Abstract
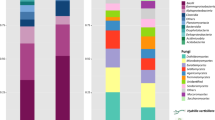
The phylogenetic composition and physiological profiles of bacterial communities in freshwater springs were evaluated during the blooming and non-blooming stages of the invasive plant species, Hydrilla verticillata. Community-level physiological profiles (CLPPs) and pyrosequencing of 16S rRNA gene amplicons were used to study potential Hydrilla mediated shifts in the physiological potential and phylogenetic composition of the bacterial community in infested systems. The results of CLPP revealed that the microbes in the Hydrilla invaded sites utilized less substrates during blooming periods than during non-blooming periods of the plant. Spearman’s rank correlation analysis showed some relationships between the relative abundances of bacterial taxa and the Biolog substrate utilization pattern. The relative abundance of the identified taxa showed some striking differences based on the blooming status of Hydrilla and to a lesser extent on site variation. The relative abundance of Actinobacteria, Bacteriodetes, and Verrucomicrobia was generally higher during Hydrilla blooms, while Deltaproteobacteria was generally higher during non-blooming stages of Hydrilla. The detected genera also varied based on the blooming stages of the plant. Based on the findings, it appears that Hydrilla alters the phylogenetic composition and structure of the bacterial community during the blooming stage.





Similar content being viewed by others
References
Allgaier, M. & H.-P. Grossart, 2006. Diversity and seasonal dynamics of Actinobacteria populations in four lakes in northeastern Germany. Applied and Environmental Microbiology 72: 3489–3497.
Angeloni, N. L., et al., 2006. Effects of an invasive cattail species (Typha × glauca) on sediment nitrogen and microbial community composition in a freshwater wetland. FEMS Microbiology Letters 263: 86–92.
AWWA, APHA, 1998. Standard Methods for the Examination of Water and Wastewater, 20th ed. APHA, Washington, DC.
Blackburn, R. D., et al., 1969. Identification and distribution of certain similar-appearing submersed aquatic weeds in Florida. Hyacinth Control Journal 8: 17–21.
Campbell, C. D., S. J. Grayston & D. J. Hirst, 1997. Use of rhizosphere carbon sources in sole carbon source tests to discriminate soil microbial communities. Journal of Microbiological Methods 30: 33–41.
Cao, H., J. B. Cowart & J. K. Osmond, 1999. Water sources of Wakulla Springs, Wakulla County, Florida: physical and uranium isotopic evidences. Southeastern Geology 39: 51–55.
Chao, A., 1984. Nonparametric estimation of the number of classes in a population. Scandinavian Journal of Statistics 11: 265–270.
Chao, A. & S. M. Lee, 1992. Estimating the number of classes via sample coverage. Journal of the American Statistical Association 87: 210–217.
Classen, A. T., et al., 2006. Community-level physiological profiles of bacteria and fungi: plate type and incubation temperature influences on contrasting soils. FEMS Microbiology Ecology 44: 319–328.
Coles, S. L., et al., 1999. Historical and recent introductions of non-indigenous marine species into Pearl Harbor, Oahu, Hawaiian Islands. Marine Biology 135: 147–158.
Costa, R., et al., 2005. Effects of site and plant species on rhizosphere community structure as revealed by molecular analysis of microbial guilds. FEMS Microbiology Ecology 56: 236–249.
D’Antonio, C. M. & P. M. Vitousek, 1992. Biological invasions by exotic grasses, the grass/fire cycle, and global change. Annual Review of Ecology and Systematics 23: 63–87.
Dowd, S. E., et al., 2008. Polymicrobial nature of chronic diabetic foot ulcer biofilm infections determined using bacterial tag encoded FLX amplicon pyrosequencing (bTEFAP). PLoS One 3: e3326.
Du, H., et al., 2006. Diversity and distribution of pigmented heterotrophic bacteria in marine environments. FEMS Microbiology Ecology 57: 92–105.
Duda, J. J., et al., 2003. Differences in native soil ecology associated with invasion of the exotic annual chenopod, Halogeton glomeratus. Biology and Fertility of Soils 38: 72–77.
Edgar, R. C., et al., 2011. UCHIME improves sensitivity and speed of chimera detection. Bioinformatics 27: 2194–2200.
EPA, 1986. Test Methods for Evaluating Solid Waste, Physical/Chemical Methods. US EPA, Cincinnati, OH, USA.
Fiehn, O., et al., 2000. Metabolite profiling for plant functional genomics. Nature Biotechnology 18: 1157–1161.
Garbeva, P., et al., 2005. Effect of above-ground plant species on soil microbial community structure and its impact on suppression of Rhizoctonia solani AG3. Environmental Microbiology 8: 233–246.
Garland, J. L. & A. L. Mills, 1991. Classification and characterization of heterotrophic microbial communities on the basis of patterns of community-level sole-carbon-source utilization. Applied and Environmental Microbiology 57: 2351–2359.
Gomez, E., J. Garland & M. Conti, 2004. Reproducibility in the response of soil bacterial community-level physiological profiles from a land use intensification gradient. Applied Soil Ecology 26: 21–30.
Goodfellow, M. & S. T. Williams, 1983. Ecology of actinomycetes. Annual Reviews in Microbiology 37: 189–216.
Gordon-Bradley, N., D. Lymperopoulo & H. N. Williams, 2013. Differences in bacterial community structure on Hydrilla vertlicillata and Vallisnera americana in a Freshwater Spring. Microbes and Environment 29: 67–73.
Hahn, D. R., 2003. Alteration of microbial community composition and changes in decomposition associated with an invasive intertidal macrophyte. Biological Invasions 5: 45–51.
Hammer, Ø., D. A. T. Harper & P. D. Ryan, 2001. PAST-Palaeontological Statistics. www.uv.es/~pardomv/pe/2001_1/past/pastprog/past.pdf. Accessed on 25.7.2009.
Han, X., et al., 2007. Effects of vegetation type on soil microbial community structure and catabolic diversity assessed by polyphasic methods in North China. Journal of Environmental Sciences 19: 1228–1234.
He, D., L. Ren & Q. Wu, 2012. Epiphytic bacterial communities on two common submerged macrophytes in Taihu Lake: diversity and host-specificity. Chinese Journal of Oceanology and Limnology 30: 237–247.
Hipskind, J., et al., 1992. A fungitoxic phenolic compound in Hydrilla verticillata. Biological Control 2: 51–58.
Huse, S. M., et al., 2010. Ironing out the wrinkles in the rare biosphere through improved OTU clustering. Environmental Microbiology 12: 1889–1898.
Insam, H., 1997. A new set of substrates proposed for community characterization in environmental samples. In Insam, H. & A. Rangger (eds), Microbial Communities, Functional versus Structural Approaches. Springer, Berlin: 259–260.
Kourtev, P. S., J. G. Ehrenfeld & M. Häggblom, 2003. Experimental analysis of the effect of exotic and native plant species on the structure and function of soil microbial communities. Soil Biology and Biochemistry 35: 895–905.
Langeland, K. A., 1996. Hydrilla verticillata (LF) Royle (Hydrocharitaceae), “The perfect aquatic weed”. Castanea 61: 293–304.
Newton, R. J., et al., 2011. A guide to the natural history of freshwater lake bacteria. Microbiology and Molecular Biology Reviews 75: 14–49.
Nielsen, J. L., et al., 2006. Detection of activity among uncultured actinobacteria in a drinking water reservoir. FEMS Microbiology Ecology 55: 432–438.
Pernthaler, J., et al., 1998. Seasonal community and population dynamics of pelagic bacteria and archaea in a high mountain lake. Applied and Environmental Microbiology 64: 4299–4306.
Posey, M. H., 1988. Community changes associated with the spread of an introduced seagrass, Zostera japonica. Ecology 69: 974–983.
Ravit, B., J. G. Ehrenfeld & M. M. Haggblom, 2003. A comparison of sediment microbial communities associated with Phragmites australis and Spartina alterniflora in two brackish wetlands of New Jersey. Estuaries 26: 465–474.
Rheims, H., et al., 1999. Molecular monitoring of an uncultured group of the class Actinobacteria in two terrestrial environments. Journal of Microbiological Methods 36: 65–75.
Roeselers, G., et al., 2011. Evidence for a core gut microbiota in the zebrafish. The ISME Journal 5: 1595–1608.
Rohwer, F., et al., 2001. Diversity of bacteria associated with the Caribbean coral Montastraea franksi. Coral Reefs 20: 85–91.
Schloss, P. D., D. Gevers & S. L. Westcott, 2011. Reducing the effects of PCR amplification and sequencing artifacts on 16S rRNA-based studies. PloS One 6: e27310.
Söderberg, K. H., P. A. Olsson & E. Bååth, 2002. Structure and activity of the bacterial community in the rhizosphere of different plant species and the effect of arbuscular mycorrhizal colonisation. FEMS Microbiology Ecology 40: 223–231.
Stackebrandt, E., F. A. Rainey & N. L. Ward-Rainey, 1997. Proposal for a new hierarchic classification system, Actinobacteria classis nov. International Journal of Systematic Bacteriology 47: 479–491.
Vitousek, P. M., et al., 1997. Human domination of Earth’s ecosystems. Science 277: 494–499.
Wang, Q., et al., 2007. Naive Bayesian classifier for rapid assignment of rRNA sequences into the new bacterial taxonomy. Applied and Environmental Microbiology 73: 5261–5267.
Wieland, G., R. Neumann & H. Backhaus, 2001. Variation of microbial communities in soil, rhizosphere, and rhizoplane in response to crop species, soil type, and crop development. Applied and Environmental Microbiology 67: 5849–5854.
Windham, L. & J. G. Ehrenfeld, 2003. Net impact of a plant invasion on nitrogen-cycling processes within a brackish tidal marsh. Ecological Applications 13: 883–896.
Wu, J. Y., et al., 2010. Effects of polymerase, template dilution and cycle number on PCR based 16 S rRNA diversity analysis using the deep sequencing method. BMC Microbiology 10: 255.
Xiao, Y., et al., 2007. Chemical composition of Hydrilla verticillata (L. f.) Royle in Taihu Lake. Chinese Journal of Chemistry 25: 661–665.
Yao, H., et al., 2000. Microbial biomass and community structure in a sequence of soils with increasing fertility and changing land use. Microbial Ecology 40: 223–237.
Zwart, G., et al., 2002. Typical freshwater bacteria: an analysis of available 16S rRNA gene sequences from plankton of lakes and rivers. Aquatic Microbial Ecology 28: 141–155.
Acknowledgments
This project was supported by funding from the National Science Foundation HBCU-RISE Program (NSF Grant No. HRD-0932137). We are grateful to Scott Savery from the Wakulla Spring State Park and Scott Barrett Dyer from the Florida Department of Environmental Protection for their help with sample collection from the Wakulla Spring State Park.
Author information
Authors and Affiliations
Corresponding author
Additional information
Handling Editor: Stefano Amalfitano.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Gordon-Bradley, N., Li, N. & Williams, H.N. Bacterial community structure in freshwater springs infested with the invasive plant species Hydrilla verticillata . Hydrobiologia 742, 221–232 (2015). https://doi.org/10.1007/s10750-014-1988-0
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10750-014-1988-0