chest X-ray images, CT scans, and ultrasound images, and validate the results with radiologist’s observations. This is obligatory to propose an acceptable real-time application for automatic detection of COVID-19 using imaging modalities. Deep learning approaches to detect COVID-19 could be improved, if more clinical information can be collected from images comprising multiple disease symptoms. Currently, major deep learning approaches focus only on the posterior-anterior (PA) view of X-rays. Hence, it cannot differentiate other views of X-rays such as anterior–posterior (AP), lateral, etc. Further research studies can consider these factors. Future deep learning models must seek to distinguish COVID-19 cases from other similar viral cases, e.g. SARS, MERS, and from varieties of common pneumonia.
Future research studies can primarily focus on development of hybrid models utilizing deep learning architectures for segmentation and feature extraction purpose along with machine learning classifiers for binary or multiclass COVID-19 classification, since these hybrid models does not require large size dataset. Deep learning approaches lack transparency and interpretability since it is impossible to determine which imaging features are being considered to determine the output. Even the heat-map that is used to visualize the essential regions in the scans cannot determine which unique features are used to establish the output. A substantial overlap exists between how the lung reacts to various offends and appearance of diseases in the lung that depend on host factors, e.g. age, drug reactivity, immune status, underlying comorbidities. Hence, multidisciplinary models may be required because no single method can differentiate all lung diseases form imaging appearance on chest X-rays and chest CT scans. Future deep learning models must also consider determining severity degree of COVID-19 besides detecting it in order to monitor and treat patients effectively.
Availability of Data and Material
CT scan dataset can be provided on request, whereas the other chest X-ray dataset is publically available.
Code Availability
The code will be provided on request.
References
Paules CI, Marston HD, Fauci AS. Coronavirus infections—more than just the common cold. JAMA. 2020;323(8):707–8.
Nishiura H, Linton NM, Akhmetzhanov AR. Serial interval of novel coronavirus (COVID-19) infections. Int J Infect Dis. 2020. https://doi.org/10.1016/j.ijid.2020.02.060.
Perlman S, Netland J. Coronaviruses post-SARS: update on replication and pathogenesis. Nat Rev Microbiol. 2009;7(6):439–50.
Chan JF, To KK, Tse H, Jin DY, Yuen KY. Interspecies transmission and emergence of novel viruses: lessons from bats and birds. Trends Microbiol. 2013;21(10):544–55.
Chen Y, Liu Q, Guo D. Emerging coronaviruses: genome structure, replication, and pathogenesis. J Med Virol. 2020;92(4):418–23.
Stoecklin SB, Rolland P, Silue Y, Mailles A, Campese C, Simondon A, et al. First cases of coronavirus disease 2019 (COVID-19) in France: surveillance, investigations and control measures. Eurosurveillance. 2020;25(6):2000094.
Zhang N, Wang L, Deng K, Liang R, Su M, He C, Hu L, Su Y, Ren J, Yu F, Du L, Jiang S. Recent advances in the detection of respiratory virus infection in humans. J Med Virol. 2020. https://doi.org/10.1002/jmv.25674.
Xu Z, Shi L, Wang Y, Zhang J, Huang L, Zhang C, Liu S, Zhao P, Liu H, Zhu L, Tai Y, Bai C, Gao T, Song J, Xia P, Dong J, Zhao J, Wang FS. Pathological findings of COVID-19 associated with acute respiratory distress syndrome. Lancet Respir Med. 2020;8(4):420–2. https://doi.org/10.1016/S2213-2600(20)30076-X.
Muhammad LJ, Islam MM, Usman SS, Ayon SI. Predictive data mining models for novel coronavirus (COVID-19) infected patients’ recovery. SN Comput Sci. 2020;1(4):206. https://doi.org/10.1007/s42979-020-00216-w.
Corman VM, Landt O, Kaiser M, Molenkamp R, Meijer A, Chu DK, Bleicker T, Brünink S, Schneider J, Schmidt ML, Mulders DG, Haagmans BL, van der Veer B, van den Brink S, Wijsman L, Goderski G, Romette JL, Ellis J, Zambon M, Peiris M, Goossens H, Reusken C, Koopmans M, Drosten C. Detection of 2019 novel coronavirus (2019-nCoV) by real-time RT-PCR. Euro surveillance: bulletin Europeen sur les maladies transmissibles European communicable disease bulletin. 2020;25(3):2000045. https://doi.org/10.2807/1560-7917.ES.2020.25.3.2000045.
Lan L, Xu D, Ye G, Xia C, Wang S, Li Y, Xu H. Positive RT-PCR test results in patients recovered from COVID-19. JAMA. 2020;323(15):1502–3. https://doi.org/10.1001/jama.2020.2783.
Wang W, Xu Y, Gao R, Lu R, Han K, Wu G, Tan W. Detection of SARSCoV-2 in different types of clinical specimens. JAMA. 2020. https://doi.org/10.1001/jama.2020.3786.
Movassagh AA, Alzubi JA, Gheisari M, Rahimi M, Mohan S, Abbasi AA, Nabipour N. Artificial neural networks training algorithm integrating invasive weed optimization with differential evolutionary model. J Ambient Intell Humaniz Comput. 2021;1–9. https://doi.org/10.1007/s12652-020-02623-6.
ALzubi JA, Bharathikannan B, Tanwar S, Manikandan R, Khanna A, Thaventhiran C. Boosted neural network ensemble classification for lung cancer disease diagnosis. Appl Soft Comput. 2019;80:579–591. https://doi.org/10.1016/j.asoc.2019.04.031.
Li Y, Xia L. Coronavirus disease 2019 (COVID-19): role of chest CT in diagnosis and management. AJR. 2020;214:1280–6. https://doi.org/10.2214/AJR.20.22954.
Fang Y, Zhang H, Xie J, Lin M, Ying L, Pang P, Ji W. Sensitivity of chest CT for COVID-19: comparison to RT-PCR. Radiology. 2020. https://doi.org/10.1148/radiol.2020200432.
Self WH, Courtney DM, McNaughton CD, Wunderink RG, Kline JA. High discordance of chest x-ray and computed tomography for detection of pulmonary opacities in ED patients: implications for diagnosing pneumonia. Am J Emerg Med. 2013;31(2):401–405. https://doi.org/10.1016/j.ajem.2012.08.041.
Yang W, Sirajuddin A, Zhang X, Liu G, Teng Z, Zhao S, Lu M. The role of imaging in 2019 novel coronavirus pneumonia (COVID-19). Eur Radiol. 2020;30(9):4874–82. https://doi.org/10.1007/s00330-020-06827-4.
Chung M, Bernheim A, Mei X, Zhang N, Huang M, Zeng X, Cui J, Xu W, Yang Y, Fayad ZA, Jacobi A, Li K, Li S, Shan H. CT Imaging features of 2019 novel coronavirus (2019-nCoV). Radiology. 2020;295(1):202–207. https://doi.org/10.1148/radiol.2020200230.
Huang C, Wang Y, Li X, Ren L, Zhao J, Hu Y, Zhang L, Fan G, Xu J, Gu X, Cheng Z, Yu T, Xia J, Wei Y, Wu W, Xie X, Yin W, Li H, Liu M, Xiao Y, et al. Clinical features of patients infected with 2019 novel coronavirus in Wuhan, China. Lancet, London, England. 2020;395(10223):497–506. https://doi.org/10.1016/S0140-6736(20)30183-5.
Kundu S, Elhalawani H, Gichoya JW, Kahn CE How might AI and chest imaging help unravel COVID-19’s mysteries?. Radiol Artif Intell. 2020;2(3). https://doi.org/10.1148/ryai.2020200053
Thevenot J, Lopez MB, Hadid A. A survey on computer vision for assistive medical diagnosis from faces. IEEE J Biomed Health Inform. 2020;22(5):1497–511. https://doi.org/10.1109/JBHI.2017.2754861.
Islam MM, Iqbal H, Haque MR, Hasan MK. Prediction of breast cancer using support vector machine and K-Nearest neighbors. IEEE Region 10 Humanitarian Technology Conference. Dhaka, Bangladesh. 2017. https://doi.org/10.1109/R10-HTC.2017.8288944.
Haque MR, Islam MM, Iqbal H, Reza MS, Hasan MK. Performance evaluation of random forests and artificial neural networks for the classification of liver disorder. Int Conf Comput Commun Chem Mater Electr Eng (IC4ME2). Rajshahi, Bangladesh. 2018. https://doi.org/10.1109/IC4ME2.2018.8465658.
Hasan MK, Islam MM, Hashem MMA. Mathematical model development to detect breast cancer using multigene genetic programming. 5th Int Conf Inform Electr Vision (ICIEV). Dhaka, Bangladesh. 2016. https://doi.org/10.1109/ICIEV.2016.7760068.
Islam AS, Islam MM. Diabetes prediction: a deep learning approach. Int J Inform Eng Electr Business. 2019;11(2):21–7. https://doi.org/10.5815/ijieeb.2019.02.03.
Ayon SI, Islam MM, Hossain MR. Coronary artery heart disease prediction: a comparative study of computational intelligence techniques. IETE J Res. 2020. https://doi.org/10.1080/03772063.2020.1713916.
Rahaman A, Islam MM, Islam MR, Sadi MS, Nooruddin S. Developing IoT based smart health monitoring systems: a review. Rev Intell Artif. 2019;33:435–40. https://doi.org/10.18280/ria.330605.
Islam MM, Rahaman A, Islam MR. Development of smart healthcare monitoring system in IoT environment. SN Comput Sci. 2020;1:185. https://doi.org/10.1007/s42979-020-00195-y.
Jiang X. Feature extraction for image recognition and computer vision. Proc 2nd IEEE Int Conf Comput Sci Inf Techno ICCSIT. 2009. https://doi.org/10.1109/ICCSIT.2009.5235014.
Kim TK, Yi PH, Hager GD, Lin CT. Refining dataset curation methods for deep learning-based automated tuberculosis screening. J Thoracic Dis. 2020;12(9):5078–5085. https://doi.org/10.21037/jtd.2019.08.34.
Wong HYF, Lam HYS, Fong AH, Leung ST, Chin TW, Lo CSY, Lui MM, Lee JCY, Chiu KW, Chung TW, Lee EYP, Wan EYF, Hung IFN, Lam TPW, Kuo MD, Ng MY. Frequency and distribution of chest radiographic findings in patients positive for COVID-19. Radiology. 2020;296(2):E72–8. https://doi.org/10.1148/radiol.2020201160.
Zech JR, Badgeley MA, Liu M, Costa AB, Titano JJ, Oermann EK. Variable generalization performance of a deep learning model to detect pneumonia in chest radiographs: a cross-sectional study. PLOS MED. 2018. https://doi.org/10.1371/journal.pmed.1002683.
Hurt B, Kligerman S, Hsiao A. Deep learning localization of pneumonia: 2019 coronavirus (COVID-19) outbreak. J Thorac Imaging. 2020;35(3):W87–9. https://doi.org/10.1097/RTI.0000000000000512.
Yi PH, Kim TK, Lin CT. Generalizability of deep learning tuberculosis classifier to COVID-19 chest radiographs: new tricks for an old algorithm? J Thorac Imaging. 2020;35(4):W102–4. https://doi.org/10.1097/RTI.0000000000000532.
Xia C, Li X, Wang X, Kong B, Chen Y, Yin Y, Cao K, Song Q, Lyu S, Wu X. A multi-modality network for cardiomyopathy death risk prediction with CMR images and clinical information. Med Image Comput Comput Assist Interv. 2019:577–585. https://doi.org/10.1007/978-3-030-32245-8_64.
Kong B, Wang X, Junjie B, Lu Y, Gao F, Kunlin C, Xia J, Song Q, Yin Y. Learning tree-structured representation for 3D coronary artery segmentation. Comput Med Imaging Graph. 2019. https://doi.org/10.1016/j.compmedimag.2019.101688.
Ye H, Gao F, Yin Y, Guo D, Zhao P, Lu Y, Wang X, Bai J, Cao K, Song Q, Zhang H, Chen W, Guo X, Xia J. Precise diagnosis of intracranial hemorrhage and subtypes using a three-dimensional joint convolutional and recurrent neural network. Eur Radiol. 2019;29(11):6191–201. https://doi.org/10.1007/s00330-019-06163-2.
Kermany DS, Goldbaum M, Cai W, Valentim C, Liang H, Baxter SL, McKeown A, Yang G, Wu X, Yan F, Dong J, Prasadha MK, Pei J, Ting M, Zhu J, Li C, Hewett S, Dong J, Ziyar I, Shi A, et al. Identifying medical diagnoses and treatable diseases by image-based deep learning. Cell. 2018;172(5):1122-1131.e9. https://doi.org/10.1016/j.cell.2018.02.010.
Rajaraman S, Candemir S, Kim I, Thoma G, Antani S. Visualization and interpretation of convolutional neural network predictions in detecting pneumonia in pediatric chest radiographs. Applied sciences, Basel Switzerland. 2018;8(10):1715. https://doi.org/10.3390/app8101715.
Depeursinge A, Chin AS, Leung AN, Terrone D, Bristow M, Rosen G, Rubin DL. Automated classification of usual interstitial pneumonia using regional volumetric texture analysis in high-resolution computed tomography. Invest Radiol. 2015;50(4):261–7. https://doi.org/10.1097/RLI.0000000000000127.
Anthimopoulos M, Christodoulidis S, Ebner L, Christe A, Mougiakakou S. Lung pattern classification for interstitial lung diseases using a deep convolutional neural network. IEEE Trans Med Imaging. 2016;35(5):1207–16. https://doi.org/10.1109/TMI.2016.2535865.
Bhandare A, Bhide M, Gokhale P, Chandavarkar R. Applications of convolutional neural networks. Int J Comput Sci Inform Technol. 2016;7:2206–15.
Suzuki K. Overview of deep learning in medical imaging. Radiol Phys Technol. 2017;10(3):257–73. https://doi.org/10.1007/s12194-017-0406-5.
Banihabib ME, Bandari R, Valipour M. Improving daily peak flow forecasts using hybrid Fourier-series autoregressive integrated moving average and recurrent artificial neural network models. AI. 2020;1(2):263–275. https://doi.org/10.3390/ai1020017.
Ucar F, Korkmaz D. Covidiagnosis-net: deep bayes-squeezenet based diagnostic of the coronavirus disease. (covid-19) from x-ray images. Medi Hypothesis J. 2019;2020.
Hammoudi K, Benhabiles H, Melkemi M, Dornaika F, Arganda-Carreras I, Collard D, Scherpereel A. Deep learning on chest X-ray images to detect and evaluate pneumonia cases at the era of COVID-19. 2020. https://arxiv.org/pdf/2004.03399.pdf.
Rajaraman S, Siegelman J, Alderson PO, Folio LS, Folio LR, Antani SK. Iteratively pruned deep learning ensembles for COVID-19 detection in chest X-rays. IEEE Access. 2020;8:115041–50. https://doi.org/10.1109/ACCESS.2020.3003810.
Hall LO, Paul R, Goldgof DB, Goldgof GM. Finding COVID-19 from chest X-rays using deep learning on a small dataset. 2020. https://arxiv.org/ftp/arxiv/papers/2004/2004.02060.pdf.
Rahimzadeh M, Atta A. A new modified deep convolutional neural network for detecting Covid-19 from X-ray images. 2020. https://arxiv.org/pdf/2004.08052.pdf.
Zhang J, Xie Y, Li Y, Shen C, Xia Y. COVID-19 screening on chest X-ray images using deep learning based anomaly detection. 2020. https://arxiv.org/pdf/2003.12338.pdf.
Hemdan EE, Shouman MA and Karar ME. COVIDX-Net: a framework of deep learning classifiers to diagnose COVID-19 in X-ray images. 2020. https://arxiv.org/ftp/arxiv/papers/2003/2003.11055.pdf.
Wang S, Zha Y, Li W, Wu Q, Li X, Niu M, Wang M, Qiu X, Li H, Yu H, Gong W, Bai Y, Li L, Zhu Y, Wang L, Tian J. A Fully automatic deep learning system for COVID-19 diagnostic and prognostic analysis. Eur Respir J. 2020;56:2000775. https://doi.org/10.1183/13993003.00775-2020.
Zheng C, Deng X, Fu Q, Zhou Q. Deep learning-based detection for COVID-19 from chest CT using weak label. IEEE Trans Med Imaging. 2020. https://doi.org/10.1109/TMI.2020.2995965.
Apostolopoulos ID, Sokratis AI, Mpesiana TA. Extracting possibly representative COVID-19 biomarkers from X-ray images with deep learning approach and image data related to pulmonary diseases. Journal of Medical and Biological Engineering. 2020;40:462–9. https://doi.org/10.1007/s40846-020-00529-4.
Fu M, Yi SL, Zeng Y, Ye F, Li Y, Dong X, Ren YD, Luo L, Pan JS, Zhang Q. Deep learning based recognizing COVID-19 and other common infectious diseases of the lung by chest CT scan images. 2020. https://doi.org/10.1101/2020.03.28.20046045.
Ardakani AB, Kanafi AR, Acharya UR, Khadem N, Mohammadi A. Application of deep learning technique to manage COVID-19 in routine clinical practice using CT images: results of 10 convolutional neural networks. Comput Biol Med. 2020;121: 103795. https://doi.org/10.1016/j.compbiomed.2020.103795.
Rehman A, Naz S, Khan A, Zaib A, Razzak I. Improving corona virus (Covid-19) diagnosis using deep transfer learning. medRxiv. 2020. https://doi.org/10.1101/2020.04.11.20054643.
Khalifa NEM, Taha Aboul MHN, Hassanien E, Elghamrawy S. Detection of coronavirus (COVID-19) associated pneumonia based on generative adversarial networks and a fine-tuned deep transfer learning model using chest X-ray dataset. 2022. arXiv:2004.01184.
Loey M, Smarandache F, Khalifa NEM. Within the lack of chest COVID-19 X-ray dataset: a novel detection model based on GAN and deep transfer learning. Symmetry. 2020;12(4):651. https://doi.org/10.3390/sym12040651.
Shan F, Gao Y, Wang J, Shi W, Shi N, Han M, Xue Z, Shen D, Shi Y. Lung infection quantification of COVID-19 in CT images with deep learning. 2020. https://arxiv.org/abs/2003.04655.
Hu S, Gao Y, Niu Z, Jiang Y, Li L, Xiao X, Wang M, Fang EF, Smith WM, Xia J, Ye H, Yang G. Weakly supervised deep learning for COVID-19 infection detection and classification from CT images. IEEE Access. 2020;8:118869–83. https://doi.org/10.1109/ACCESS.2020.3005510.
Wang L, Lin ZQ, Wong A. COVID-Net: a tailored deep convolutional neural network design for detection of COVID-19 cases from chest X-ray images. Scientific Reports, Nature. 2020;10:19549. https://doi.org/10.1038/s41598-020-76550-z.
Li L, Qin L, Xu Z, Yin Y, Wang X, Kong B, Bai J, Lu Y, Fang Z, Song Q, Cao K, Liu D, Wang G, Xu Q, Fang X, Zhang S, Xia J, Xia J. Using artificial intelligence to detect COVID-19 and community-acquired pneumonia based on pulmonary CT: evaluation of the diagnostic accuracy. Radiology. 2020;296(2):E65–71. https://doi.org/10.1148/radiol.2020200905.
Minaee S, Kafieh R, Sonka M, Yazdani S, Soufi GJ. Deep-COVID: predicting COVID-19 from chest X-ray images using deep transfer learning. Med Image Anal. 2020;101794. https://doi.org/10.1016/j.media.2020.101794.
Basu S, Mitra S, Shah N. Deep learning for screening COVID-19 using chest X-ray images. 2020. https://arxiv.org/pdf/2004.10507.pdf.
Khalifa NEM, Smarandache F, Manogaran G, Loey M. A study of the neutrosophic set significance on deep transfer learning models: an experimental case on a limited COVID-19 chest X-ray dataset. Cognitive Computing. 2021. https://doi.org/10.1007/s12559-020-09802-9.
Cohen JP, Dao L, Roth K, Morrison P, Bengio Y, Abbasi AF, Shen B, Mahsa HK, Ghassemi M, Li H, Duong T. Predicting COVID-19 pneumonia severity on chest X-ray with deep learning. Cureus. 2020;12(7):e9448. https://doi.org/10.7759/cureus.9448.
Ying S, Zheng S, Li L, Zhang X, Zhang X, Huang Z, Chen J, Zhao H, Wang R, Chong Y, Shen J, Zha Y, Yang Y. Deep learning enables accurate diagnosis of novel coronavirus (COVID-19) with CT images. MedRxiv. 2020. https://doi.org/10.1101/2020.02.23.20026930.
Wang S, Kang B, Ma J, Zeng X, Xiao M, Guo J, Cai M, Yang J, Li Y, Meng X, Xu B. A deep learning algorithm using CT images to screen for Corona Virus Disease (COVID-19). medRxiv preprint. 2020. https://doi.org/10.1101/2020.02.14.20023028.
Narin A, Ceren K, Ziynet P. Automatic detection of coronavirus disease (COVID-19) using X-ray images and deep convolutional neural networks. 2020. arXiv:2003.10849v3
Jin C, Chen W, Cao Y, Xu Z, Tan Z, Zhang X, Deng L, Zheng C, Zhou J, Shi H, Feng J. Development and evaluation of an AI system for COVID-19 diagnosis. Nat Commun. 2020;11(1):5088. https://doi.org/10.1038/s41467-020-18685-1.
Xu X, Jiang X, Ma C, Du P, Li X, Lv S, Yu L, Chen Y, Su J, Lang G, Li Y, Zhao H, Xu K, Ruan L Wu W. Deep learning system to screen coronavirus disease 2019 pneumonia. Engineering. 202010(6):1122–1129. https://doi.org/10.1016/j.eng.2020.04.010.
Huang L, Han R, Ai T, Yu P, Kang H, Tao Q, Xia L. Serial quantitative chest CT assessment of COVID-19: deep-learning approach. radiology: cardiothoracic imaging 2020; 2(2):e200075. https://doi.org/10.1148/ryct.2020200075.
Farooq M, Hafeez A. COVID-ResNet: A deep learning framework for screening of COVID19 from radiographs. 2020. arXiv:2003.14395v1.
Chen J, Wu L, Zhang J, Zhang L, Gong D, Zhao Y, Chen Q, Huang S, Yang M, Yang X, Hu S, Wang Y, Hu X, Zheng B, Zhang K, Wu H, Dong Z, Xu Y, Zhu Y, Chen X, Zhang M, Yu L, Cheng F, Yu H. Deep learning-based model for detecting 2019 novel coronavirus pneumonia on high-resolution computed tomography. Sci Rep. 2020. 5;10(1):19196. https://doi.org/10.1038/s41598-020-76282-0.
Asnaoui KEL, Chawki Y, Idri A. Automated methods for detection and classification pneumonia based on X-ray images using deep learning. 2020. arXiv:2003.14363v1.
Chowdhury MEH, Rahman T, Khandakar A, Mazhar R, Kadir MA, Mahbub ZB, Islam KR, Khan MS, Iqbal A, Al-Emadi N, Reaz MBI, Islam MT. Can AI help in screening viral and COVID-19 pneumonia? IEEE Access. 2020;8:132665–76. https://doi.org/10.1109/ACCESS.2020.3010287.
Apostolopouos ID, Mpesiana TA. Covid-19: automatic detection from X-ray images utilizing transfer learning with convolutional neural networks. Phys Eng Sci Med. 2020;43:635–40. https://doi.org/10.1007/s13246-020-00865-4.
Afshar P, Heidarian S, Naderkhani F, Oikonomou A, Plataniotis KN, Mohammadi A. COVID-CAPS: a capsule network-based framework for identification of COVID-19 cases from X-ray images. Pattern Recogn Lett. 2020;138:638–43. https://doi.org/10.1016/j.patrec.2020.09.010.
Butt C, Gill J, Chun D, Babu BA, Deep learning system to screen coronavirus disease,. pneumonia. Appl Intell. 2019;2020:1–7. https://doi.org/10.1007/s10489-020-01714-3.
Ozturk T, Talo M, Yildirim EA, Baloglu UB, Yildirim O, Rajendra AU. Automated detection of COVID-19 cases using deep neural networks with X-ray images. Comput Biol Med. 2020;121: 103792. https://doi.org/10.1016/j.compbiomed.2020.103792.
Shah V, Keniya R, Shridharani A, Pubjabi M, Shah J. Diagnosis of COVID-19 using CT scan images and deep learning techniques. Emerg Radiol. 2021. https://doi.org/10.1007/s10140-020-01886-y.
Javaheri T, Homayounfar M, Amoozgar Z, Reiazi R, Homayounieh F, Abbas E, Laali A, Radmard AR, Gharib MH, Mousavi S, Ghaemi O, Babaei R, Mobin HK, Hosseinzadeh M, Jahanban-Esfahlan R, Seidi K, Kalra MK, Zhang G, Chitkushev LT, Haibe-Kains B, Malekzadeh R, Rawassizadeh R. CovidCTNet: an open-source deep learning approach to diagnose covid-19 using small cohort of CT images. NPJ digital medicine. 2021;4(1):29. https://doi.org/10.1038/s41746-021-00399-3.
Wang S, Kang B, Ma J, Zeng X, Xiao M, Guo J, Cai M, Yang J, Li Y, Meng X, Xu B. A deep learning algorithm using CT images to screen for corona virus disease (COVID-19). Eur Radiol. 2021. https://doi.org/10.1007/s00330-021-07715-1.
Saad W, Shalaby WA, Shokair M, El-Samie FA, Dessouky M, Abdellatef E. COVID-19 classification using deep feature concatenation technique. J Ambient Intell Humaniz Comput. 2021. https://doi.org/10.1007/s12652-021-02967-7.
Serte S, Demirel H. Deep learning for diagnosis of COVID-19 using 3D CT scans. Comput Biol Med. 2021. https://doi.org/10.1016/j.compbiomed.2021.104306.
Morozov SP, Andreychenko AE, Pavlov NA, Vladzymyrskyy AV, Ledikhova NV, Gombolevskiy VA, Blokhin IA,Gelezhe PB, Gonchar AV, Chernina VY. Mosmeddata: chest CT scans with covid-19 related findings dataset. 2020. https://mosmed.ai/datasets/covid19_1110.
Singh D, Kumar V, Kaur M. Densely connected convolutional networks-based COVID-19 screening model. Appl Intell. 2021. https://doi.org/10.1007/s10489-020-02149-6.
Pathak Y, Shukla PK, Arya KV. Deep bidirectional classification model for covid-19 disease infected patients. IEEE/ACM Trans Comput Biol Bioinf. 2020. https://doi.org/10.1109/TCBB.2020.3009859.
Singh D, Kumar V, Vaishali, Kaur M. Classification of COVID-19 patients from chest CT images using multi-objective differential evolution-based convolutional neural networks. European Journal of Clinical Microbiology & Infectious Diseases: Official Publication of the European Society of Clinical Microbiology. 2021;39(7):1379–1389. https://doi.org/10.1007/s10096-020-03901-z.
Li L, Huang H, Jin X. AE-CNN classification of pulmonary tuberculosis based on CT images. 9th International Conference on Information Technology in Medicine and Education (ITME), Hangzhou, China. 2018;39–42. https://doi.org/10.1109/ITME.2018.00020.
Kedia P, Anjum, Katarya R. CoVNet-19: a deep learning model for the detection and analysis of COVID-19 patients. Appl Soft Comput. 2021. https://doi.org/10.1016/j.asoc.2021.107184.
Rahman T, Chowdhury M, Khandakar A. Covid19-radiography-database. 2020. https://www.kaggle.com/tawsifurrahman/covid19-radiography-database.
Chung AG. COVID-19 chest X-ray. 2020. https://github.com/agchung.
Larxel. COVID-19 X rays. 2020. https://www.kaggle.com/andrewmvd/convid19-x-rays.
Elgendi M, Nasir MN, Tang Q, Smith D, Grenier JP, Batte C, Spieler B, Leslie WD, Menon C, Fletcher RR, Howard N, Ward R, Parker W, Nicolaou S. The effectiveness of image augmentation in deep learning networks for detecting COVID-19: a geometric transformation perspective. Front Med. 2021. https://doi.org/10.3389/fmed.2021.629134.
Praveen. CoronaHack-Chest X-Ray-Dataset. 2020. https://www.kaggle.com/praveengovi/coronahack-chest-xraydataset.
Ieracitano C, Mammone N, Versaci M, Varone G, Ali AR, Armentano A, Calabrese G, Ferrarelli A, Turano L, Tebala C, Hussain Z, Sheikh Z, Sheikh A, Sceni G, Hussain A, Morabito FC. A fuzzy-enhanced deep learning approach for early detection of Covid-19 pneumonia from portable chest X-ray images. Neurocomputing. 2022;481:202–15. https://doi.org/10.1016/j.neucom.2022.01.055.
Tang Z, Zhao W, Xie X, Zhong Z, Shi F, Liu J, Shen D. Severity assessment of coronavirus disease 2019 (COVID-19) using quantitative features from chest CT images. 2020. https://arxiv.org/abs/2003.11988v1.
Barstugan M , Ozkaya U, Ozturk S. Coronavirus (COVID-19) Classification using CT images by machine learning methods. 2020. https://arxiv.org/abs/2003.09424.
Sethy, PK, Behera, SK. Detection of coronavirus disease (COVID-19) based on deep features. Preprints. 2020;2020030300. https://doi.org/10.20944/preprints202003.0300.v1.
Al-Karawi D, Al-Zaidi S, Polus N, Jassim S. Machine learning analysis of chest CT scan images as a complementary digital test of coronavirus (COVID-19) patients. 2020. https://doi.org/10.1101/2020.04.13.20063479v1.
Ozkaya U, Ozturk S, Barstugan M. Coronavirus (COVID-19) Classification using deep features fusion and ranking technique. 2020. https://arxiv.org/ftp/arxiv/papers/2004/2004.03698.pdf.
Alom MZ, Shaifur Rahman MM, Nasrin MS, Taha TM, Asari VK. COVID_MTNet: COVID-19 detection with multi-task deep learning approaches. 2020. https://arxiv.org/ftp/arxiv/papers/2004/2004.03747.pdf.
Kumar R, Arora R, Bansal V, Sahayasheela VJ, Buckchash H, Imran J, Narayanan N, Pandian GN, Raman B. Accurate prediction of COVID-19 using chest X-ray images through deep feature learning model with SMOTE and machine learning classifiers. MedRxiv. 2020. https://doi.org/10.1101/2020.04.13.20063461v1.full.pdf.
Cohen JP. COVID-19 image data collection. 2020. https://github.com/ieee8023/covid-chestxray-dataset.
Zhao J, Zhang Y, He X, Xie P. COVID-CT-Dataset: a CT scan dataset about COVID-19. 2020. arXiv:2003.13865.
Jun M, Cheng G, Yixin W, Xingle A, Jiantao G, Ziqi Y, Minqing Z, Xin L, Xueyuan D, Shucheng C, Hao W, Sen M, Xiaoyu Y, Ziwei N, Chen L, Lu T, Yuntao Z, Qiongjie Z, Guoqiang D, Jian H. COVID-19 CT lung and infection segmentation dataset. Zenodo. 2020. https://doi.org/10.5281/zenodo.3757476.
Gunraj H, Wang L, Wong A. COVIDNet-CT: a tailored deep convolutional neural network design for detection of COVID-19 cases from chest CT images. Front Med. 2020;7: 608525. https://doi.org/10.3389/fmed.2020.608525.
Afshar P, Heidarian S, Enshaei N, Naderkhani F, Rafiee MJ, Oikonomou A, Fard FB, Samimi K, Plataniotis KN, Mohammadi A. COVID-CT-MD: COVID-19 computed tomography (CT) scan dataset applicable in machine learning and deep learning. 2020. arXiv:2009.14623.
El-Shafai, W, Abd El-Samie F. Extensive COVID-19 X-ray and ct chest images dataset. Mendeley Data, V3. 2020. https://doi.org/10.17632/8h65ywd2jr.3.
Simonyan K, Zisserman A. Very deep convolutional networks for large scale image recognition. arXiv. 2015. https://arxiv.org/abs/1409.1556.
Liu T, Wang Z. HiCNN: a very deep convolutional neural network to better enhance the resolution of Hi-C data. Bioinformatics. 2019;35(21):4222–8. https://doi.org/10.1093/bioinformatics/btz251.
He K, Zhang X, Ren S, Sun J. Deep residual learning for image recognition. In CVPR. 2016;770–778. https://arxiv.org/abs/1512.03385.
Huang G, Liu Z, Van Der Maaten L, Weinberger KQ. Densely connected convolutional networks. In Proceedings of IEEE Conference Computer Vision Pattern Recognition (CVPR). 2017;22612269.
Kingma DP. Adam: a method for stochastic optimization. 2015. http://arxiv.org/abs/1412.6980.
Srivastava N, Hinton G, Krizhevsky A, Sutskever I, Salakhutdinov R. Dropout: a simple way to prevent neural networks from over fitting. J Mach Learn Res. 2014;15(56):1929–58.
Selvaraju RR, Cogswell M, Das A, Vedantam R, Parikh D, Batra D. Grad-CAM: visual explanations from deep networks via gradient-based localization. Int J Comput Vision. 2020;128:336–59. https://doi.org/10.1007/s11263-019-01228-7.
Acknowledgements
We would like to thank Dr. Pushraj Bhatele (Chief Radiologist) of MP MRI and CT scan center at Netaji Subhash Chandra Bose Medical College, Jabalpur, Madhya Pradesh, India for providing the clinical COVID-19 positive and Normal CT scan images for this research study. We are also thankful to all the radiologists for providing their valuable support in terms of providing knowledge and validate this work. We would also like to extend our gratitude to the Sanya Hospital and Diagnostics Pvt. Ltd. Delhi for supporting this research study.
Funding
This research study is supported by the Indian Radiological and Imaging Association.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Ethical Approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Consent to Participate
This article does not contain any studies with human participants or animals performed by any of the authors.
Consent for Publication
All the authors have given their consent.
Conflict of Interest
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Bhatele, K.R., Jha, A., Tiwari, D. et al. COVID-19 Detection: A Systematic Review of Machine and Deep Learning-Based Approaches Utilizing Chest X-Rays and CT Scans. Cogn Comput (2022). https://doi.org/10.1007/s12559-022-10076-6
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s12559-022-10076-6