Abstract
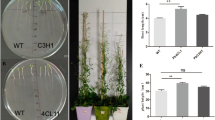
4-Coumarate/coenzyme A ligases (4CLs) play key roles in the biosynthesis of plant secondary compounds and act at the divergence point from general phenylpropanoid metabolism to several major branch pathways, such as lignins and flavonoids. In this study, five 4CL genes in Populus tomentosa were cloned and characterized. Quantitative polymerase chain reaction (qPCR) and promoter-β-glucuronidase analysis showed that the expression of Pto4CLs is significantly divergent and certainly overlapping. The five Pto4CL recombinant proteins also have different substrate specificities and catalytic turnover rates. Pto4CL4 has the highest substrate affinity and catalytic turnover rate compared with other Pto4CL proteins. All five Pto4CL recombinant proteins had no detectable activity towards sinapate. Deletion of V338 residues in Pto4CL mutants resulted in activity towards sinapate. The low accumulation of sinapate and high proportion of S unit lignin implies that the ability of 4CL to catalyze the activation of sinapate is not necessary for lignin biosynthesis. Overexpression of the five Pto4CLs in transgenic tobacco using the 35S promoter revealed significantly increased lignin contents and decreased hydroxycinnamate derivative contents in phloem, xylem, root, and leaf tissues. An increased content of naringenin in Pto4CL4 transgenic tobacco roots was also observed. Our results suggest that the substrate specificity and divergent and overlapping expression patterns of Pto4CL isoforms may play important roles in regulating the lignin and flavonoid biosynthesis.






Similar content being viewed by others
References
Alberstein M, Eisenstein M, Abeliovich H (2012) Removing allosteric feedback inhibition of tomato 4-coumarate:CoA ligase by directed evolution. Plant J 69:57–69
Allina SM, Pri-Hadash A, Theilmann DA, Ellis BE, Douglas CJ (1998) 4-Coumarate:coenzyme A ligase in hybrid poplar properties of native enzymes, cDNA cloning, and analysis of recombinant enzymes. Plant Physiol 116:743–754
Besseau S, Hoffmann L, Geoffroy P, Lapierre C, Pollet B, Legrand M (2007) Flavonoid accumulation in Arabidopsis repressed in lignin synthesis affects auxin transport and plant growth. Plant Cell 19:148–162
Costa MA, Bedgar DL, Moinuddin SGA, Kim KW, Cardenas CL, Cochrane FC, Shockey JM, Helms GL, Amakura Y, Takahashi H (2005) Characterization in vitro and in vivo of the putative multigene 4-coumarate:CoA ligase network in Arabidopsis: syringyl lignin and sinapate/sinapyl alcohol derivative formation. Phytochemistry 66:2072–2091
Cukovica D, Ehlting J, Ziffle JAV, Douglas CJ (2001) Structure and evolution of 4-coumarate:coenzyme A ligase (4CL) gene families. Biol Chem 382:645–654
Ehlting J, Büttner D, Wang Q, Douglas CJ, Somssich IE, Kombrink E (1999) Three 4-coumarate:coenzyme A ligases in Arabidopsis thaliana represent two evolutionarily divergent classes in angiosperms. Plant J 19:9–20
Endler A, Martens S, Wellmann F, Matern U (2008) Unusually divergent 4-coumarate:CoA ligases from Ruta graveolens L. Plant Mol Biol 67:335–346
Gui J, Shen J, Li L (2011) Functional characterization of evolutionarily divergent 4-coumarate: coenzyme A ligases in rice. Plant Physiol 157:574–586
Hamada K, Tsutsumi Y, Yamauchi K, Fukushima K, Nishida T (2003) Treatment of poplar callus with ferulic and sinapic acids I: incorporation and enhancement of lignin biosynthesis. J Wood Sci 49:333–338
Hamberger B, Hahlbrock K (2004) The 4-coumarate:CoA ligase gene family in Arabidopsis thaliana comprises one rare, sinapate-activating and three commonly occurring isoenzymes. Proc Natl Acad Sci U S A 101:2209–2214
Harding SA, Leshkevich J, Chiang VL, Tsai CJ (2002) Differential substrate inhibition couples kinetically distinct 4-coumarate :coenzyme A ligases with spatially distinct metabolic roles in quaking aspen. Plant Physiol 128:428–438
Hu WJ, Kawaoka A, Tsai CJ, Lung J, Osakabe K, Ebinuma H, Chiang VL (1998) Compartmentalized expression of two structurally and functionally distinct 4-coumarate:CoA ligase genes in aspen (Populus tremuloides). Proc Natl Acad Sci U S A 95:5407–5412
Hu WJ, Harding SA, Lung J, Popko JL, Ralph J, Stokke DD, Tsai CJ, Chiang VL (1999) Repression of lignin biosynthesis promotes cellulose accumulation and growth in transgenic trees. Nat Biotechnol 17:808–812
Hu Y, Gai Y, Yin L, Wang X, Feng C, Feng L, Li D, Jiang XN, Wang DC (2010) Crystal structures of a Populus tomentosa 4-coumarate:CoA ligase shed light on its enzymatic mechanisms. Plant Cell 22:3093–3104
Jiang H, Wood KV, Morgan JA (2005) Metabolic engineering of the phenylpropanoid pathway in Saccharomyces cerevisiae. Appl Environ Microbiol 71:2962–2969
Kajita S, Hishiyama S, Tomimura Y, Katayama Y, Omori S (1997) Structural characterization of modified lignin in transgenic tobacco plants in which the activity of 4-coumarate:coenzyme A ligase is depressed. Plant Physiol 114:871–879
Kaneko M, Hwang E, Ohnishi Y, Horinouchi S (2003) Heterologous production of flavanones in Escherichia coli: potential for combinatorial biosynthesis of flavonoids in bacteria. J Ind Microbiol Biotechnol 30:456–461
Knobloch KH, Hahlbrock K (1977) 4-Coumarate:CoA ligase from cell suspension cultures of Petroselinum hortense Hoffm: partial purification, substrate specificity, and further properties. Arch Biochem Biophys 184:237–248
Lapierre C, Monties B, Rolando C (1986) Thioacidolysis of poplar lignins: identification of monomeric syringyl products and characterization of guaiacyl-syringyl lignin fractions. Holzforschung 40:113–118
Lee D, Douglas CJ (1996) Two divergent members of a tobacco 4-coumarate:CoA ligase (4CL) gene family (cDNA structure, gene inheritance and expression, and properties of recombinant proteins). Plant Physiol 112:193–205
Lee D, Meyer K, Chapple C, Douglas CJ (1997) Antisense suppression of 4-coumarate:coenzyme A ligase activity in Arabidopsis leads to altered lignin subunit composition. Plant Cell 9:1985–1998
Leonard E, Chemler J, Lim KH, Koffas MA (2006) Expression of a soluble flavone synthase allows the biosynthesis of phytoestrogen derivatives in Escherichia coli. Appl Microbiol Biotechnol 70:85–91
Lindermayr C, Möllers B, Fliegmann J, Uhlmann A, Lottspeich F, Meimberg H, Ebel J (2002) Divergent members of a soybean (Glycine max L.) 4-coumarate:coenzyme A ligase gene family. Eur J Biochem 269:1304–1315
Lindermayr C, Fliegmann J, Ebel J (2003) Deletion of a single amino acid residue from different 4-coumarate:CoA ligases from soybean results in the generation of new substrate specificities. J Biol Chem 278:2781–2786
Liu Z, Zhang D, Liu D, Li F, Lu H (2013) Exon skipping of AGAMOUS homolog PrseAG in developing double flowers of Prunus lannesiana (Rosaceae). Plant Cell Rep 32:227–237
Lu H, Zhao Y, Jiang X (2004) Stable and specific expression of 4-coumarate:coenzyme A ligase gene (4CL1) driven by the xylem-specific Pto4CL1 promoter in the transgenic tobacco. Biotechnol Lett 26:1147–1152
Morita H, Mori T, Wanibuchi K, Kato R, Sugio S, Abe I (2011) Crystallization and preliminary X-ray analysis of 4-coumarate:CoA ligase from Arabidopsis thaliana. Acta Crystallogr Sect F: Struct Biol Cryst Commun 67:409–411
Naoumkina MA, Zhao Q, Gallego-Giraldo L, Dai X, Zhao PX, Dixon RA (2010) Genome-wide analysis of phenylpropanoid defence pathways. Mol Plant Pathol 11:829–846
Pan X, Li H, Wei H, Su W, Jiang X, Lu H (2012) Analysis of the spatial and temporal expression pattern directed by the Populus tomentosa 4-coumarate:CoA ligase Pto4CL2 promoter in transgenic tobacco. Mol Biol Rep 40:2309–2317
Rastogi S, Kumar R, Chanotiya CS, Shanker K, Gupta MM, Nagegowda DA, Shasany AK (2013) 4-Coumarate:CoA ligase partitions metabolites for eugenol biosynthesis. Plant Cell Physiol 54:1238–1252
Schneider K, Hövel K, Witzel K, Hamberger B, Schomburg D, Kombrink E, Stuible HP (2003) The substrate specificity-determining amino acid code of 4-coumarate:CoA ligase. Proc Natl Acad Sci U S A 100:8601–8606
Soltani BM, Ehlting J, Hamberger B, Douglas CJ (2006) Multiple cis-regulatory elements regulate distinct and complex patterns of developmental and wound-induced expression of Arabidopsis thaliana 4CL gene family members. Planta 224:1226–1238
Stuible HP, Kombrink E (2001) Identification of the substrate specificity-conferring amino acid residues of 4-coumarate:coenzyme A ligase allows the rational design of mutant enzymes with new catalytic properties. J Biol Chem 276:26893–26897
Sun H, Li Y, Feng S, Zou W, Guo K, Fan C, Si S, Peng L (2013) Analysis of five rice 4-coumarate:coenzyme A ligase enzyme activity and stress response for potential roles in lignin and flavonoid biosynthesis in rice. Biochem Biophys Res Commun 430:1151–6
Tian X, Xie J, Zhao Y, Lu H, Liu S, Qu L, Li J, Gai Y, Jiang X (2013) Sense-, antisense- and RNAi-4CL1 regulate soluble phenolic acids, cell wall components and growth in transgenic Populus tomentosa Carr. Plant Physiol Biochem 65:111–119
Voelker SL, Lachenbruch B, Meinzer FC, Jourdes M, Ki C, Patten AM, Davin LB, Lewis NG, Tuskan GA, Gunter L, Decker SR, Selig MJ, Sykes R, Himmel ME, Kitin P, Shevchenko O, Strauss SH (2010) Antisense down-regulation of 4CL expression alters lignification, tree growth, and saccharification potential of field-grown poplar. Plant Physiol 154:874–886
Wagner A, Donaldson L, Kim H, Phillips L, Flint H, Steward D, Torr K, Koch G, Schmitt U, Ralph J (2009) Suppression of 4-coumarate:CoA ligase in the coniferous gymnosperm Pinus radiata. Plant Physiol 149:370–383
Wu H, Liu C, Li M, Zhao M, Gu D, Xu Y (2013) Effects of monogalactoglycerolipid deficiency and diacylglycerol acyltransferase overexpression on oil accumulation in transgenic tobacco. Plant Mol Biol Rep 31:1077–1088
Xu B, Escamilla-Treviño LL, Sathitsuksanoh N, Shen Z, Shen H, Zhang YH, Dixon RA, Zhao B (2011) Silencing of 4-coumarate:coenzyme A ligase in switchgrass leads to reduced lignin content and improved fermentable sugar yields for biofuel production. New Phytol 192:611–25
Yamauchi K, Yasuda S, Hamada K, Tsutsumi Y, Fukushima K (2003) Multiform biosynthetic pathway of syringyl lignin in angiosperms.Planta 216:496–501
Acknowledgments
This work was supported by the National Natural Science Foundation of China (31170574 and 30671697) and the 111 Project (Project No. B13007).
Author information
Authors and Affiliations
Corresponding author
Additional information
Guodong Rao and Xiang Pan contributed equally to this work.
The English in this document has been checked by at least two professional editors, both native speakers of English. For a certificate, please see: http://www.textcheck.com/certificate/VDpuSO
Electronic supplementary material
Below is the link to the electronic supplementary material.
ESM 1
(PDF 34 kb)
Rights and permissions
About this article
Cite this article
Rao, G., Pan, X., Xu, F. et al. Divergent and Overlapping Function of Five 4-Coumarate/Coenzyme A Ligases from Populus tomentosa . Plant Mol Biol Rep 33, 841–854 (2015). https://doi.org/10.1007/s11105-014-0803-4
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11105-014-0803-4