Abstract
Purpose
Long noncoding RNAs (lncRNAs) are outstanding as novel cancer biomarkers with great prospects. Herein, we focused on summarizing the overall diagnostic evaluation of lncRNAs for hepatocellular carcinoma (HCC).
Methods
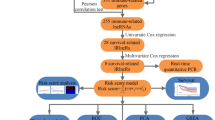
Relevant literature was collected from the online databases. The Quality Assessment for Studies of Diagnostic Accuracy checklist was used to assess the quality of included studies. The pooled sensitivity, specificity, and diagnostic odds ratio (DOR) were plotted using random-effects models. Summary receiver operating characteristic curve and the area under the curve (AUC) were used to estimate the overall test performance. Statistical analysis was performed by STATA 14.0 and Meta-DiSc 1.4 software.
Results
Ten studies with a total of 820 HCC patients and 785 healthy controls were included. For overall lncRNAs, the pooled sensitivity, specificity, and DOR to predict HCC patients were 80% [95% confidence interval (CI) 77–82%], 79% (95% CI 76–81%), and 27.66 (95% CI 14.26–53.63), respectively, corresponding to an AUC of 0.91.
Conclusions
LncRNAs were a high diagnostic value for HCC and its expression could potentially be used as auxiliary biomarker in confirming HCC.




Similar content being viewed by others
References
Honeyman JN, Simon EP, Robine N, Chiaroni-Clarke R, Darcy DG, Lim II, et al. Detection of a recurrent DNAJB1-PRKACA chimeric transcript in fibrolamellar hepatocellular carcinoma. Science. 2014;343:1010–4.
Anestopoulos I, Voulgaridou GP, Georgakilas AG, Franco R, Pappa A, Panayiotidis MI. Epigenetic therapy as a novel approach in hepatocellular carcinoma. Pharmacol Therapeut. 2015;145:103–19.
Collins JF. Long noncoding RNAs and hepatocellular carcinoma. Gastroenterology. 2015;148:291–4.
Xue M, Chen W, Li X. Urothelial cancer associated 1: A long noncoding RNA with a crucial role in cancer. J Cancer Res Clin Oncol. 2016;142:1407–19.
Chang Y, Zhang K, Hu Z, Qi H, Shi Z, Han X, et al. Hypoxia-regulated lncRNAs in cancer. Gene. 2016;575:1–8.
Evans JR, Feng FY, Chinnaiyan AM. The bright side of dark matter: LncRNAs in cancer. J Clin Invest. 2016;126:2775–82.
Yu S, Li G, Wang Z, Wang Z, Chen C, Cai S, et al. The prognostic value of pSTAT3 in gastric cancer: a meta-analysis. J Cancer Res Clin Oncol. 2016;142:649–57.
Serghiou S, Kyriakopoulou A, Ioannidis JP. Long noncoding RNAs as novel predictors of survival in human cancer: a systematic review and meta-analysis. Mol Cancer. 2016;15:50.
Chen J, Chen Y, Gu L, Li X, Gao Y, Lyu X, et al. LncRNAs act as prognostic and diagnostic biomarkers in renal cell carcinoma: a systematic review and meta-analysis. Oncotarget. 2016;7(45):74325–36. doi:10.18632/oncotarget.11101.
Whiting PF, Rutjes AW, Westwood ME, Mallett S, Deeks JJ, Reitsma JB, et al. QUADAS-2: a revised tool for the quality assessment of diagnostic accuracy studies. Ann Intern Med. 2011;155:529–36.
Chu H, Guo H. A unification of models for meta-analysis of diagnostic accuracy studies. Biostatistics. 2009;10:201–3.
Higgins JPT, Thompson SG. Quantifying heterogeneity in a meta-analysis. Stat Med. 2002;21:1539–58.
Hao H, Chen L, Huang D, Ge J, Qiu Y, Hao L. Meta-analysis of alkaline phosphatase and prognosis for osteosarcoma. Eur J Cancer Care (Engl). 2016. doi:10.1111/ecc.12536.
Deeks JJ, Macaskill P, Irwig L. The performance of tests of publication bias and other sample size effects in systematic reviews of diagnostic test accuracy was assessed. J Clin Epidemiol. 2005;58:882–93.
Konishi H, Ichikawa D, Yamamoto Y, Arita T, Shoda K, Hiramoto H, et al. Plasma level of metastasis-associated lung adenocarcinoma transcript 1 is associated with liver damage and predicts development of hepatocellular carcinoma. Cancer Sci. 2016;107:149–54.
Jing W, Gao S, Zhu M, Luo P, Jing X, Chai H, et al. Potential diagnostic value of lncRNA SPRY4-IT1 in hepatocellular carcinoma. Oncol Rep. 2016;36:1085–92.
El-Tawdi AH, Matboli M, El-Nakeep S, Azazy AE, Abdel-Rahman O. Association of long noncoding RNA and c-JUN expression in hepatocellular carcinoma. Expert Rev Gastroenterol Hepatol. 2016;10:869–77.
Kamel MM, Matboli M, Sallam M, Montasser IF, Saad AS, El-Tawdi AH. Investigation of long noncoding RNAs expression profile as potential serum biomarkers in patients with hepatocellular carcinoma. Transl Res. 2016;168:134–45.
Ma W, Wang H, Jing W, Zhou F, Chang L, Hong Z, et al. Downregulation of long non-coding RNAs JPX and XIST is associated with the prognosis of hepatocellular carcinoma. Clin Res Hepatol Gastroenterol. 2016. doi:10.1016/j.clinre.2016.09.002 (In Press).
Lu J, Xie F, Geng L, Shen W, Sui C, Yang J. Investigation of serum lncRNA-uc003wbd and lncRNA-AF085935 expression profile in patients with hepatocellular carcinoma and HBV. Tumour Biol. 2015;36:3231–6.
Li J, Wang X, Tang J, Jiang R, Zhang W, Ji J, et al. HULC and linc00152 act as novel biomarkers in predicting diagnosis of hepatocellular carcinoma. Cell Physiol Biochem. 2015;37:687–96.
Tang J, Jiang R, Deng L, Zhang X, Wang K, Sun B. Circulation long non-coding RNAs act as biomarkers for predicting tumorigenesis and metastasis in hepatocellular carcinoma. Oncotarget. 2015;6:4505–15.
Wang K, Guo WX, Li N, Gao CF, Shi J, Tang YF, et al. Serum LncRNAs profiles serve as novel potential biomarkers for the diagnosis of HBV-Positive hepatocellular carcinoma. PLoS ONE. 2015;10:e144934.
Xie H, Ma H, Zhou D. Plasma HULC as a promising novel biomarker for the detection of hepatocellular carcinoma. Biomed Res Int. 2013;2013:136106.
Aznar-Oroval E, Mancheno-Alvaro A, Garcia-Lozano T, Sanchez-Yepes M. Likelihood ratio and Fagan’s nomogram: 2 basic tools for the rational use of clinical laboratory tests. Rev Calid Asist. 2013;28:390–1.
Fusar-Poli P, Cappucciati M, Rutigliano G, Heslin M, Stahl D, Brittenden Z, et al. Diagnostic stability of ICD/DSM first episode psychosis diagnoses: meta-analysis. Schizophr Bull. 2016;42:1395–406.
Li CH, Chen Y. Targeting long non-coding RNAs in cancers: progress and prospects. Int J Biochem Cell Biol. 2013;45:1895–910.
Esposti DD, Hernandez-Vargas H, Voegele C, Fernandez-Jimenez N, Forey N, Bancel B, et al. Identification of novel long non-coding RNAs deregulated in hepatocellular carcinoma using RNA-sequencing. Oncotarget. 2016;7:31862–77.
Ouyang SS, Zhang P, Wang J, Huang ZK, Liao L. Expression of long non-coding RNA colon cancer associated transcript 2 and its clinicopathologic significance in oral squamous cell carcinoma. Zhonghua Kou Qiang Yi Xue Za Zhi. 2016;51:286–91.
Nie F, Yu X, Huang M, Wang Y, Xie M, Ma H, et al. Long noncoding RNA ZFAS1 promotes gastric cancer cells proliferation by epigenetically repressing KLF2 and NKD2 expression. Oncotarget. 2016. doi:10.18632/oncotarget.9611.
Cheng Y, Jutooru I, Chadalapaka G, Corton JC, Safe S. The long non-coding RNA HOTTIP enhances pancreatic cancer cell proliferation, survival and migration. Oncotarget. 2015;6:10840–52.
Qu S, Yang X, Song W, Sun W, Li X, Wang J, et al. Downregulation of lncRNA-ATB correlates with clinical progression and unfavorable prognosis in pancreatic cancer. Tumour Biol. 2016;37:3933–8.
Zhao JH, Sun JX, Song YX, Chen XW, Yang YC, Ma B, et al. A novel long noncoding RNA-LOWEG is low expressed in gastric cancer and acts as a tumor suppressor by inhibiting cell invasion. J Cancer Res Clin Oncol. 2016;142:601–9.
Chauhan R, Lahiri N. Tissue- and serum-associated biomarkers of hepatocellular carcinoma. Biomark Cancer. 2016;8:37–55.
Wang Z, Gou W, Liu M, Sang W, Chu H, Zhang W. Expression of p53 and HSP70 in chronic hepatitis, liver cirrhosis, and early and advanced hepatocellular carcinoma tissues and their diagnostic value in hepatocellular carcinoma: an immunohistochemical study. Med Sci Monit. 2015;21:3209–15.
Kudo M, Izumi N, Sakamoto M, Matsuyama Y, Ichida T, Nakashima O, et al. Survival analysis over 28 years of 173,378 patients with hepatocellular carcinoma in Japan. Liver Cancer. 2016;5:190–7.
Li Z, You K, Li J, Wang Y, Xu H, Gao B, et al. Madecassoside suppresses proliferation and invasiveness of HGF-induced human hepatocellular carcinoma cells via PKC-cMET-ERK1/2-COX-2-PGE2 pathway. Int Immunopharmacol. 2016;33:24–32.
Smekalova EM, Kotelevtsev YV, Leboeuf D, Shcherbinina EY, Fefilova AS, Zatsepin TS, et al. LncRNA in the liver: prospects for fundamental research and therapy by RNA interference. Biochimie. 2016;131:159–72.
Parasramka MA, Maji S, Matsuda A, Yan IK, Patel T. Long non-coding RNAs as novel targets for therapy in hepatocellular carcinoma. Pharmacol Ther. 2016;161:67–78.
Cui Z, Chen Y, Xiao Z, Hu M, Lin Y, Chen Y, et al. Long noncoding RNAs as auxiliary biomarkers for gastric cancer screening: a pooled analysis of individual studies. Oncotarget. 2016;7:25791–800.
Acknowledgements
We thank Elsevier’s English Language Editing service for editing the manuscript. This work was supported by grants from the National Natural Science Foundation of China (81560475).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors have no conflicts to disclose.
Additional information
C. Zheng and H. Hao equally contributed to this work.
Rights and permissions
About this article
Cite this article
Zheng, C., Hao, H., Chen, L. et al. Long noncoding RNAs as novel serum biomarkers for the diagnosis of hepatocellular carcinoma: a systematic review and meta-analysis. Clin Transl Oncol 19, 961–968 (2017). https://doi.org/10.1007/s12094-017-1626-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12094-017-1626-1