Abstract
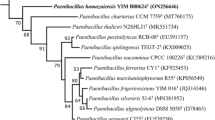
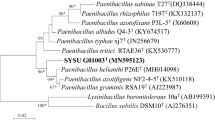
A novel strain, designated strain CSA42T, was isolated from the tomb of emperor Yang of Sui in Yangzhou, Jiangsu province, China. Strain CSA42T was observed to be Gram-stain positive, strictly aerobic, rod-shaped, spore-forming and motile. The optimum conditions for growth were found to be 30 °C, pH 8.0 and without NaCl. Phylogenetic analysis, based on 16S rRNA gene sequences, revealed strain CSA42T to be closely related to Paenibacillus larvae DSM 7030T (94.7%), Paenibacillus doosanensis CAU 1055T (94.4%) and Paenibacillus gansuensis B518T (94.2%). The major cellular fatty acids were identified as anteiso-C15:0, anteiso-C17:0 and iso-C16:0. MK-8 was found to be the only respiratory quinone. The polar lipids were found to be comprised of diphosphatidylglycerol, phosphatidylethanolamine, phosphatidylglycerol and two aminophospholipids. The cell wall peptidoglycan was found to contain meso-diaminopimelic acid and ribose as the only whole cell sugar. The genomic G+C content of strain CSA42T was determined to be 47.6 mol%. The low DNA–DNA relatedness values between strain CSA42T and the reference strain P. larvae KACC 11540T and many phenotypic properties support the classification of strain CSA42T (=KACC 18941T =CCTCC AB 2016201T) as the type strain of a novel species of the genus Paenibacillus, for which the name Paenibacillus tumbae sp. nov. is proposed. An emended description of the genus Paenibacillus based on the new data is also given.

Similar content being viewed by others
References
Achouak W, Normand P, Heulin T (1999) Comparative phylogeny of rrs and nifH genes in the Bacillaceae. Int J Syst Bacteriol 49:961–967
Ash C, Priest FG, Collins MD (1993) Molecular identification of rRNA group 3 bacilli (Ash, Farrow, Wallbanks and Collins) using a PCR probe test. Proposal for the creation of a new genus Paenibacillus. Antonie Van Leeuwenhoek 64:253–260
Ash C, Priest FG, Collins MD (1994) Paenibacillus gen. nov. In Validation of the publication of new names and new combinations previously effectively published outside the IJSB. List no. 51. Int J Syst Bacteriol 44:852
Berge O, Guinebretiere MH, Achouak W, Normand P, Heulin T (2002) Paenibacillus graminis sp. nov. and Paenibacullus odorifer sp. nov., isolated from plant roots, soil and food. Int J Syst Evol Microbiol 52:607–616
Cao Y, Chen F, Li Y, Wei S, Wang G (2015) Paenibacillus ferrarius sp. nov., isolated from iron mineral soil. Int J Syst Evol Microbiol 65:165–170
Chung YR, Kim CH, Hwang I, Chun J (2000) Paenibacillus koreensis sp. nov., a new species that produces an iturin-like antifungal compound. Int J Syst Evol Microbiol 50:1495–1500
Collins MD (1985) Isoprenoid quinone analysis in classification and identification. In: Goodfellow M, Minnikin DE (eds) Chemical methods in bacterial systematics. Academic Press, London, pp 267–287
De Ley J, Cattoir H, Reynaerts A (1970) The quantitative measurement of DNA hybridization from renaturation rates. Eur J Biochem 12:133–142
Felsenstein J (1985) Confidence limits on phylogenies: an approach using the bootstrap. Evolution 39:783–791
Fitch WM (1971) Toward defining the course of evolution: minimum change for a specific tree topology. Syst Zool 20:406–416
Gordon RE, Barnett DA, Handerhan JE, Pang CH-N (1974) Nocardia coeliaca, Nocardia autotrophica, and the nocardin strain. Int J Syst Bacteriol 24:54–63
Guo GN, Zhou X, Chen ZL, Yang ZW, Li XD, Li YH (2016a) Paenibacillus marchantiophytorum sp. nov., isolated from the liverwort Herbertus sendtneri. Int J Syst Evol Microbiol 66:755–761
Guo X, Zhou S, Wang Y-W, Wang H-M, Kong D-L, Zhu J, Dong W-W, He M-X, Zhao B-Q, Hu G-Q, Ruan Z-Y (2016b) Paenibacillus salinicaeni sp. nov., isolated from saline silt sample. Antonie Van Leeuwenhoek 109:721–728
Huang Z, Dai W, Zhou Z, Wang G, Lin G, Yan X, Zhao F (2016) Paenibacillus terreus sp. nov., isolated from forest soil. Int J Syst Evol Microbiol 66:243–247
Jin H-J, Lv J, Chen S-F (2011) Paenibacillus sophorae sp. nov., a nitrogen-fixing species isolated from the rhizosphere of Sophora japonica. Int J Syst Evol Microbiol 61:767–771
Kim OS, Cho YJ, Lee K, Yoon SH, Kim M, Na H, Park SC, Jeon YS, Lee JH, Yi H, Won S, Chun J (2012) Introducing EzTaxon-e: a prokaryotic 16S rRNA gene sequence database with phylotypes that represent uncultured species. Int J Syst Evol Microbiol 62:716–721
Kim J-H, Kang H, Kim W (2014a) Paenibacillus doosanensis sp. nov., isolated from soil. Int J Syst Evol Microbiol 64:1271–1277
Kim M, Oh HS, Park SC, Chun J (2014b) Towards a taxonomic coherence between average nucleotide identity and 16S rRNA gene sequence similarity for species demarcation of prokaryotes. Int J Syst Evol Microbiol 64:346–351
Kimura M (1980) A simple method for estimating evolutionary rates of base substitutions through comparative studies of nucleotide sequences. J Mol Evol 16:111–120
Kämpfer P, Falsen E, Lodders N, Martin K, Kassmannhuber J, Busse H-J (2012) Paenibacillus chartarius sp. nov., isolated from a paper mill. Int J Syst Evol Microbiol 62:1342–1347
Lee SD (2016) Paenibacillus cavernae sp. nov., isolated from soil of a natural cave. Int J Syst Evol Microbiol 66:598–603
Lee JC, Kim CJ, Yoon KH (2011) Paenibacillus telluris sp. nov., a novel phosphate-solubilizing bacterium isolated from soil. J Microbiol 49:617–621
Lee HW, Roh SW, Yim KJ, Shin NR, Lee J, Whon TW, Kim JY, Hyun DW, Kim D, Bae JW (2013) Paenibacillus marinisediminis sp. nov., a bacterium isolated from marine sediment. J Microbiol 51:312–317
Li J, Lu Q, Liu T, Zhou S, Yang G, Zhao Y (2014) Paenibacillus guangzhouensis sp. nov., an Fe(III)- and humus-reducing bacterium from a forest soil. Int J Syst Evol Microbiol 64:3891–3896
Lim J-M, Jeon CO, Lee J-C, Xu L-H, Jiang C-L, Kim C-J (2006) Paenibacillus gansuensis sp. nov., isolated from desert soil of Gansu province in China. Int J Syst Evol Microbiol 56:2131–2134
Logan NA, Berge O, Bishop AH, Busse H-J, De Vos P, Fritze D, Heyndrickx M, Kämpfer P, Rabinovitch L, Salkinoja-Salonen MS, Seldin L, Ventosa A (2009) Proposed minimal standards for describing new taxa of aerobic, endospore-forming bacteria. Int J Syst Evol Microbiol 59:2114–2121
Ma Y-C, Chen S-F (2008) Paenibacillus forsythia sp. nov., a nitrogen-fixing species isolated from rhizosphere soil of Forsythia mira. Int J Syst Evol Microbiol 58:319–323
Ma Y, Zhang J, Chen S (2007) Paenibacillus zanthoxyli sp. nov., a novel nitrogen-fixing species isolated from the rhizosphere of Zanthoxylum simulans. Int J Syst Evol Microbiol 57:873–877
Mesbah M, Premachandran U, Whitman WB (1989) Precise measurement of the G+C content of deoxyribonucleic acid by high performance liquid chromatography. Int J Syst Bacteriol 39:159–167
Mishra AK, Lagier J-C, Rivet R, Raoult D, Fournier P-E (2012) Non-contiguous finished genome sequence and description of Paenibacillus senegalensis sp. nov. Stand Genomic Sci 7:70–81
Moon J, Kim J (2014) Isolation of Paenibacillus pinesoli sp. nov. from forest soil in Gyeonggi-Do, Korea. J Microbiol 52:273–277
Rameshkumar N, Lang E, Tanaka N (2016) Description of Vogesella oryzae sp. nov., isolated from the rhizosphere of saline tolerant pokkali rice. Syst Appl Microbiol 39:20–24
Rhuland LE, Work E, Denman RF, Hoare DS (1955) The behavior of the isomers of α, ε-diaminopimelic acid on paper chromatograms. J Am Chen Soc 77:4844–4846
Romanenko LA, Tanaka N, Svetashev VI, Kalinovskaya NI (2013) Paenibacillus profundus sp. nov., a deep sediment bacterium that produces isocoumarin and peptide antibiotics. Arch Microbiol 195:247–254
Saitou N, Nei M (1987) The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol Biol Evol 4:406–425
Sasser M (1990) Identification of bacteria by gas chromatography of cellular fatty acids. MIDI technical note 101. MIDI Inc, Newark
Shen P, Fan XR, Li GW (1999) Experiment of microbiology. Higher Education Press, Beijing
Shida O, Takagi H, Kadowaki K, Nakamura LK, Komagata K (1997) Transfer of Bacillus alginolyticus, Bacillus chondroitinus, Bacillus curdlanolyticus, Bacillus glucanolyticus, Bacillus kobensis, and Bacillus thiaminolyticus to the genus Paenibacillus and emended description of the genus Paenibacillus. Int J Syst Bacteriol 47:289–298
Stackebrandt E, Goebel BM (1994) Taxonomic note: a place for DNA–DNA reassociation and 16S rRNA sequence analysis in the present species definition in bacteriology. Int J Syst Bacteriol 44:846–849
Sukweenadhi J, Kim Y-J, Lee KJ, Koh S-C, Hoang V-A, Nguyen N-L, Yang D-C (2014) Paenibacillus yonginensis sp. nov., a potential plant growth promoting bacterium isolated from humus soil of Yongin forest. Antonie Van Leeuwenhoek 106:935–945
Švec P, Vancanneyt M, Seman M, Snauwaert C, Lefebvre K, Sedláček I, Swings J (2005) Evaluation of (GTG)5-PCR for identification of Enterococcus spp. FEMS Microbiol Lett 247:59–63
Tamura K, Stecher G, Peterson D, Filipski A, Kumar S (2013) MEGA 6: molecular evolutionary genetics analysis version 6.0. Mol Biol Evol 30:2725–2729
Tindall BJ (1990a) A comparative study of the lipid composition of Halobacterium saccharovorum from various sources. Syst Appl Microbiol 13:128–130
Tindall BJ (1990b) Lipid composition of Halobacterium lacusprofundi. FEMS Microbiol Lett 66:199–202
Van der Maarel MJEC, Veen A, Wijbenga DJ (2000) Paenibacillus granivorans sp. nov., a new Paenibacillus species which degrades native potato starch granules. Syst Appl Microbiol 23:344–348
Wang L, Baek SH, Cui Y, Lee HG, Lee ST (2012) Paenibacillus sediminis sp. nov., a xylanolytic bacterium isolated from a tidal flat. Int J Syst Evol Microbiol 62:1284–1288
Wang LY, Li J, Li QX, Chen SF (2013) Paenibacillus beijingensis sp. nov., a nitrogen-fixing species isolated from wheat rhizosphere soil. Antonie Van Leeuwenhoek 104:675–683
Wayne LG, Brenner DJ, Colwell RR, Grimont PAD, Kandler O, Krichevsky MI, Moore LH, Moore WEC, Murray RGE, Stackebrandt E, Starr MP, Trüper HG (1987) Report of the ad-hoc-committee on reconciliation of approaches to bacterial systematic. Int J Syst Bacteriol 37:463–464
Xiang W, Wang G, Wang Y, Yao R, Zhang F, Wang R, Wang D, Zheng S (2014) Paenibacillus selenii sp. nov., isolated from selenium mineral soil. Int J Syst Evol Microbiol 64:2662–2667
Zhou Y, Gao S, Wei D-Q, Yang L-L, Huang X, He J, Zhang Y-J, Tang S-K, Li W-J (2012) Paenibacillus thermophiles sp. nov., a novel bacterium isolated from a sediment of hot spring in Fujian Province, China. Antonie Van Leeuwenhoek 102:601–609
Acknowledgements
The authors thank Dr Soon-Wo Kwon, Korean Agricultural Culture Collection (KACC), for kindly providing strain Paenibacillus larvae KACC 11540T. The authors also thank the editor and reviewers for giving many suggestions to improve this manuscript. This work was supported by National Nature Science Foundation of China (41201251, 41101229 and 51478103).
Author information
Authors and Affiliations
Corresponding authors
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Huang, Z., Zhao, F. & Li, YH. Isolation of Paenibacillus tumbae sp. nov., from the tomb of the emperor Yang of the Sui dynasty, and emended description of the genus Paenibacillus . Antonie van Leeuwenhoek 110, 357–364 (2017). https://doi.org/10.1007/s10482-016-0807-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10482-016-0807-1