Abstract
This paper describes about the surface plasmon resonance (SPR) based high affinity detection of recombinant antigen H1N1 (rH1N1HA). The modification of gold SPR chip with 4-MBA and anti-rH1N1HA were in situ characterized by SPR and electrochemical impedance spectroscopy (EIS). By using kinetic evaluation software, K D and Bmax values were calculated and found to be 0.278 pM and 84.51 mo, respectively for the interaction of rH1N1HA with immobilized anti-rH1N1HA. In addition, thermodynamic parameters such as ∆G (−49.78 kJ/mol), ∆H (98.25 kcal/mol), and ∆S (72.59 cal/mol.K) were determined first time for the interaction between anti-rH1N1HA and rH1N1HA, and these values revealed that this interaction is spontaneous, endothermic, and entropy driven one. The kinetics and thermodynamic results of this study revealed that the interaction between the immobilized anti-rH1N1HA with rH1N1HA is more effective than that of interaction of anti-rH1N1HA with the immobilized rH1N1HA. The sensing linearity observed for rH1N1HA is from 0.015 pg to 1.5 ng/ml. The present SPR-based approach may be used in the hospitals equipped with SPR for the early screening of clinical samples in a label-free, real-time manner.
Similar content being viewed by others
Introduction
Endemic, epidemic, and pandemic are three words that demote to the spread of infectious diseases among a population, but on different scales. An infection is endemic when it affects a very specific and limited group of individuals at a constant and stable rate. An epidemic occurs when new cases of a certain disease in a given human population, during a given period, substantially exceed the normal rate. Pandemic infections indicate a far higher number of people with a much larger affected region than an epidemic [1–3].
In the case of influenza A virus (H1N1) or swine influenza virus (SIV), it seems suitable to use the term “pandemic” where the number of people with the disease is still small, but the rate of infection is above normal and this disease is present in several continents [4].
In April 2009, a novel influenza A virus (H1N1) was noticed in Mexico and the USA, from which the virus quickly spread across the world [5]. By May 2010, the H1N1 pandemic had caused over 18,000 deaths and had a noteworthy universal economic impact [6]. The 2009 H1N1 influenza pandemic was the first pandemic of the 21st century and a major global public health crisis.
Influenza A viruses are segmented single-stranded RNA viruses from the family Orthomyxoviridae. The majority of influenza A viruses infect a broad variety of avian and mammalian species [7]. Influenza viruses are divided into three types: A, B, and C; among these, C is not as important clinically as the A and B types. Virus type A is further subdivided into various subtypes based on the structure and conformation of its two most important envelope glycoproteins: hemagglutinin (HA) and neuraminidase (NA) [8]. HA, the most abundant surface protein consists of three identical monomers with 3 to 9 N-linked glycosylation sites. The type of glycosylation and the glycan structure depend on the virus subtype [9]. HA mediates both receptor (glycan) binding and membrane fusion for cell entry. NA functions as the receptor-demolishing enzyme in virus release [10, 11].
HA is a homo-trimer composed of disulfide-linked polypeptide chains, HA1 and HA2, formed due to cleaving the host’s enzymes at the proteolytic site; the former is the main module of the HA antigen. At present, 16 sub-types of HAs and nine subtypes of NAs (N1–N9) have been recognized in avian species (H1–H16) [12–15]. However, the previous four decades have witnessed an increasing number of individual cases of avian influenza virus infections, including H5N1, H7N2, H7N7, and H9N2 sub-types totalling about 50 confirmed transmissions [16–24].
Conventional methods of virus diagnosis including virus isolation, polymerase chain reaction (PCR), and enzyme-linked immunosorbent assay (ELISA) require extensive sample handling, laboratory infrastructure, and long sample-to-result time [25]. Classical methods of influenza virus diagnostics are based on virus isolation, culture, and subtype identification by immunoassays, possibly followed by an in vivo experiment to determine pathogenicity [26]. This process requires 3 to 7 days of virus culture in addition to transporting the sample to the laboratory [27]. The foregoing methods necessitate sophisticated bench-top instrumentation, to avoid tedious and time-consuming sample preparation and inclusion of trained personnel. Furthermore, in most of the cases, the amount of sample available for testing is often very small, e.g., blood and cerebrospinal fluid from a neonate. Hence, it is essential to develop a simple, sensitive, and reliable antigen detection technique for the detection of H1N1 virus.
For the direct detection of bimolecular interactions, surface plasmon resonance (SPR) immunosensors have attracted a lot of attention [28] because of its high sensitivity, label-free, and real-time monitoring ability [29]. A novel assay for influenza virus quantification of hemagglutinin (HA) by single-radial immunodiffusion (SRID) using SPR is reported [30]. In addition, SPR technique particularly by combining it with fluorescence spectroscopy was also reported [31, 32].
The use of fluorescence detection schemes in combination with the resonant excitation of surface plasmons was shown to increase the sensitivity for bioanalyte monitoring considerably. The applicability of a new technique, microwave-accelerated surface plasmon-coupled luminescence (MA-SPCL) for fast and sensitive bioassays in buffer, serum, and whole blood using quantum dots as luminescence reporters is also demonstrated [33]. Moreover, SPR sensing of anti-rH1N1HA was reported by our team by immobilizing rH1N1HA on the SPR chip [34]. In continuation to our earlier studies on the development of SPR detection methodologies for BWAs [35–38] in this present work, SPR-based methodology is developed for the detection of rH1N1HA in phosphate buffered saline (PBS) medium using 4-mercaptobenzoic acid (4-MBA) modified gold chip immobilized with anti-rH1N1HA. To the best of our knowledge, so far, no SPR report is available for the sensing of rH1N1HA with its immobilized anti-rH1N1HA on 4-MBA modified gold SPR chip. This modified gold chip was characterized by SPR and electrochemical impedance spectroscopy (EIS). Parameters affecting the response of SPR were optimized and finally, equilibrium constant (K D) and maximum binding capacity of analyte (Bmax) were calculated, and also for the first time, thermodynamic parameters such as change in Gibb’s free energy (ΔG), change in enthalpy (ΔH), and change in entropy (ΔS) involved in the interaction between immobilized anti-rH1N1HA and rH1N1HA were also deduced in this study.
Materials and Methods
Chemicals and Reagents
N-(3-dimethylaminopropyl)-N-ethylcarbodiimide hydrochloride (EDC), N-hydroxysuccinimide (NHS), phosphate buffered saline (PBS), sodium acetate, ethanolamine, and hydrochloric acid (HCl) were of Fluka grade and obtained from Sigma-Aldrich, Bangalore, Karnataka, India. Glacial acetic acid, glycine, sodium hydroxide (NaOH), and methanol (MeOH) were supplied by Sigma-Aldrich, Bangalore, Karnataka, India. 4-MBA Aldrich purchased from Sigma-Aldrich, Bangalore, Karnataka, India was used for the modification of the SPR gold chip (XanTec Bioanalytics GmbH, Metrowingerplatz, Germany). All chemicals and reagents used were of analytical grade, and purification was performed wherever necessary before use.
Moreover, rH1N1HA antigen (M.Wt. 75 kDa) was prepared by the trained biologists as reported earlier [34] in our establishment in addition to the raising of anti-rH1N1HA (M.Wt. 160 kDa). The concentration of rH1N1HA antigen (15 mg/ml) and anti-rH1N1HA (5 mg/ml) were used in this study.
Different buffer solutions were used in this study depending on pH [acetate buffer (pH 4.0–5.5), PBS buffer (pH 6.0–7.5), and glycine–NaOH buffer (pH 8.0–9.0)] for the optimization of pH. All solutions were prepared using water from a Milli-Q system (Millipore India, Bangalore, Karnataka, India) throughout the experiment.
Instruments
The biomolecular interactions were explored using a two channel cuvette-based electrochemical surface plasmon resonance system (Autolab ESPRIT, Ecochemie B.V., The Netherlands) and diode laser is a source with a fixed wavelength of 670 nm in conjunction with a scanning mirror to modulate the plane polarized light beam on the SPR substrate. The outcome of the SPR measurement was automatically monitored using a PC with data acquisition software version 4.3.1. All kinetic data were obtained using kinetic evaluation software version 5.0 (Ecochemie B.V.). Bare gold chip was modified with 4-MBA by using spin coater (Autolab ESPRIT, Ecochemie B.V., The Netherlands). The pH of the buffers was measured with a EUTECH instruments pH metre (pH 1500, Ayer Rajah Crescent, Singapore). All experiments were carried out at 25 °C unless otherwise specified, and the temperature of cuvette was controlled by a Julabo HE-4 (Julabo Labortechnik GmbH, Seelbach, Germany) water bath.
Preparation of 4-MBA Modified Gold Chip
Bare gold SPR chip was modified with 0.01 M methanolic solution of 4-MBA using a spin coater. Firstly 75 μl of 4-MBA solution was dispensed on bare gold chip and spread at 100 rpm. After this, spin speed was increased up to 2,500 rpm and kept for 5 min to spread the 4-MBA homogenously on the gold SPR chip.
Immobilization of Anti-rH1N1HA on 4-MBA Modified Gold SPR Chip
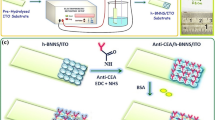
Prior to the immobilization of anti-rH1N1HA on 4-MBA modified gold SPR chip, 50 μl of PBS buffer (pH 7.5) was passed every 120 s interval for 60 s in order to get a stable baseline in both the channel. 4-MBA modified gold SPR chip was chemically activated by the injection of 75 μl of a 1:1 mixture of 400 mM EDC and 100 mM NHS. Subsequently, 75 μl of anti-rH1N1HA (1:100 dilution in 10 mM PBS) was injected in channel 1 for 1,800 s to get an effective immobilization of anti-rH1N1HA on the activated 4-MBA modified gold SPR chip surface. Following anti-rH1N1HA immobilization, the remaining active sites were blocked with 75 μl of 1,000 mM ethanolamine. Afterwards, 10 mM HCl was injected to achieve regeneration of immobilized sensor surface. For negative control measurements, the modified gold surface was activated with EDC/NHS and then quenched with ethanolamine (channel 2) as mentioned above and was used as blank control surface. This anti-rH1N1HA that was immobilized on the modified gold SPR chip was interacted with different concentration of rH1N1HA, in order to see and also deduce kinetic and thermodynamic parameters.
Biosensing Protocol
Binding studies was performed using different dilutions of rH1N1HA in PBS. The entire SPR sensing methodology was executed by a sequence of automatic procedure (baseline, association, dissociation, regeneration, and back to baseline steps). A sample solution containing a selected concentration of rH1N1HA in the PBS was injected in both channels from the 384 well microtiter plate and then association step was performed for 500 s and dissociation step was performed for 400 s followed by the regeneration of the sensor surface by addition of 10 mM HCl for 120 s. PBS (pH 7.5) was used as the running buffer solution throughout the study. For SPR sensing, an aliquot of the analyte solution (75 μL) was injected and mixed at 16.7 μL/s and this protocol was adopted with 1:12,800, 1:6,400, 1:3,200, 1:1,600, 1:800, and 1:400 dilution of rH1N1HA. The resultant data in these experiments were further utilized for the calculation kinetic parameters and also for plotting the calibration graph.
Optimization of Experimental Parameters
Effect of Temperature on SPR Response
Temperature is a well-known physical parameter to influence sensitivity of SPR measurement, and temperature variation allows to find out the temperature dependence of molecular interactions and also for optimizing detection conditions [39]. The determination of the reaction constants at different temperatures also gives the opportunity to access the thermodynamic parameters for anti-rH1N1HA and rH1N1HA interaction. The two-channel SPR system having a reference channel to compensate for bulk refractive index (RI) changes, non-specific binding, and minor temperature variations, while the temperature controller device (water bath) provides the temperature stability necessary for monitoring small changes in RI [40]. Changes in temperature affect the materials present on the SPR sensor chip and RI of aqueous sample solutions. Optimal temperature for binding of rH1N1HA (1:12,800 dilution) on immobilized anti-rH1N1HA was investigated by varying the temperature between 10 and 37 °C with a 3 °C increment.
Effect of pH on SPR Response
In order to know the effect of pH on the change in SPR angle, pH variation study was carried out using different buffers in the pH range from 4.0 to 9.0 with 0.5 increments. To maintain the pH, acetate buffer (pH 4.0 to 5.5), PBS buffer (pH 6.0 to 7.5), and glycine—NaOH buffer (pH 8.0 to 9.0) were used for the interaction of rH1N1HA with immobilized anti-rH1N1HA. The pH variation study was conducted from 4.0 to 9.0 as below pH 4.0 and above pH 9.0 protein losses its activity [38, 41].
Result and Discussion
Immobilization of Anti-rH1N1HA on 4-MBA Modified Gold SPR Sensor Chip
The most important requirement for the effective immobilization of ligand on the SPR gold chip surface is that the pH of the ligand solution should be above pH 3.5 and below the 0.5 unit of isoelectric point of the ligand on the gold chip surface and so that the ligand carry opposite net charges. In addition, the electrostatic interactions involved in preconcentration are favored by low ionic strength in the coupling buffer [42]. Figure 1 represents the stepwise immobilization of anti-rH1N1HA on 4-MBA modified chip, and this process comprises of nine steps. Figure 1, in the first step, stabilization of baseline was performed for 120 s and the SPR angle observed for this process was −580 mo. In the second step, for chemical binding between the 4-MBA present on the gold SPR chip surface and free amino groups of anti-rH1N1HA, activation of carboxyl groups on 4-MBA modified gold SPR chip was carried out for 900 s with EDC-NHS which showed an angle change from −580 to −84 mo. In the third step, washing was conducted with PBS and the SPR angle shifted from −84 to −397.39 mo and this is due to the conversion of 4-MBA to its ester. In the fourth step, anti-rH1N1HA was injected on 4-MBA modified gold chip, allowed to interact 1800s and an increase in SPR angle is observed from −397.39 to 101.95 mo. In the fifth step, washing was performed, and this process resulted in the SPR angle change from 101.95 to 55.89 mo. The sixth step was performed to prevent non-specific binding and also for the blocking of unreacted NHS-ester groups on 4-MBA chip; 1,000 mM ethanolamine was used for blocking and allowed to react with sensor surface for 600 s; the angle change involved here was from 55.89 to 495.33 mo. In seventh step, washing was performed for 30 s as discussed in the experimental part which resulted in the SPR angle change from 495.33 to −104.46 mo. In the eighth step, regeneration was carried out with 0.1 M hydrochloric acid for 120 s and the SPR angle change was from −104.46 to −94.65 mo. At last in ninth step, back to baseline process was conducted for 60 s, and the SPR angle change for this step was from −94.65 to 20.62 mo. The important process involved in Fig. 1 is also represented as Scheme 1.
SPR Characterization
The SPR angle of 4-MBA modified chip is shifted from −583 to 20.62 mo due to the covalent attachment of anti-rH1N1HA on 4-MBA adsorbed SPR chip results in a shift in the SPR angle, and this modification allows monitoring of the mass coverage (antigen interaction) at the surface with a high accuracy [43, 44]. Thus, the shift in SPR angle is attributed to the formation of thin layer of anti-rH1N1HA on 4-MBA modified gold SPR chip. The changes of SPR curve after anti-rH1N1HA immobilization on 4-MBA modified gold SPR chip is shown in Fig. 2. Angle change after ligand coupling observed is 454.52 mo on 4-MBA modified gold SPR chip after anti-rH1N1HA immobilization; this reveals the covalent attachment of 3.79 ng/mm2 of anti-rH1N1HA on 4-MBA modified gold chip [45].
Electrochemical Impedance Spectroscopic Characterization
EIS was used to characterize the electron transfer properties of the 4-MBA modified SPR gold chip. In Fig. 3, the semicircle diameter at higher frequency corresponds to the charge transfer limited process (R ct) [46]. In addition to the above, R ct is most influential and direct parameter to reflect the in situ changes which are occurring on the modified chip/electrolyte interface. Hence, EIS studies were conducted, obtained the data, and fit, and simulation method was adopted to find out the R ct value for bare gold SPR chip, modified gold SPR chip with 4-MBA after immobilization of anti-rH1N1HA and after interaction of rH1N1HA with anti-rH1N1HA immobilized 4-MBA modified gold SPR chip, the R ct values are found to be 0.683, 390, and 535 kΩ, respectively. Figure 3a is showing the EIS spectra for the bare gold SPR chip, Fig. 3b is showing EIS spectra for anti-rH1N1HA immobilized on 4-MBA modified gold SPR chip, and Fig. 3c is showing EIS spectra for after interaction of rH1N1HA with immobilized anti-rH1N1HA on 4-MBA modified gold SPR chip. As illustrated in Fig. 3, after immobilization of anti-rH1N1HA on 4-MBA, R ct increases significantly, showing increase in the semicircle diameter on the Nyquist plot. This is due to the presence of anti-rH1N1HA and 4-MBA on gold SPR chip surface [47]. It is observed from Fig. 3c that the Nyquist diagram includes a squeezed semicircle portion at higher frequencies, which corresponds to the electron transfer limited process, followed by a linear part characteristic of the lower frequency attributable to a diffusion-limited process [48].
Interaction of rH1N1HA With the Immobilized Anti-rH1N1HA on Gold SPR Chip
The anti-rH1N1HA immobilized on the SPR sensor chip was utilized for the sensing of different concentrations of rH1N1HA as discussed in the experimental part, and the results are depicted as SPR sensorgram in Fig. 4, and this figure exhibit concentration dependant angle changes with various concentrations of rH1N1HA from 0.015 pg to 1.5 ng/ml. Calibration graph was constructed using the response of angle changes (Fig. 4) against log concentration of antigen and presented in Fig. 5. The limit of detection (LOD) of the present method was calculated experimentally and was found to be 0.015 pg/ml rH1N1HA, and this is the minimum concentration which showed the response during the interaction with the immobilized anti-rH1N1HA on 4-MBA modified chip. It is noteworthy to compare here the result of the present work with our previous report [34], where we have immobilized the rH1N1HA on 4-MBA modified SPR chip and detected anti-rH1N1HA; the LOD found for that interaction was 0.5 ng/ml, and this value is very less than that of the value obtained in the present study 0.015 pg/ml. The findings of the present study and our earlier report [34] implies that SPR detection of rH1N1HA using anti-rH1N1HA immobilized SPR chip is better and more sensitive than that of the SPR detection of anti-rH1N1HA using rH1N1HA immobilized SPR chip (reason for this observation is discussed in Evaluation of kinetics involved in between the interaction of rH1N1HA with immobilized anti-rH1N1HA below).
Evaluation of Kinetics Involved in Between the Interaction of rH1N1HA With Immobilized Anti-rH1N1HA
The affinity interactions between immobilized antibody and antigen were characterized by the equilibrium constant (K D). The data were fitted using a simple 1:1 interaction model [49]. A + B = AB, where “A” is the injected analyte, “B” is the immobilized ligand, and “AB” is the analyte–ligand complex formed during the interaction process. In the SPR system, the signal R is proportional to the amount of (AB) and the Rmax is proportional to the initial (B). Hence, in this study, kinetic parameters such as K D and Bmax value were calculated for binding of rH1N1HA with immobilized anti-rH1N1HA using the software and were found to be 0.278 pM and 84.51 mo, respectively. This low K D value (if K D ≤10 nM, then it represents the high affinity interactions) represents high affinity interaction of the antigen with the immobilized anti-rH1N1HA [50]. This K D value was compared with the K D value of our earlier report [34], where we have immobilized the rH1N1HA on 4-MBA modified SPR chip and interacted anti-rH1N1HA; the K D value found for that interaction was 49.7 nM, and that value is very less than that of the value obtained in the present study 0.278 pM. These findings (present study and our earlier report [34]) implies that SPR interaction of rH1N1HA with anti-rH1N1HA immobilized SPR chip is having high affinity and the SPR interaction of anti-rH1N1HA with rH1N1HA immobilized SPR chip is having low affinity based on the K D values. This is the reason for the good LOD obtained in the present study for the SPR detection of rH1N1HA.
Evaluation of Thermodynamic Parameters
The K D obtained from kinetic evaluation software for the binding of rH1N1HA with immobilized anti-rH1N1HA was further exploited first time to calculate the thermodynamic parameters such as change in Gibb’s free energy (ΔG), change in enthalpy (ΔH), and change in entropy (ΔS) associated with binding using Van’t Hoff equation [51, 52]:
where R indicates universal gas constant, K A affinity constant, T temperature, and K 1 and K 2 are affinity constants for association at T 1 and T 2 temperature, respectively. ∆G, ∆H, and ∆S are change in Gibb’s free energy, change in the enthalpy and change in the entropy due to the interaction of rH1N1HA with immobilized anti-rH1N1HA.
The value of the change in Gibb’s free energy for binding of rH1N1HA with immobilized anti-rH1N1HA was found to be −49.78 kJ/mol at 298 K. The negative value of ∆G indicates the spontaneous interaction of rH1N1HA with immobilized anti-rH1N1HA. The calculated value of ∆H using Van’t Hoff wizard in kinetic evaluation software was found to be 98.25 kcal/mol. The positive value of ∆H reveals the interaction of rH1N1HA with immobilized anti-rH1N1HA as endothermic process [53]. The value of ∆S for binding of rH1N1HA with immobilized anti-rH1N1HA was calculated using software and was found to be 72.59 cal/mol.K. The observed positive value in entropy indicates that the interaction can be explained by Langmuir replacement reaction that exhibits Langmuir type isotherm [51]. Langmuir replacement reaction suggests that the observed positive entropy is because of desorption of water molecules from either antibody or antigen or both. The significance of desorption of water molecules from protein surfaces during ligand-protein binding was reported earlier [54].
Effect of Temperature and pH
Temperature variation study was carried out in order to know the effect of temperature on SPR response during the interaction of rH1N1HA with immobilized anti-rH1N1HA. Upon increasing temperature from 10 to 25 °C, an increase in SPR angle is observed as shown in Fig. 6a, and beyond 25 °C, SPR angle decreased. Hence, 25 °C is used as optimum temperature for the interaction of rH1N1HA with its immobilized anti-rH1N1HA on 4-MBA chip [55, 56]. Figure 6b shows the effect of pH on SPR angle due to the interaction of rH1N1HA with its immobilized anti-rH1N1HA. It is observed from Fig. 6b that SPR angle increased as increase in pH up to 7.5 and then decreased up to pH 9.0. This observation is probably due to the pH-dependent structural changes and electrostatic interactions on the sensor disc between antibody and antigen as reported earlier [57]. All these observations suggests that in pH 7.5 PBS buffer, rH1N1HA, and anti-rH1N1HA interaction is more effective and thereby resulted in more angle change; hence, pH 7.5 was preferred in this study.
a Effect of temperature on the interaction of rH1N1HA with the immobilized anti-rH1N1HA on 4-MBA modified gold chip. Temperature range 10 °C to 37 °C, pH 7.5 and rH1N1HA (dilution 1:12,800). b Effect of pH on the interaction of rH1N1HA with the immobilized anti-rH1N1HA; pH range 4.0 to 9.0, temperature 25 °C and rH1N1HA (dilution 1:12,800)
Conclusion
A label-free, real-time SPR detection methodology for rH1N1HA was developed using a 4-MBA modified SPR gold chip. The modification of gold chip with 4-MBA and anti-rH1N1HA was confirmed in situ by SPR and EIS. The detection time for rH1N1HA by SPR is found to be less than 10 min. By using kinetic evaluation software, K D and Bmax values were calculated and found to be 0.278 pM and 84.51 mo, respectively. Moreover, thermodynamic parameters such as ∆G, ∆H, and ∆S were determined first time for the interaction between rH1N1HA and anti-rH1N1HA, and the values revealed that this interaction is spontaneous, endothermic, and entropy-driven one. This study also indicates that rH1N1HA interaction with anti-rH1N1HA immobilized SPR chip is having high affinity when compared to anti-rH1N1HA interaction with rH1N1HA immobilized SPR chip based on K D and thermodynamic parameters. The present SPR-based approach may used in the hospitals equipped with SPR for the early screening of clinical samples in a label-free real-time manner.
References
Porta M (2012) Dictionary of Epidemiology. Oxford University, p 179.
Roos R (2010) WHO says H1N1 pandemic is over, Center for Infectious Disease Research and Policy. University of Minnesota, Minnesota
Palese P (2004) Influenza: old and new threats. Nat Med. doi:10.1038/nm1141
Barry JM (2004) The great influenza: the epic story of the greatest plague in history. Viking Penguin, New York
Fraser C et al (2009) Pandemic potential of a strain of influenza A (H1N1): early findings. Science 324:1557–1561
World Health Organization. Pandemic (H1N1) 2009 (2010) update (28 May 2010).
Wright P, Neumman G, Kawaoka Y (2006) Orthomyxoviruses. In: Knipe D, Howley P (eds) Fields virology. Lippincott Williams & Wilkins, Philadelphia, pp 1691–1740
Schulze IT (1997) Effects of glycosylation on the properties and functions of influenza virus hemagglutinin. J Infect Dis 176:24–28
Deom CM, Schulze IT (1985) Oligosaccharide composition of an influenza virus hemagglutinin with host-determined binding properties. J Biol Chem 260:14771–14774
World Health Organization (1980) WHO Bull 58:585–591
Suenaga E, Mizuno H, Penmetcha KKR (2012) Monitoring influenza hemagglutinin and glycan interactions using surface plasmon resonance. Biosens Bioelectron 32:195–201
Fouchier RA, Munster V, Wallensten A, Bestebroer TM, Herfst S, Smith D, Rimmelzwaan GF, Olsen B, Osterhaus AD (2005) Characterization of a novel influenza A virus hemagglutinin subtype (H16) obtained from black-headedgulls. J Virol 79:2814–2822
Garten RJ et al (2009) Antigenic and genetic characteristics of swine-origin 2009 A (H1N1) influenza viruses circulating in humans. Science 325:197–201
Scholtissek C, Rohde W, Von Hoyningen V, Rott R (1978) On the origin of the human influenza virus subtypes H2N2 and H3N2. Virology 87:13–20
Kawaoka Y, Bean WJ, Webster RG (1989) Evolution of the hemagglutinin of equine H3 influenza viruses. Virology 169:283–292
De Jong JC, Claas EC, Osterhaus AD, Webster RG, Lim WL (1997) A pandemic warning? Nature 389:554. doi:10.1038/39218
Taubenberger JK, Morens DM, Fauci AS (2007) The next influenza pandemic: can it be predicted? JAMA 297:2025–2027
Skehel JJ, Bizebard T, Bullough PA, Hughson FM, Knossow M, Steinhauer DA, Wharton SA, Wiley DC (1995) Membrane fusion by influenza hemagglutinin. Cold Spring Harb Symp Quant Biol 60:573–580
Skehel JJ, Wiley DC (2000) Receptor binding and membrane fusion in virus entry: the influenza hemagglutinin. Annu Rev Biochem 69:531–569
Choi YK, Goyal SM, Kang SW, Farnham MW, Joo HS (2002) Detection and subtyping of swine influenza H1N1, H1N2 and H3N2 viruses in clinical samples using two multiplex RT-PCR assays. J Virol Methods 102:53–59
Wang K, Holtz KM, Anderson K, Chubet R, Mahmoud W, Cox MM (2006) Expression and purification of an influenza hemagglutinin—one step closer to a recombinant protein-based influenza vaccine. Vaccine 24:2176–2185
Karlsson R, Stahlberg R (1995) Surface plasmon resonance detection and multisport sensing for direct monitoring of interactions involving low-molecular weight analytes and for determination of low affinities. Anal Biochem 228:274–280
Boa Y, Bolotov P, Dernovoy D, Kiryutin B, Zaslavsky L, Tatusova T, Ostell J, Lipman D (2008) The influenza virus resource at the National Center for Biotechnology Information. J Virol 82:596–601
Swine influenza A (H1N1) infection in two children - Southern California, Centers for Disease Control and Prevention (2009) Morb Mortal Wkly Rep 58:400–402. http://www.cdc.gov/mmwr/preview/mmwrhtml/mm58d0421a1.htm
Lee BW, Bey RF, Baarsch MJ, Simonson RR (1993) ELISA method for detection of influenza A infection in swine. J Vet Diagn Investig 5:510–515
Storch GA (2000) Essentials of diagnostic virology. The University of Michigan. Churchill Livingstone, New York
Kukol A, Li P, Estrela P, Ferrigno PK, Migliorato P (2008) Label-free electrical detection of DNA hybridization for the example of influenza virus gene sequences. Anal Biochem 374:143–153
Loyprasert S, Thavarungkul P, Asawatreratanakul P, Wongkittisuka B, Limsa- kul C, Kanatharana P (2008) Label-free capacitive immunosensor for microcystin-LR using self-assembled thiourea monolayer incorporated with Ag nanoparticles on gold electrode. Biosens Bioelectron 24:78–86
Toyama S, Shoji A, Yoshida Y, Yamauchi S, Ikariyama Y (1998) Surface design of SPR-based immunosensor for the effective binding of antigen or antibody in the evanescent field using mixed polymer matrix. Sensors Actuators B 52:65–71
Nilsson CE, Abbas S, Bennemo M, Larsson A, Hamalainen MD, Frostell-Karlsson A (2010) A novel assay for influenza virus quantification using surface plasmon resonance. Vaccine 28:759–766
Liebermann T, Knoll W (2000) Surface-plasmon field-enhanced fluorescence spectroscopy. Colloids Surf A 171:115–130
Aslan K, Previte MJR, Zhang Y, Geddes CD (2008) Microwave-accelerated surface plasmon-coupled directional luminescence 2: a platform technology for ultra fast and sensitive target DNA detection in whole blood. J Immunol Methods 331:103–113
Aslan K, Malyn SN, Geddes CD (2007) Microwave-accelerated surface Plasmon-coupled directional luminescence: application to fast and sensitive assays in buffer, human serum and whole blood. J Immunol Methods 323:55–64
Athmaram TN, Saraswat S, Sikarwar B, Verma SK, Singh AK, Boopathi M (2014) Characterization of pandemic influenza A (H1N1) virus hemagglutinin specific polyclonal antibodies for biosensor applications. J Med Virol 86:363–371
Gupta G, Sharma PK, Sikarwar B, Merwyn S, Kaushik S, Boopathi M, Agarwal GS, Singh B (2012) Surface plasmon resonance immunosensor for the detection of Salmonella typhi antibodies in buffer and patient serum. Biosens Bioelectron 36:95–102
Gupta G, Bhaskar ASB, Tripathi BK, Pandey P, Boopathi M, Lakshmana Rao PV, Singh B, Vijayaraghavan R (2011) Supersensitive detection of T-2 toxin by the in situ synthesized π-conjugated molecularly imprinted nanopatterns. An in situ investigation by surface plasmon resonance combined with electrochemistry. Biosens Bioelectron 26:2534–2540
Gupta G, Kumar A, Boopathi M, Thavaselvam D, Singh B, Vijayaraghavan R (2011) Rapid and quantitative determination of biological warfare agent Brucella abortus CSP-31 using surface plasmon resonance. Anal Bioanal Electrochem 3:26–37
Gupta G, Singh PK, Boopathi M, Kamboj DV, Singh B, Vijayaraghavan R (2010) Surface plasmon resonance detection of biological warfare agent Staphylococcal enterotoxin B using high affinity monoclonal antibody. Thin Solid Films 519:1171–1177
Ozdemir SK, Turhan-Sayan G (2003) Temperature effects on surface plasmon resonance: design considerations for an optical temperature sensor. J Lightwave Technol 21:805–814
Sharma AK, Gupta BD (2006) Theoretical model of a fiber optic sensor based on surface plasmon resonance for temperature detection. Opt Fiber Technol 12:87–100
Rogers RK, Mulchandani A (1998) Affinity biosensors: techniques and protocols. Totowa, New Jersey, p 48
Biacore Sensor Surface Handbook BR-1005-71 Edition AB
Fagerstam LG, Frostell-Karlsson A, Karlsson R, Persson B, Ronnberg I (1992) Biospecific interaction analysis using surface plasmon resonance detection applied to kinetic, binding site and concentration analysis. J Chromatogr 597:397–410
Lundstrom I (1994) Real-time biospecific interaction analysis. Biosens Bioelectron 9:725–736
Stenberg E, Persson B, Roos H, Urbaniczky C (1991) Quantitative determination of surface concentration of protein with surface plasmon resonance using radio labeled proteins. J Colloid Interface Sci 143:513–526
Bard AJ, Faulkner R (1980) Electrochemical methods: fundamentals and applications. Wiley & Sons, New York, p 368
Huang Y, Bell MC, Suni II (2008) Impedance biosensor for peanut protein Ara h 1. Anal Chem 80:9157–9161
Yu X, Xu D, Xu DR, Liu Z (2006) An impedance biosensor array for label-free detection of multiple antigen-antibody reactions. Front Biosci 11:983–990
Liu JT, Chen LY, Shih MC, Chang Y, Chen WY (2008) The investigation of recognition interaction between phenylboronate monolayer and glycated hemoglobin using surface plasmon resonance. Anal Biochem 375:90–96
Wassaf D, Kuang G, Kopacz K, Wu QL, Nguyen Q, Toews M, Cosic J, Jacques J, Wiltshire S, Lambert J, Pazmany CC, Hogan S, Ladner RC, Nixon AE, Sexton DJ (2006) High-throughput affinity ranking of antibodies using surface plasmon resonance microarrays. Anal Biochem 351:241–253
Savara A, Schmidt CM, Geiger FM, Weitz E (2009) Adsorption entropies and enthalpies and their implications for adsorbate dynamics. J Phys Chem A 113:2806–2815
Glasstone SD (1947) Thermodynamics for chemists. Van Nostrand Company, New York, p 288
Cabilio NR, Omanovic S, Roscoe S (2000) Electrochemical studies of the effect of temperature and pH on the adsorption of α-lactalbumin at Pt. Langmuir 16:8480–8488
Gregory RB (1995) Protein–Solvent Interactions. In: Marcel D (ed) New York, Chapter 11.
Moreira CS, Lima AMN, Neff H, Thirstrup C (2008) Temperature-dependent sensitivity of surface plasmon resonance sensors at the gold–water interface. Sensors Actuators B 134:854–862
Gupta G, Singh PK, Boopathi M, Kamboj DV, Singh B, Vijayaraghavan R (2010) Molecularly imprinted polymer for the recognition of biological warfare agent Staphylococcal enterotoxin B based on surface plasmon resonance. Thin Solid Films 519:1115–1121
Paynter S, Russell DA (2002) Surface plasmon resonance measurement of pH-induced responses of immobilized biomolecules: conformational change of electrostatic interaction effects? Anal Biochem 309:85–95
Acknowledgments
The authors thank Prof. Dr. M. P. Kaushik, Director, Defence Research and Development Establishment, DRDO, Gwalior474002, India for his keen interest and encouragement.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Sikarwar, B., Sharma, P.K., Saraswat, S. et al. Surface Plasmon Resonance Immunosensor for Recombinant H1N1 Protein. Plasmonics 10, 77–85 (2015). https://doi.org/10.1007/s11468-014-9780-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11468-014-9780-6