Abstract
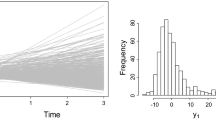
For psychiatric diseases, established mechanistic models are lacking and alternative empirical mathematical structures are usually explored by a trial-and-error procedure. To address this problem, one of the most promising approaches is an automated model-free technique that extracts the model structure directly from the statistical properties of the data. In this paper, a linear-in-parameter modelling approach is developed based on principal component analysis (PCA). The model complexity, i.e. the number of components entering the PCA-based model, is selected by either cross-validation or Mallows’ Cp criterion. This new approach has been validated on both simulated and clinical data taken from a Phase II depression trial. Simulated datasets are generated through three parametric models: Weibull, Inverse Bateman and Weibull-and-Linear. In particular, concerning simulated datasets, it is found that the PCA approach compares very favourably with some of the popular parametric models used for analyzing data collected during psychiatric trials. Furthermore, the proposed method performs well on the experimental data. This approach can be useful whenever a mechanistic modelling procedure cannot be pursued. Moreover, it could support subsequent semi-mechanistic model building.










Similar content being viewed by others
References
Mould DR, Denman NG, Duffull S (2007) Using disease progression models as a tool to detect drug effect. Clin Pharmacol Ther 82:81–86. doi:10.1038/sj.clpt.6100228
Shang EY, Gibbs MA, Landen JW, Krams M, Russell T, Denman NG, Mould DR (2009) Evaluation of structural models to describe the effect of placebo upon the time course of major depressive disorder. J Pharmacokinet Pharmacodyn 36:63–80. doi:10.1007/s10928-009-9110-3
Gomeni R, Merlo-Pich E (2006) Bayesian modelling and ROC analysis to predict placebo responders using clinical score measured in the initial weeks of treatment in depression trials. Br J Clin Pharmacol 63:595–613. doi:10.1111/j.1365-2125.2006.02815.x
Nucci G, Gomeni R, Poggesi I (2009) Model-based approaches to increase efficiency of drug development in schizophrenia: a can’t miss opportunity. Expert Opin Drug Discov 4:837–856. doi:10.1517/17460440903036073
Santen G, Danhof M, Della Pasqua O (2008) Evaluation of treatment response in depression studies using a Bayesian parametric cure rate model. J Psychiatr Res 42:1189–1197. doi:10.1016/j.jpsychires.2007.11.009
Holford N, Li J, Benincosa L, Birath M (2002) Population disease progress models for the time course of HAMD score in depressed patients receiving placebo in anti-depressant clinical trials. In: Population Approach Group in Europe (PAGE) 11th Meeting, Abstract 311. http://www.page-meeting.org/?abstract=311. Accessed 23 Jan 2012
Reddy VP, Kozielska M, Johnson M, Vermeulen A, de Greef R, Liu J, Groothuis GMM, Danhof M, Proost JH (2011) Structural models describing placebo treatment effects in schizophrenia and other neuropsychiatric disorders. Clin Pharmacokinet 50:429–450. doi:10.2165/11590590-000000000-00000
Karlsson M, Holford N (2008) A tutorial on visual predictive checks. In: Population Approach Group in Europe (PAGE) 17th Meeting, Abstract 1434. http://www.page-meeting.org/default.asp?abstract=1434. Accessed 14 Mar 2012
Jolliffe IT (2002) Principal component analysis. Springer, New York
Price AL, Patterson NJ, Plenge RM, Weinblatt ME, Shadick NA, Reich D (2006) Principal components analysis corrects for stratification in genome-wide association studies. Nat Genet 38:904–909. doi:10.1038/ng1847
Price AL, Zaitlen NA, Reich D, Patterson N (2010) New approaches to population stratification in genome-wide association studies. Nat Rev Genet 11:459–463. doi:10.1038/nrg2813
Raychaudhuri S, Stuart JM, Altman RB (2000) Principal components analysis to summarize microarray experiments: application to sporulation time series. Pac Symp Biocomput 5:452–463. doi:10.1.1.21.3169
De la Torre, Black MJ (2001) Robust principal component analysis for computer vision. International Conferences on Computer Vision, IEEE. Vancouver
Ghosh-Dastidar S, Adeli H (2008) Principal component analysis-enhanced cosine radial basis function neural network for robust epilepsy and seizure detection. IEEE Trans Biomed Eng 55:512–518. doi:10.1109/TBME.2007.905490
Abdi H, Williams LJ (2010) Principal component analysis. WIREs Comput Stat. doi:10.1002/wics.101
Santen G, van Zwet E, Danhof M, Della Pasqua O (2008) Heterogeneity in patient response in depression: the relevance of functional data analysis. In: To fail or not to fail: clinical trials in depression. Doctoral thesis, Leiden University, pp 111–126
El Yazaji M, Battas O, Agoub M, Moussaoui D, Gutknecht C, Dalery J, d’Amato T, Saoud M (2002) Validity of the depressive dimension extracted from principal component analysis of the PANSS in drug-free patients with schizophrenia. Schizophr Res 56:121–127. doi:10.1016/S0920-9964(01)00247-X
Rowland M, Sheiner LB, Steimer JL (1985) Variability in drug therapy: description, estimation and control. Raven Press, New York
Hamilton M (1960) A rating scale for depression. J Neurol Neurosur Psychiat 23:56-62. doi:10.1136/jnnp.23.1.56
Gomeni R, Lavergne A, Merlo-Pich E (2009) Modeling placebo response in depression trials using a longitudinal model with informative dropout. Eur J Pharm Sci 36:4–10. doi:10.1016/j.bbr.2011.03.031
Berry MW (1992) Large scale sparse singular value computations. Int J Supercomput Appl 6:13–49. doi:10.1.1.37.8591
Russu A, Poggesi I, Gomeni R, De Nicolao G (2011) Bayesian population modeling of Phase I dose escalation studies: Gaussian process versus parametric approaches. IEEE Trans Biomed Eng 58:3156–3164. doi:10.1109/TBME.2011.2164614
Jackson JE (1991) A user’s guide to principal components. Wiley, New York
Peres-Neto PR, Jackson DA, Somers KM (2005) How many principal components? Stopping rules for determining the number of non-trivial axes revisited. Comput Stat Data Anal 49:974–997. doi:10.1016/j.csda.2004.06.015
Cattell RB (1966) The scree test for the number of factors. Multivar Behav Res 1:245–276. doi:10.1207/s15327906mbr0102_10
Mallows CL (1973) Some comments on Cp. Technometrics 15:661–675. doi:10.2307/1267380
Mallows CL (1995) More comments on Cp. Technometrics 37:362–372. doi:10.2307/1269729
Soderstrom T, Stoica P (1989) System identification. Prentice-Hall International, Hemel Hempstead
R Development Core Team (2011) R: a language and environment for statistical computing. R Foundation for Statistical Computing, Vienna.
Molenberghs G, Thijs H, Jansen I, Beunckens C (2004) Analyzing incomplete longitudinal clinical trial data. Biostatistics 5:445–464. doi:10.1093/biostatistics/kxh001
Hu C, Sale ME (2003) A joint model for nonlinear longitudinal with informative dropout. J Pharmacokinet Pharmacodyn 30:83–103. doi:10.1023/A:1023249510224
Harville DA (1977) Maximum likelihood approaches to variance component estimation and to related problems. J Am Stat Assoc 72:320–338. doi:10.2307/2286796
Conflict of interest
The authors declare that they have no conflict of interest.
Author information
Authors and Affiliations
Corresponding author
Appendix
Appendix
How to obtain the covariance matrix of the unobservable response
The following preposition shows that the SVD of the covariance matrix \( \Sigma_{Y}\) provides also the principal vectors of the covariance matrix \(\Sigma_{X}\) of the unobservable true response \( X_i, i = 1, \ldots, M.\)
Proposition
Let U X = U Y and D X = D Y − σ 2 I. Then, the SVD of \( \Sigma_X \) is \( \Sigma_X = U_X D_X U_X^T\).
Proof
In view of (16),
□
Hyperparameter estimation
If the variance matrix \( \Sigma_{\overline{X}} \) of the fixed effect parameters \( \overline{X} \) were finite, the marginal likelihood would be given by
Hence, the ML estimate \( \varvec{\lambda}^{ML} \) would be
In order to obtain the ML estimate of \( \varvec{\lambda}\), the fixed effect parameter vector \( \overline{X} \) must be integrated out of the joint density \( p({\bf Y},\overline{X})\). Indeed,
where \( p(\overline{X}|\varvec{\lambda}) \) is the prior density for the fixed-effect vector \( \overline{X}\).
In the proposed model, an improper prior for \( \overline{X} \) is assumed, that is \( \Sigma_{ \overline{X}} \) is infinite. Fortunately, the improper, flat nature of \( p(\overline{X}|\varvec{\lambda}) \) does not hinder the closed-form calculation of a function \( G(\varvec{\lambda}) \) proportional to the marginal likelihood:
By rearranging the integral (31) as explained in [22], the result is
where \( \Sigma = {\varvec{\widetilde{\Phi}}} {\varvec{\Omega}} {\varvec{\widetilde{\Phi}}}^T + \sigma^2 {\bf I}_{Mn}, \overline{\Sigma} = ({\varvec{\overline{\Phi}}}^T \Sigma^{-1} {\varvec{\overline{\Phi}}})^{-1} \) and \( \gamma^T \gamma = {\bf Y}^T \Sigma^{-1} {\bf Y} - {\bf Y}^T \Sigma^{-1} \varvec{\overline{\Phi}} \widetilde{\Sigma} \varvec{\overline{\Phi}}^T \Sigma^{-1} {\bf Y}\).
Hence, the ML estimation problem is reformulated as
Estimation of fixed and random effects
Once the hyperparameter vector has been estimated, as specified in the previous subsection, then the fixed and random effects, \( \overline{X} \) and \( \varvec{\eta}\), are easily estimated applying the standard conditional posterior formula for the model (21):
Connection with Restricted Maximum Likelihood approach
In this work, parameters are estimated according to an established approach. As shown by Harville [32] in a frequentist framework, the same point estimates of λ, \(\overline{X}\) and η would be obtained by resorting to Restricted Maximum Likelihood estimation.
Rights and permissions
About this article
Cite this article
Marostica, E., Russu, A., Gomeni, R. et al. A PCA approach to population analysis: with application to a Phase II depression trial. J Pharmacokinet Pharmacodyn 40, 213–227 (2013). https://doi.org/10.1007/s10928-013-9304-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10928-013-9304-6