Abstract
As human populations in Africa expand, humans encroach and modify wildlife habitats for farming, fishing, tourism, or settlement. Anthropogenic activities in shared environments may promote transmission of zoonotic pathogens between humans, domestic animals, and wildlife. Between July 2012 and February 2014, we evaluated Salmonella prevalence, serovars, genotypes, and antibiotic resistant phenotypes in resident and migratory birds utilizing human-impacted habitats in northwestern Lake Victoria and protected habitats in Queen Elisabeth National Park. Salmonella occurrence in the urban environment was assessed by sampling storm-water and wastewater from a channel that drains Kampala City into Lake Victoria. Salmonella was detected in 4.3% pooled bird fecal samples, and 57.1% of environmental samples. While birds in impacted and protected areas shared serovars, the genotypes were distinct. We found distinct strains in birds and the environment suggesting some strains in birds are host adapted, and strains circulating in the environment may not necessarily disseminate to birds. Conversely, birds in both impacted and protected areas shared strains with the urban environment, suggesting Salmonella disseminates between impacted environments and birds across sites. Overall, more strains were observed in the urban environment compared to birds, and poses risk of Salmonella reemergence in birds and transmission across species and space.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Populations of some Afro-Palearctic migratory birds have been declining in Europe for decades (Sanderson et al. 2006). In Uganda, a decline in wild bird (hereafter bird) populations has been observed since 2004 (Pomeroy and Asasira 2011). While the exact cause of these declines is unclear, several interconnected factors may contribute including increasing human population resulting in habitat loss, increasing climatic variability affecting breeding and wintering grounds, and decreasing food availability in key habitats (Sanderson et al. 2006; Vickery et al. 2014). Another factor is the potential for pathogen introduction into bird populations when humans encroach and modify wildlife habitats into agricultural, fishing, or urban areas. Increasing habitat overlap may promote pathogen transmission between humans and wildlife (Daszak et al. 2000). Also, multiple migratory bird species from various breeding grounds congregate at higher densities at stopover and wintering grounds and intermingle with resident birds. Such phenomena may promote pathogen exposure and transmission across species and space (Altizer et al. 2011).
Zoonotic pathogens can threaten bird populations sharing habitats with humans. Non-typhoidal Salmonella affects humans worldwide (Majowicz et al. 2010) and multiple animal hosts, and causes morbidity and mortality in birds under natural and human influenced conditions (Pennycott et al. 1998; Velarde et al. 2012; Giovannini et al. 2013). For example, provisioning has been shown to attract high bird densities at feeding sites leading to fecal contamination, pathogen transmission, and reemergence of salmonellosis (Pennycott et al. 1998). Salmonella is also frequently isolated from apparently healthy scavenging birds that are attracted to refuse pits and human sewage outfalls (Benskin et al. 2009). Asymptomatic birds may disseminate Salmonella to susceptible individuals through fecal shedding, shared environments, and via direct contact.
Few studies have compared Salmonella prevalence across diverse bird species under non-epidemic conditions (Hernandez et al. 2003); urban and rural settings (Hamer et al. 2012); or different habitat protection types. These studies have mostly been conducted at European and North American sites (Pennycott et al. 2006; Hughes et al. 2008; Hamer et al. 2012). Studies at African sites are important because Afro-Palearctic migratory birds spend a considerable part of their annual cycle in Africa. For instance, Uganda has wetland habitats of international importance for resident and migratory birds (Byaruhanga and Nalwanga 2006). The bird habitats in northwestern Lake Victoria are closely associated with Kampala city, and evidence of microbial contamination in the urban environment has been documented (Afema et al. 2016). There are calls for research and conservation efforts to understand and mitigate factors causing Afro-Palearctic migrant bird population declines (Vickery et al. 2014).
This study investigated whether birds utilizing human-impacted habitats in northwestern Lake Victoria had higher risk of acquiring Salmonella compared to birds in Queen Elisabeth national park (QENP). The study used noninvasive methods to collect fecal samples from birds. Salmonella occurrence in the urban environment was assessed by sampling storm-water and wastewater from a channel that drains Kampala City into Lake Victoria. This study describes Salmonella occurrence, antibiotic-resistant phenotypes, serovar distribution and genotypic structure of common serovars, and infer transmission.
Materials and Methods
Study Area
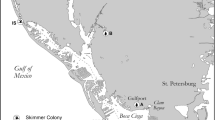
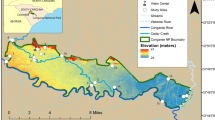
This study was conducted in Lake Victoria region and in QENP (Fig. 1). Lutembe Bay and Nakiwogo are important bird areas in northwestern Lake Victoria (LVBA). Lutembe Bay is located on outer Murchison Bay and is spatially close to Kampala, the capital and largest city in Uganda. Lutembe Bay has many small muddy islets surrounded by shallow waters that birds use for roosting (Byaruhanga and Nalwanga 2006). Nakiwogo is located close to Entebbe town. It covers a greater geographic area and has less human impact compared to Lutembe Bay and birds roost on rocky outcrops in the lake and the shorelines. Fishing communities exist at both Lutembe Bay and Nakiwogo.
Map of Uganda showing study area. Inset on the upper right is northwestern Lake Victoria showing spatial relationship between Lutembe Bay, Nakiwogo, Nakivubo Channel (NC), Murchison Bay (MB), and the main cities. The Queen Elisabeth (QENP) sampling sites (Bwenda Bay and Kazinga Channel) and their spatial relationships are shown on the lower right. Maps obtained from: ArcGIS\Packages\0ef9b6b4700942ef9adaecfb0b3857cd\commondata\data0\Ug_Districtboundaries2006.shp. http://services2.arcgis.com/D6cgxWzepFC1quJH/arcgis/rest/services/Uganda_Rivers/FeatureServer. ArcGIS\Packages\2ac6a57b073749d4b2ab2ba90efabb55\commondata\data0\Ug_Waterbodies.shp.
Nakivubo Channel (NC) is approximately 12-km-long channel network constructed to drain storm-water from Kampala City. This channel also receives secondary effluent from wastewater treatment plant, and abattoir run-off and it empties into inner Murchison Bay in Lake Victoria. Impairment of water quality because of high fecal coliform counts has been documented in NC and Murchison Bay (Kansiime and van Bruggen 2001).
QENP is a savanna park located in western Uganda. Human activity in the park is restricted to tourism, staff and permitted fishing villages. Bwenda Bay is located on Lake Edward close to Katwe town. The bay has shallow waters that are shielded from the lake by wetland plants. Kazinga Channel is a 33 km stretch of water that connects Lake George and Lake Edward. Kazinga Channel is popular for tourism and birds commonly roost on the southern shoreline. Fishing is permitted in some parts of the channel with a resident community at Kazinga fishing village.
Sampling Strategy and Sample Collection
This study employed a repeated cross-sectional sampling approach. From July 2012 to February 2013, 1 l of grab water samples from 3 points along NC was collected into sterile plastic bottles. From December 2012 to February 2013, fecal samples were collected from birds at Lutembe Bay, Nakiwogo, Bwenda Bay and Kazinga Channel. Birds were identified during sampling or photographed and identified later using a guidebook (Stevenson and Fanshawe 2006). A cotton-tipped swab was used to collect a single fresh fecal dropping on roosting grounds, and 5 fecal swabs were pooled into a 50-ml tube containing 45 ml Rappaport–Vasilliadis selective enrichment broth (R10, Hardy Diagnostics, USA). At each site, samples were collected 1–3 times per month. From October 2013 to February 2014, samples were collected once a month from all sites. Samples were placed in a cool box containing cold packs and processed within 24 h of collection at the Microbiology Laboratory, College of Veterinary Medicine, Makerere University or a field laboratory in Mweya, QENP.
Culture Methods
Samples in R10 were incubated at 42°C for 18–24 h, then 100 μl of R10 enriched broth was spread onto xylose lysine tergitol-4 (XLT-4) or xylose lysine deoxycholate (XLD) agar plates (Difco Laboratories) using sterile beads, and incubated at 37°C for 18–24 h. Based on selective plates indicating Salmonella growth, original enriched broths suspected to contain Salmonella were subjected to serial dilutions (Sanderson et al. 1995) and re-plated onto XLD or XLT-4 plates to obtain well-isolated colonies. An average of 9 suspect Salmonella colonies were picked from each plate and stabbed into brain heart infusion agar (Hardy Diagnostics, USA) banking tubes and stored at room temperature. Water samples from NC were processed using previously described methods (Berge et al. 2006) with minor modifications (Afema et al. 2016). All isolates were shipped to Washington State University (WSU) for further processing in compliance with Uganda National Council of Science and Technology materials transfer agreement with College of Veterinary Medicine, Makerere University.
Serotyping
Suspect Salmonella isolates were screened by PCR amplification of the invA gene (Yoshida et al. 2007). PCR-positive samples were then evaluated using a genovar method and assigned a genovar code (Leader et al. 2009). All isolates were subsequently subjected to the Kauffman–White serotyping scheme (Popoff et al. 2004).
Multiple Locus Variable Number Tandem Repeats Analysis (MLVA)
Salmonella Typhimurium were genotyped using MLVA (Lindstedt et al. 2003, 2004) according to PulseNet, Centers for Disease Control and Prevention protocol. Briefly, DNA was prepared as boiled cell lysate and seven variable number tandem repeats (VNTR) loci (ST3, ST5, ST7, STTR10pl, ST2, ST6, and ST8) were amplified. The PCR products were separated by capillary electrophoresis at the WSU Laboratory for Biotechnology and Bioanalysis. The copy number (allele) for each VNTR locus was determined as follows: [(observed fragment size − offset)/repeat size]. Based on alleles at the 7 loci, isolates were assigned as a unique MLVA type. The data were analyzed using PHYLOViZ software (Francisco and Vaz 2012).
Pulsed Field Gel Electrophoresis (PFGE)
We used PulseNet PFGE protocols (Ribot et al. 2006) to subtype non-S. Typhimurium serovars recovered from at least two sampling sites. DNA was cleaved with restriction endonuclease XbaI (Fermentas, Thermo Fisher Scientific). PFGE bands were assigned and analyzed using BioNumerics 6.6 software (Applied Maths, Austin, TX, USA). The Dice coefficient was used to assess band pattern similarity and isolates with ≥90% similarity index were considered to be closely related. Cluster analysis was performed by the unweighted pair group method with arithmetic mean, band matching tolerance of 2%, and relaxed doublet matching.
Antibiotic Resistance Tests
Isolates were tested for resistance to a panel of 15 antibiotics (amikacin, 30 μg; amoxicillin/clavulanic acid, 20/10 μg; ampicillin, 10 μg; cefotaxime, 30 μg; cefoxitin, 30 μg; ceftiofur, 30 μg; chloramphenicol, 30 μg; ciprofloxacin, 5 μg; gentamicin, 10 μg; kanamycin, 30 μg; nalidixic acid, 10 μg; streptomycin, 10 μg; sulfisoxazole, 250 μg; tetracycline, 30 μg; and trimethoprim/sulfamethoxazole, 1.25/23.75 μg;) using a disk diffusion assay (Bauer et al. 1966). Isolates were classified as resistant or susceptible to each antibiotic using Clinical and Laboratory Standards Institute definitions for Enterobacteriaceae (Watts 2008) except for streptomycin where the National Antimicrobial Resistance Monitoring Systems breakpoint was used (CDC 2013). Isolates with intermediate resistance were interpreted as susceptible. A single resistance profile was created for each isolate by concatenating resistance to 15 antibiotics.
Results
Birds Observed During Sample Collection
The species observed during sample collection are shown in Table 1. We mostly observed mixed species flocks; therefore, the species from which a fecal sample was collected could not be ascertained. The species that were present at all sites during most sampling occasions include black-winged stilt, little egret, gull-billed tern, grey heron, little ringed plover, Egyptian geese, and grey-headed gull. African skimmers were observed only at Kazinga Channel during all sampling occasions. Pied avocet, lesser and greater flamingos were only observed at Bwenda Bay. The common Squacco heron, long-toed lapwing, saddle-billed stork, and black-tailed Godwit were only observed at Lutembe, while black kite, barn swallow, whiskered tern, and yellow wagtail were only observed at Nakiwogo area.
Salmonella Detection in Bird Feces and NC Water
Salmonella was isolated from 17 of 395 pooled bird fecal samples (Table 2). The prevalence was similar across impacted and protected sites. In contrast, Salmonella was far more common (57.1%) in NC water. From each positive pooled fecal sample and water sample, an average of 7 (1–12) and 10 (3–16) PCR confirmed Salmonella isolates were obtained, respectively. A total of 121 isolates from birds and 166 from NC water were used in subsequent analyses.
Serovar Distribution
From the 287 Salmonella isolates, 18 serovars were identified (Fig. 2). No single serovar was detected across all five sampling sites. Salmonella Chandans, Salmonella Heidelberg, Salmonella Newport, and Salmonella Stanleyville were shared across three sampling sites and were always present at Lutembe Bay and NC. Salmonella Newport was shared between birds in LVBA and NC water, but not birds at QENP. All serovars found in birds at LVBA also occurred in NC. Serovars unique to sites were also detected.
Multiple Locus Variable Number Tandem Repeats Analysis
We performed MLVA on 29 S. Typhimurium isolates from birds at Nakiwogo and 7 isolates from NC water and detected 10 genotypes. A minimum spanning tree of the genotypes was constructed using the goeBURST algorithm to observe genotypic relationships. At the single locus variant level, the genotypes from birds clustered into 3 clonal complexes, and each complex was composed of isolates from a pooled sample (Fig. 3a). With a full goeBURST algorithm, two bird-associated complexes from samples collected within 3 months clustered together and were distinct from another bird associated complex from a sample collected about 9 months earlier (Fig. 3b). The genotypes from NC differed from each other and the bird genotypes by at least 4 alleles.
a, b Minimum spanning tree of S. Typhimurium MLVA genotypes constructed using the goeBURST algorithm. There are 10 genotypes represented by blue (birds) or red (NC water) circles. The size of the circle is proportional to the number of isolates on a log scale. The number inside the circle is the assigned MLVA type. The 7 genotypes from bird isolates cluster into 3 clonal complexes at the single locus variant level (a). A full tree with all the genotypes connected and the number of locus variants along the connecting paths is shown in b.
Pulsed Field Gel Electrophoresis
We performed PFGE on selected isolates from 8 serovars recovered from at least 2 sampling sites (Table 3). When we compared PFGE patterns of serovars shared between bird sampling sites, there was little evidence of shared PFGE types across sites. For instance, S. Chandans and S. Heidelberg were shared by birds at Lutembe Bay and Kazinga Channel but had distinct PFGE patterns. Similarly, S. Stanleyville was common to birds at Lutembe Bay and Bwenda Bay but had distinct PFGE patterns. Nonetheless, S. Newport with similar PFGE types was observed in birds at Lutembe Bay and Nakiwogo.
When we compared PFGE patterns in serovars common to birds and NC, interestingly, similar PFGE types were shared by birds at each site and NC. For example, S. Chandans, S. Senftenberg, and S. Stanleyville with indistinguishable PFGE profiles (B, J, and N, respectively) were common to Lutembe Bay birds and NC. Also, S. Newport with similar PFGE types was recovered from NC, and birds at Lutembe Bay and Nakiwogo. Although S. Newport, S. Senftenberg, and S. Stanleyville with similar PFGE types were found in Lutembe Bay birds and NC, other PFGE types were exclusive to NC (H, K, L, and Q). Furthermore, although S. Heidelberg was common to Lutembe Bay birds and NC, distinct PFGE patterns were observed in birds (E) and NC (C).
We had expected to observe PFGE differences between QENP birds and NC, but shared serovars between NC and Bwenda Bay (S. II 42:r:- and S. Stanleyville), and NC and Kazinga Channel (S. Heidelberg and S. Zanzibar) had similar PFGE patterns. The only shared serovar with PFGE pattern unique to protected area birds was S. Chandans.
Antibiotic Resistance
The Salmonella recovered from birds were either susceptible to all 15 antibiotics (pan-susceptible) or had single resistance to sulfisoxazole, except one S. Virchow isolate from Lutembe Bay with a multi-drug-resistant profile (Table 4). In NC, 53% of the isolates were pan-susceptible, 19% were resistant to sulfisoxazole, and the remaining isolates were distributed across 17 profiles including multi-drug-resistant types. The multi-drug-resistant profiles were mostly associated with Salmonella Kentucky with streptomycin, sulfisoxazole, tetracycline, ciprofloxacin, and nalidixic acid resistance profile.
Discussion
Uganda has a high diversity of resident and migratory birds and several habitats that are crucial for their conservation (NatureUganda 2010). We assessed Salmonella shedding in apparently healthy water-birds exposed to habitats that varied in the impact of humans on those habitats. Because of the proximity of LVBA to Kampala city, and the relative remoteness of QENP from urban influence, we hypothesized LVBA birds had higher risk of acquiring and shedding environmental-source Salmonella than QENP birds. However, we observed no significant difference in Salmonella pooled prevalence between LVBA and QENP.
One explanation is that transmission of fecal parasites (including bacteria) is influenced by intensity of habitat use (Nunn et al. 2011). We sampled between October and February which coincides with the wintering season for Afro-Palearctic migrants. We observed high bird numbers at both QENP and LVBA, and although we were unable to estimate number of birds per unit area, the habitats were in intense use relative to off season when only resident birds exist. Modeling has shown that buildup of pathogens in the environment coupled with increased contact leads to increased prevalence in a population during seasonal sharing of resources (Nunn et al. 2014). Salmonella prevalence being similar across sampled sites may be directly related to high bird densities and species richness. A way to evaluate this is to determine prevalence when migrants are away. It is possible that when only residents exist, there is less buildup of fecal material, less density dependent transmission, and the environment can be ‘cleaned’ by pathogen decay.
Another explanation for the similar prevalence between sites is that although QENP has less human impact, there is still risk of introducing pathogens from fishing and pastoral communities within and bordering the park. A modeling study showed that continuous introduction of fecal-associated pathogens at the edge of a protected area led to spread within the entire population in the reserve (Nunn et al. 2014). A third explanation is that migrant birds are the source of the Salmonella we observed. Alternatively, birds in QENP share habitat with other wild animals, and Salmonella could be transmitted between different species.
Our measure of non-typhoidal Salmonella prevalence in birds was based on pooled samples. While pooling samples is more sensitive for detecting Salmonella compared to individual samples (Arnold et al. 2009, 2011), prevalence cannot be directly determined and instead lies within a range of values (0.2–1 in our study). Our measure has good internal validity for comparing pooled prevalence across sites but makes comparison to previous studies difficult. Nonetheless, our median prevalence inference is comparable to other studies.
Salmonella estimated prevalence in our study was greater compared to that in Afro-Palearctic migrants captured at Ottenby bird observatory during fall migration from continental Europe to West Africa (Hernandez et al. 2003) or birds in Greater Chicago (Hamer et al. 2012). In contrast, our prevalence is lower than reports of 15.9% (6.4–43.5%) fecal shedding in migrating cranes at Izumi plain in Japan (Kitadai et al. 2010). A possible explanation is that prevalence is influenced by feeding ecology. Birds in the Ottenby and Chicago studies were passerines where contact with fecal material is less likely, whereas ground roosting water-birds we observed come into contact with feces on the ground or in water, and are more likely to become infected in case of environmental contamination. This is supported by a survey of Campylobacter shedding in migratory birds at Ottenby, where higher prevalence occurred in birds that foraged on invertebrates on shorelines, opportunistic feeders, gregarious, and long-lived birds (Waldenstrom et al. 2002).
This study detected 18 different serovars across study sites and found a diverse array of bird associated serovars (n = 12) compared to previous studies that mainly found S. Typhimurium (Hughes et al. 2008; Kitadai et al. 2010). While prevalence in birds across sampling sites was not significantly different, the serovar distributions were different, and no serovars were shared between all sampling sites. There was similarity between serovars found in water from NC (which contains runoff from Kampala’s urban environment and human source wastewater) and spatially associated birds in LVBA. These sites shared 6 serovars of which 4 had similar PFGE patterns. Paradoxically, a similar pattern of serovar sharing was observed for QENP birds and NC with 5 shared serovars and 4 sharing PFGE patterns. These findings suggest birds across sites were exposed to and likely acquired strains from a human impacted environment. Interestingly, there were 3 serovars shared by QENP and LVBA birds, but none shared PFGE patterns, possibly due to different environments or bird species.
Environmental samples were not collected from QENP because there was no point source reflecting human impact equivalent to NC. While QENP habitats, including water, were assumed to be relatively pristine compared to Lake Victoria region, there are human-associated non-point source impacts because of fishing villages within the park. The sharing of serovars between NC and QENP birds may suggest that birds at QENP are exposed to human-source Salmonella. Alternatively, birds that are attracted to human waste may introduce Salmonella to bird areas. For instance, Black-winged stilts were observed at all sites on most sampling occasions, and at human waste stabilization ponds.
Salmonella Typhimurium was only detected in NC water and birds at Nakiwogo. Although it is cited as the most commonly recovered serovar (Reed et al. 2003; Hughes et al. 2008, 2010), it represented only 8.3% of the serovars we recovered. The genotypes in birds were distinct from those in NC, and most bird isolates were pan-susceptible whereas those from NC were multi-drug resistant. Despite the known wide host range of S. Typhimurium, our findings, consistent with other studies, corroborate the conclusion that bird strains are host adapted (Hughes et al. 2008, 2010).
Salmonella Stanleyville was common in our study shared between NC, Lutembe Bay (LVBA), and Bwenda Bay (QENP). In a related study, we found similar strains of S. Stanleyville in wastewater from a livestock slaughterhouse and human effluent from a wastewater treatment plant (Afema et al. 2016). These facilities discharge wastewater into NC and eventually into Lake Victoria. Finding similar strains of S. Stanleyville in Lutembe Bay birds and NC water suggests birds could have acquired them through environmental exposure. Finding similar strains of S. Stanleyville in NC water and birds in Bwenda Bay suggests birds that utilize protected and impacted areas could have been exposed to pathogens in urban environments. Alternatively, although human impact in protected areas may be low, human fecal pathogens may still circulate in the environment.
Salmonella Chandans was observed in birds from both conserved and impacted areas, but this serovar is rare in humans and livestock (Madden et al. 2007). As expected, identical PFGE and antibiotic resistance patterns occurred in LVBA and NC but were distinct from those in QENP.
This study used a noninvasive sampling technique which still provides adequate initial insights into Salmonella ecology in birds. However capturing and obtaining samples from known species would have provided information on risk factors associated with Salmonella shedding in different guilds and age groups. Also, sampling was limited to October–March when migrants occur at these sites. Sampling throughout the year would have provided information on Salmonella occurrence during the non-migratory season. In addition, we used PFGE and MLVA to establish genotypic relationships; however, phylogenetic analysis of single-nucleotide polymorphisms based on whole genome sequence data would have provided better resolution of strain variations.
Future collaborations with ornithologists to capture birds would be invaluable to obtain health and biological data for monitoring zoonotic pathogens. Monitoring Salmonella in lake water and soils on roosting grounds would be invaluable in understanding Salmonella ecology. Furthermore, investigating Salmonella strains in humans residing in fishing and urban communities would provide insights into transmission processes at these human–wildlife interfaces. Satellite tracking and microbiological assessments would help in understanding the spatial extent of pathogen spread by birds.
Conclusions
Salmonellosis continues to exert a huge burden on public health worldwide (Majowicz et al. 2010) and epidemics with mortalities have occurred in birds (Velarde et al. 2012; Giovannini et al. 2013). We have shown that Salmonella is shed by birds in both conserved and human-impacted areas, more strains exist in the urban environment than in birds at all sites, some strains are adapted to birds, and that several strains in birds are indistinguishable from strains in the environment. Our results indicate that there is risk of different strains disseminating between the urban environment and birds across sites. Furthermore, Salmonella could reemerge particularly in birds at LVBA because diverse species of resident and migratory birds congregate at high densities and the spatial connection with Kampala city exposes birds to human source pathogens. While the significance of Salmonella shedding on bird health is unknown, it may affect population vitality during stressful periods or when coinfections occur (Klaassen et al. 2012). It is also possible that birds could spread strains acquired from impacted environments to humans, other species and areas and this calls for additional studies.
References
Afema, J. A., Byarugaba D.K., ShaH, D. H., Atukwase, E., Nambi, M. and Sischo, W. M. (2016) Potential sources and transmission of Salmonella and Antimicrobial resistance in Kampala, Uganda. PLoS One. 11(3):e0152130 (DOI: 10.1371/journal.pone.0152130).
Altizer,S., Bartel, R. and Han,B.A. (2011) Animal migration and infectious disease risk. Science 331, 296-302.
Arnold,M.E., Carrique-mas,J.J., McLaren,I. and Davies,R.H. (2011) A comparison of pooled and individual bird sampling for detection of Salmonella in commercial egg laying flocks. Preventive Veterinary Medicine. 99, 176-184.
Arnold,M.E., Mueller-doblies,D., Carrique-mas, J.J. and Davies,R.H. (2009) The estimation of pooled-sample sensitivity for detection of Salmonella in turkey flocks. J. Appl. Microbiol. 107, 936-943.
Bauer,A.W., Kirby,W.M., Sherris,J.C. and Turck,M. (1966) Antibiotic susceptibility testing by a standardized single disk method. A. J. Clin. Path. 45, 493-496.
Benskin,C.M., Wilson,K., Jones,K. and Hartley,I.R. (2009) Bacterial pathogens in wild birds: a review of the frequency and effects of infection. Biological Reviews 84, 349-373.
Berge,A.C.B., Dueger,E.L. and Sischo,W.M. (2006) Original article: Comparison of Salmonella enterica serovar distribution and antibiotic resistance patterns in wastewater at municipal water treatment plants in two California cities. J. Appl. Microbiol. 101, 1309-1316.
Byaruhanga, A. and Nalwanga, D. (2006) Ten years of continuous waterbird monitoring at Lutembe Bay, Lake Victoria, Uganda. In Waterbirds around the world. ed. Boere,G.C., Galbraith C.A. and Stroud D.A. pp. 457-458. Edinburgh: The Stationery Office.
CDC (2013) National Antimicrobial Resistance Monitoring System for Enteric Bacteria (NARMS): Human Isolates Final Report, 2011. Atlanta, GA: U.S. Department of Health and Human Services, CDC, 2013.
Daszak, P., Cunningham, A.A. and Hyatt, A.D. (2000) Emerging infectious diseases of wildlife–threats to biodiversity and human health. Science 287, 443-449.
Francisco AP, Vaz C (2012) PHYLOViZ: phylogenetic inference and data visualization for sequence based typing methods. BMC Bioinformatics 13, 1.
Giovannini,S., Pewsner,M., Hussy,D., Hachler,H., Ryser Degiorgis,M.P., von,H.J. and Origgi,F.C. (2013) Epidemic of salmonellosis in passerine birds in Switzerland with spillover to domestic cats. Vet. Pathol. 50, 597-606.
Hamer,S.A., Lehrer,E. and Magle,S.B. (2012) Wild birds as sentinels for multiple zoonotic pathogens along an urban to rural gradient in greater Chicago, Illinois. Zoonoses Public Health 59, 355-364.
Hernandez,J., Bonnedahl,J., Waldenstrom,J., Palmgren,H. and Olsen,B. (2003) Salmonella in birds migrating through Sweden. Emerg. Infect. Dis. 9, 753-755.
Hughes, L.A., Shopland, S., Wigley, P., Bradon, H., Leatherbarrow, A.H., Williams, N.J., Bennett, M., de, P.E., Lawson, B., Cunningham, A.A. and Chantrey, J. (2008) Characterisation of Salmonella enterica serotype Typhimurium isolates from wild birds in northern England from 2005–2006. BMC Veterinary Research 4:1.
Hughes,L.A., Wigley,P., Bennett,M., Chantrey,J. and Williams,N. (2010) Multi-locus sequence typing of Salmonella enterica serovar Typhimurium isolates from wild birds in northern England suggests host-adapted strain. Lett. Appl. Microbiol. 51, 477-479.
Kansiime,F. and van Bruggen,J.J. (2001) Distribution and retention of faecal coliforms in the Nakivubo wetland in Kampala, Uganda. Water Sci. Technol. 44, 199-206.
Kitadai,N., Obi,T., Takase,K., Ninomiya,N. and Murase,T. (2010) Salmonella isolated from the feces of migrating cranes at the Izumi Plain (2002-2008): Serotype, antibiotic sensitivity and PFGE type. J. Vet. Med. Sci. 72, 939-942.
Klaassen,M., Hoye,B.J., Nolet,B.A. and Buttemer,W.A. (2012) Ecophysiology of avian migration in the face of current global hazards. Philos. Trans. R. Soc. Lond. B Biol. Sci. 367, 1719-1732.
Leader,B.T., Frye,J.G., Hu,J., Fedorka-Cray,P.J. and Boyle,D.S. (2009) High-throughput molecular determination of Salmonella enterica serovars by use of multiplex PCR and capillary electrophoresis analysis. J. Clin. Microbiol. 47, 1290-1299.
Lindstedt,B.A., Heir,E., Gjernes,E. and Kapperud,G. (2003) DNA fingerprinting of Salmonella enterica subsp. enterica serovar Typhimurium with emphasis on phage type DT104 based on variable number of tandem repeat loci. J. Clin. Microbiol. 41, 1469-1479.
Lindstedt,B.A., Vardund,T., Aas,L. and Kapperud,G. (2004) Multiple-locus variable-number tandem-repeats analysis of Salmonella enterica subsp. enterica serovar Typhimurium using PCR multiplexing and multicolor capillary electrophoresis. J. Microbiol. Methods. 59, 163-172.
Madden,R.H., Murray,K.A. and Gilmour,A. (2007) Carriage of four bacterial pathogens by beef cattle in Northern Ireland at time of slaughter. Lett. Appl. Microbiol. 44, 115-119.
Majowicz,S.E., Fazil,A., Musto,J., Kirk,M., Scallan,E., Angulo,F.J., Hoekstra,R.M., Jones,T.F. and O’Brien,S.J. (2010) The global burden of nontyphoidal Salmonella gastroenteritis. Clin. Infect. Dis. 50, 882-889.
NatureUganda (2010) Important Bird Areas in Uganda, Status and Trends Report 2009. Kampala: NatureUganda.
Nunn,C.L., Thrall,P.H., Leendertz,F.H. and Boesch,C. (2011) The spread of fecally transmitted parasites in socially-structured populations. PLoS ONE 6, e21677.
Nunn,C.L., Thrall,P.H. and Kappeler,P.M. (2014) Shared resources and disease dynamics in spatially structured populations. Ecol. Model. 272, 198-207.
28. Pennycott,T.W., Park,A. and Mather,H.A. (2006) Isolation of different serovars of Salmonella enterica from wild birds in Great Britain between 1995 and 2003. Vet. Rec. 158, (24) 817-820.
29. Pennycott,T.W., Ross,H.M., McLaren,I.M. and Park,A. (1998) Causes of death of wild birds of the family Fringillidae in Britain. Vet. Rec. 143, 151-158.
Pomeroy D, Asasira J (2011) Monitoring bird populations in Uganda—the first 25 years. In: Proceedings of the 12th Pan African Ornithological Congress, 2008, Harebottle DM, Craig AJFK, Anderson MD, Rakotomanana H, Muchai M (editors), Cape Town: Animal Demography Unit, pp 113–119.
Popoff,M.Y., Bockemuhl,J. and Gheesling,L.L. (2004) Supplement 2002 (no. 46) to the Kauffmann-White scheme. Res Microbiol. 155, 568-570.
Reed,K.D., Meece,J.K., Henkel,J.S. and Shukla,S.K. (2003) Birds, migration and emerging zoonoses: west nile virus, lyme disease, influenza A and enteropathogens. Clin. Med. Res. 1, 5-12.
Ribot,E.M., Fair,M.A., Gautom,R., Cameron,D.N., Hunter,S.B., Swaminathan,B. and Barrett,T.J. (2006) Standardization of pulsed-field gel electrophoresis protocols for the subtyping of Escherichia coli O157:H7, Salmonella, and Shigella for PulseNet. Foodborne Pathog. Dis. 3, 59-67.
Sanderson,F.J., Donald,P.F., Pain,D.J., Burfield,I.J. and van Bommel,F.P.J. (2006) Long-term population declines in Afro-Palearctic migrant birds. Biol. Con. 131, 93-105.
Sanderson,M.W., Gay,J.M., Hancock,D.D., Gay,C.C., Fox,L.K. and Besser,T.E. (1995) Sensitivity of bacteriologic culture for detection of Escherichia coli O157:H7 in bovine feces. J. Clin. Microbiol. 33, 2616-2619.
Stevenson, T. and Fanshawe J. (2006) The Birds of East Africa: Kenya, Tanzania, Uganda, Rwanda, Burundi. Princeton University Press.
Velarde,R., Porrero,M.C., Serrano,E., Marco,I., Garcia,M., Tellez,S., Dominguez,L., Aymi,R. and Lavin,S. (2012) Septicemic salmonellosis caused by Salmonella Hessarek in wintering and migrating Song Thrushes (Turdus philomelos) in Spain. J. Wildl. Dis. 48, 113-121.
Vickery,J.A., Ewing,S.R., Smith,K.W., Pain,D.J., Bairlein,F., Skorpilova,J. and Gregory,R.D. (2014) The decline of Afro-Palaearctic migrants and an assessment of potential causes. Ibis 156, 1-22.
Waldenstrom,J., Broman,T., Carlsson,I., Hasselquist,D., Achterberg,R.P., Wagenaar,J.A. and Olsen,B. (2002) Prevalence of Campylobacter jejuni, Campylobacter lari, and Campylobacter coli in different ecological guilds and taxa of migrating birds. Appl. Environ. Microbiol. 68, 5911-5917.
Watts, J. L. (2008) Performance standards for antimicrobial disk and dilution susceptibility tests for bacteria isolated from animals: approved standard. Wayne, PA: CLSI.
Yoshida,C., Franklin,K., Konczy,P., McQuiston,J.R., Fields,P.I., Nash,J.H., Taboada,E.N. and Rahn,K. (2007) Methodologies towards the development of an oligonucleotide microarray for determination of Salmonella serotypes. J. Microbiol. Methods. 70, 261-271.
Acknowledgments
We thank Lindsay Parrish, Stephanie Wright, Christy Crudo, Emily Hudson, and Russ McClanaghan of Sischo lab, WSU for laboratory assistance. Thanks to Denis Byarugaba, Esther Atukwase, Maria Nambi, and the staff of Microbiology Laboratory, College of Veterinary Medicine, Makerere University. We are grateful to Jeradi Kyeyune and John Mulondo for providing boat rides during fieldwork. Chief Warden QENP Nelson Guma provided a boat and rangers during fieldwork. Thanks also to Ludwig Siefert and James Kalyewa for field assistance at QENP. Research permits were granted by Uganda National Council for Science and Technology and Uganda Wildlife Authority.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
About this article
Cite this article
Afema, J.A., Sischo, W.M. Salmonella in Wild Birds Utilizing Protected and Human Impacted Habitats, Uganda. EcoHealth 13, 558–569 (2016). https://doi.org/10.1007/s10393-016-1149-1
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10393-016-1149-1