Abstract
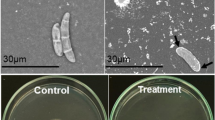
Bacillus thuringiensis is an insecticidal bacterium whose chitinolytic system may be exploited to improve the insecticidal system of Bt-crops. A nucleotide fragment of 1368 bp from B. thuringiensis serovar konkukian S4, containing the complete coding sequence of the chitin binding protein Cbp50, was cloned and sequenced. Analyses have shown the protein to contain a modular structure consisting of an N-terminal CBM33 domain, two copies of a fibronectin-like domain and a C-terminal chitin binding domain classified as CBM5. The Cbp50 protein was heterologously expressed in Escherichia coli, purified and assessed for chitin binding activity. A deletion mutant (CBD-N; containing only the N-terminal CBM33 domain) of Cbp50 was produced to determine the role of C-terminal domains in the binding activity of the protein. The full-length Cbp50 was shown to bind β-chitin most efficiently followed by α-chitin, colloidal chitin and cellulose. The polysaccharide binding activity of CBD-N was drastically decreased. The data demonstrate that both the N-terminal and C-terminal domains of Cbp50 are essential for the efficient binding of chitin. The purified Cbp50 showed antifungal activity against the phytopathogenic fungus Fusarium oxysporum and the opportunistic human pathogen Aspergillus niger. This is the first report of a modular chitin binding protein in bacteria.



Similar content being viewed by others
References
Barboza-Corona JE, Nieto-Mazzocco E, Velazquez-Robledo R, Salcedo-Hernandez R, Bautista M, Jimenez B, Ibarra JE (2003) Cloning, sequencing, and expression of the chitinase gene chiA74 from Bacillus thuringiensis. Appl Environ Microbiol 69(2):1023–1029
Bormann C, Baier D, Horr I, Raps C, Berger J, Jung G, Schwarz H (1999) Characterization of a novel, antifungal, chitin-binding protein from Streptomyces tendae Tu901 that interferes with growth polarity. J Bacteriol 181(24):7421–7429
Carrard G, Koivula A, Soderlund H, Beguin P (2000) Cellulose-binding domains promote hydrolysis of different sites on crystalline cellulose. Proc Natl Acad Sci USA 97(19):10342–10347
Chae KS, Lee IH, Choi CS, Kim HR (1999) Purification and characterization of chitin-binding proteins from the hemolymph of sweet potato hornworm, Agrius convolvuli. Comp Biochem Physiol B 124(4):475–481
Chamoy L, Nicolai M, Ravaux J, Quennedey B, Gaill F, Delachambre J (2001) A novel chitin-binding protein from the vestimentiferan Riftia pachyptila interacts specifically with beta-chitin cloning, expression, and characterization. J Biol Chem 276(11):8051–8058
Chu HH, Hoang V, Hofemeister J, Schrempf H (2001) A Bacillus amyloliquefaciens ChbB protein binds beta- and alpha-chitin and has homologues in related strains. Microbiology 147(7):1793–1803
De Bolle MF, David KM, Rees SB, Vanderleyden J, Cammue BP, Broekaert WF (1993) Cloning and characterization of a cDNA encoding an antimicrobial chitin-binding protein from amaranth, Amaranthus caudatus. Plant Mol Biol 22(6):1187–1190
Hashimoto M, Ikegami T, Seino S, Ohuchi N, Fukada H, Sugiyama J, Shirakawa M, Watanabe T (2000) Expression and Characterization of the Chitin-Binding Domain of Chitinase A1 from Bacillus circulans WL-12. J Bacteriol 182:3045–3054
Huang X, Xie W, Gong Z (2000) Characteristics and antifungal activity of a chitin binding protein from Ginkgo biloba. FEBS Lett 478(1–2):123–126
Kolbe S, Fischer S, Becirevic A, Hinz P, Schrempf H (1998) The Streptomyces reticuli alpha-chitin-binding protein CHB2 and its gene. Microbiology 144(5):1291–1297
Lehtio J, Sugiyama J, Gustavsson M, Fransson L, Linder M, Teeri TT (2003) The binding specificity and affinity determinants of family 1 and family 3 cellulose binding modules. Proc Natl Acad Sci USA 100(2):484–489
Mehmood MA, Xiao X, Hafeez FY, Yingbao G, Wang F (2009) Purification and characterization of a chitinase fom Serratia proteamaculans. World J Microbiol Biotechnol 25:1955–1961
Mehmood MA, Xiao X, Hafeez FY, Gai Y, Wang F (2010) Molecular characterization of an endochitinase from Bacillus thuringiensis subsp. konkukian. World J Microbiol Biotechnol. doi:10.1007/s11274-010-0401-z
Miller G (1959) Use of dinitrosalicylic acid reagent for detection of reducing sugar. Anal Chem 31:426–428
Nielsen KK, Nielsen JE, Madrid SM, Mikkelsen JD (1997) Characterization of a new antifungal chitin-binding peptide from sugar beet leaves. Plant Physiol 113(1):83–91
Osborn RW, De Samblanx GW, Thevissen K, Goderis I, Torrekens S, Van Leuven F, Attenborough S, Rees SB, Broekaert WF (1995) Isolation and characterization of plant defensins from seeds of Asteraceae, Fabaceae, Hippocastanaceae and Saxifragaceae. FEBS Lett 368:257–262
Perrakis A, Tews I, Dauter Z, Oppenheim AB, Chet I, Wilson KS, Vorgias CE (1994) Crystal structure of a bacterial chitinase at 2.3 A resolution. Structure 2(12):1169–1180
Pleban S, Chernin L, Chet I (1997) Chitinolytic activity of an endophytic strain of Bacillus cereus. Lett Appl Microbiol 25(4):284–288
Regev A, Keller M, Strizhov N, Sneh B, Prudovsky E, Chet I, Ginzberg I, Koncz-Kalman Z, Koncz C, Schell J, Zilberstein A (1996) Synergistic activity of a Bacillus thuringiensis delta-endotoxin and a bacterial endochitinase against Spodoptera littoralis larvae. Appl Environ Microbiol 62(10):3581–3586
Saito A, Fujii T, Yoneyama T, Redenbach M, Ohno T, Watanabe T, Miyashita K (1999) High-multiplicity of chitinase genes in Streptomyces coelicolor A3(2). Biosci Biotechnol Biochem 63(4):710–718
Schnellmann J, Zeltins A, Blaak H, Schrempf H (1994) The novel lectin-like protein CHB1 is encoded by a chitin-inducible Streptomyces olivaceoviridis gene and binds specifically to crystalline alpha-chitin of fungi and other organisms. Mol Microbiol 13(5):807–819
Schrempf H (1999) Characteristics of chitin-binding proteins from Streptomycetes. EXS 87:99–108
Selitrennikoff CP (2001) Antifungal proteins. Appl Environ Microbiol 67(7):2883–2894
Shimahara K, Takiguchi Y (1988) Preparation of crustacean chitin. Methods Enzymol 161:417–423
Tormo J, Lamed R, Chirino AJ, Morag E, Bayer EA, Shoham Y, Steitz TA (1996) Crystal structure of a bacterial family-III cellulose-binding domain: a general mechanism for attachment to cellulose. EMBO J 15(21):5739–5751
Uchiyama T, Katouno F, Nikaidou N, Nonaka T, Sugiyama J, Watanabe T (2001) Roles of the exposed aromatic residues in crystalline chitin hydrolysis by chitinase A from Serratia marcescens 2170. J Biol Chem 276(44):41343–41349
Vaaje-Kolstad G, Horn SJ, van Aalten DM, Synstad B, Eijsink VG (2005a) The non-catalytic chitin-binding protein CBP21 from Serratia marcescens is essential for chitin degradation. J Biol Chem 280(31):28492–28497
Vaaje-Kolstad G, Houston DR, Riemen AH, Eijsink VG, van Aalten DM (2005b) Crystal structure and binding properties of the Serratia marcescens chitin-binding protein CBP21. J Biol Chem 280(12):11313–11319
Vaaje-Kolstad G, Westereng B, Horn SJ, Liu Z, Zhai H, Sørlie M, Eijsink VG (2010) An oxidative enzyme boosting the enzymatic conversion of recalcitrant polysaccharides. Science 330(6001):219–222
van Aalten DM, Synstad B, Brurberg MB, Hough E, Riise BW, Eijsink VG, Wierenga RK (2000) Structure of a two-domain chitotriosidase from Serratia marcescens at 1.9-A resolution. Proc Natl Acad Sci USA 97(11):5842–5847
Zeltins A, Schrempf H (1995) Visualization of alpha-chitin with a specific chitin-binding protein (CHB1) from Streptomyces olivaceoviridis. Anal Biochem 231(2):287–294
Zeltins A, Schrempf H (1997) Specific interaction of the Streptomyces chitin-binding protein CHB1 with alpha-chitin–the role of individual tryptophan residues. Eur J Biochem 246(2):557–564
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Mehmood, M.A., Xiao, X., Hafeez, F.Y. et al. Molecular characterization of the modular chitin binding protein Cbp50 from Bacillus thuringiensis serovar konkukian . Antonie van Leeuwenhoek 100, 445–453 (2011). https://doi.org/10.1007/s10482-011-9601-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10482-011-9601-2