Abstract
Recreational waters are primary attractions at many national and state parks where feral swine populations are established, and thus are possible hotspots for visitor exposure to feral swine contaminants. Microbial source tracking (MST) was used to determine spatial and temporal patterns of fecal contamination in Congaree National Park (CONG) in South Carolina, U.S.A., which has an established population of feral swine and is a popular destination for water-based recreation. Water samples were collected between December 2017 and June 2019 from 18 surface water sites distributed throughout CONG. Host specific MST markers included human (HF183), swine (Pig2Bac), ruminant (Rum2Bac), cow (CowM3), chicken (CL), and a marker for shiga toxin producing Escherichia coli (STEC; stx2). Water samples were also screened for culturable Escherichia coli (E. coli) as part of a citizen science program. Neither the cow nor chicken MST markers were detected during the study. The human marker was predominantly detected at boundary sites or could be attributed to upstream sources. However, several detections within CONG without concurrent detections at upstream external sites suggested occasional internal contamination from humans. The swine marker was the most frequently detected of all MST markers, and was present at sites located both internal and external to the Park. Swine MST marker concentrations ≥ 43 gene copies/mL were associated with culturable E. coli concentrations greater than the U.S. Environmental Protection Agency beach action value for recreational waters. None of the MST markers showed a strong association with detection of the pathogenic marker (stx2). Limited information about the health risk from exposure to fecal contamination from non-human sources hampers interpretation of the human health implications.
Similar content being viewed by others
Introduction
Feral swine fecal contamination in environmental waters and wastewater is recognized as a possible exposure route for disease transmission1 from zoonotic pathogens such as Hepatitis E virus, Cryptosporidium parvum, and Giardia2,3,4. While reports to date from the U.S. and other countries do not show patterns of significant disease outbreaks specifically linked to feral swine impacts on water quality, this may provide a false perception of risk. Most of the feral swine range expansion and population increases in the U.S. have largely occurred within the last 20 years5.
Feral swine have become abundant and widespread in the United States (U.S.) because of their ability to adapt to a wide range of habitats and their high reproductive potential. Commonly reported environmental impacts from feral swine include physical habitat alterations resulting from rooting behavior (i.e., disturbing surfaces with their snouts to move objects around), predation of native species, and resource competition with native species5,6,7. Despite feral swine often inhabiting and defecating in areas close to water5,8,9, the impacts of swine fecal contamination in recreational waters and potential human health risks are still poorly understood9,10,11,12.
To prevent illnesses from exposure to pathogens from fecal contamination in waters designated for drinking and recreation, state governments set regulatory limits for fecal indicator bacteria (FIB) based on U.S. Environmental Protection Agency (EPA) recommendations. When regulatory limits are exceeded, the water body is listed as impaired and authorities from the associated state environmental agency must develop a plan (Total Maximum Daily Load, TMDL) to identify the sources of contamination and the practices that will be implemented to mitigate contamination. In 2016, over 43,000 waters across the U.S. were included on the EPA’s list of impaired waters13. Nearly 4,000 water impairments were due to Escherichia coli, an EPA-recommended regulatory FIB for recreational freshwaters14. Feral swine are known to harbor zoonotic diseases that can be transferred to humans15,16. Much of the existing research on pathways of transmission of zoonotic diseases in feral swine is related to transmission to livestock, direct contact with infected animals, consumption of infected farmed or feral swine pork, and contamination of crops6,16,17,18,19. In contrast, studies of risk from recreational contact with (e.g., swimming, boating, angling) and accidental ingestion of water contaminated by feral swine are largely absent from the literature. Recreational waters are common attractions at many national and state parks, which are also habitat for feral swine. Therefore, potential exposures to feral swine-contaminated water are increasingly common park management concerns.
Congaree National Park (CONG) in South Carolina encompasses more than 26,000 acres (105.22 km) of forested floodplains along the Congaree and Wateree Rivers (https://www.nps.gov/cong). Feral swine populations have rapidly increased at CONG over the last several decades. Researchers have documented significant resource impacts20,21 and the park has pursued a comprehensive management strategy to support systematic control efforts22. Simultaneously, visitation has grown significantly, increasing over 52% from 95,619 in the year 2000 to 145,929 in 201823. Much of the visitor experience at CONG, including paddling, angling, and hiking, is focused on recreating in and around Cedar Creek. Park staff lack the capacity for systematic FIB monitoring but have recently developed plans that would outline a volunteer-led program24.
In 2016, two Cedar Creek surface-water monitoring sites within CONG boundaries were added to the South Carolina Department of Health and Environmental Control 303(d) list of impaired surface waters due to elevated E. coli concentrations (Fig. 1, sites 6 and 10)25. The capacity for park management to effectively address bacterial contamination, however, has been limited without more data regarding precisely which sources should be targeted for mitigation. While feral swine are one possible source of fecal contamination in CONG, other possible sources include upstream agricultural operations, upstream septic systems, municipal wastewater conveyances, as well as in-park septic systems and native wildlife. Furthermore, the hydrologic setting of CONG facilitates the broad-scale redistribution of water-borne contamination during flood conditions that inundate much of the property. Monitoring FIB alone24 can provide a wealth of data for informing specific advisories but cannot discriminate among sources.
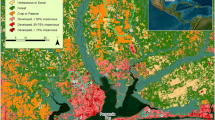
Locations sampled at Congaree National Park. Location names are listed in Table 1. Map created in ArcMap v.10.5.1 (ESRI, https://www.esri.com/).
Microbial source tracking (MST) is a potential approach for assessing the presence and relative magnitude of enteric bacteria in the environment while identifying the probable animal sources26,27,28,29,30,31,32,33. The intestinal systems of different animals such as humans, cattle, deer, or swine are microbiomes with different physiological and dietary signatures and, consequently, distinct enteric bacterial populations. Quantitative polymerase chain reaction (qPCR) is a sensitive molecular technique for detecting target DNA sequences. Bacteria from the order Bacteroidales are useful fecal-contaminant indicator microorganisms34 because they: (1) are restricted to warm-blooded animals; (2) constitute a relatively significant portion of the bacteria present in mammalian feces; (3) can be highly specific to particular species or taxonomic groups; and (4) are strict anaerobes. In warm waters their DNA may degrade beyond detection within 1 to 3 days35,36, which makes them indicative of recent contamination. Taken together, MST results can help identify mitigation targets to include in TMDLs.
In light of the challenges and opportunities at CONG, the objectives of this study were as follows: (1) measure the occurrence and concentrations of MST markers throughout CONG; (2) assess internal and external sources of fecal contamination; (3) assess the frequencies and distributions of detection of a pathogenic marker; and (4) investigate associations between MST marker concentrations and pathogenic marker presence.
Methods
Study site
Congaree National Park (CONG) in South Carolina is located in a broad floodplain between the Congaree and Wateree Rivers, and it includes over 11,000 acres (44.52 km2) of old-growth forest and over 15,000 acres (60.70 km2) of designated federal wilderness37. The Congaree River, which forms the southern park boundary (Fig. 1), drains approximately 8,290 mi2 (21,476 km2) and has varied land use throughout the watershed. Multiple major municipal wastewater systems in the Congaree River basin are within 25 miles upstream from the Park boundary. The state capitol, Columbia, has a population of over 800,000 people in the metropolitan area and is located approximately 20 mi (32.19 km) upstream of the Park. The Wateree River forms a short stretch of the park’s eastern boundary (Fig. 1) and drains over 5,600 mi2 (14,500 km2). Though not technically part of the park during normal in-bank streamflow conditions, the Congaree and Wateree Rivers (Fig. 1) are additional popular destinations for recreation37.
During normal flows on the Congaree and Wateree Rivers, the ecohydrology of the floodplain is dominated by several tributary systems including Cedar Creek, Tom’s Creek, and Dry Branch. Cedar Creek (Fig. 1) is a particularly popular destination for canoeing, kayaking, and fishing. The lower portion of Cedar Creek that flows through the park (between Wise Lake (Site 5) and the Congaree River, Fig. 1, Table 1) is the only waterbody in South Carolina with the designation of Outstanding Natural Resource Water38, a state-level designation authorized under the Clean Water Act to recognize water bodies with high ecological importance and exceptional water quality. Land use in the Cedar Creek watershed upstream of the Park is predominantly agricultural (including silviculture, poultry, horses, and farming), with some light industry and rural housing.
Sample collection and processing
One-liter water samples were collected and analyzed from 18 surface-water sites distributed throughout CONG (Fig. 1, Table 1). The sampling strategy was designed to determine if contamination was coming from inside or outside the Park by capturing primary inflows (Congaree River, Cedar Creek, Dry Branch, Myers Creek, Tom’s Creek, Wateree River) at respective upstream and downstream Park boundaries as well as to characterize a cross section of the principal hydrologic features (tributaries, wetland, oxbow lakes; Fig. 1) found within CONG. Eleven sample collection events were conducted between December 2017 and June 2019 to investigate temporal and spatial variability in sources of fecal contamination. Logistical constraints existed that included funding limitations, remote site access, and flooding thus resulting in different frequencies of sampling across study sites. The number of sites sampled during each collection event varied from 4 to 16. Five sites (Sites 3, 4, 6, 9, and 10; Fig. 1) were targeted for repeated sampling (sampled during each sample collection event as conditions allowed) to address temporal variability in sources; the remaining 13 sites were sampled intermittently (one to four times; Table 1) to inform spatial variability in sources.
Grab samples for molecular analysis were stored in coolers on ice until they were transferred to a refrigerator at the laboratory until processing, which occurred within 48 h of sample collection. Samples were filtered on a vacuum manifold through a 47-mm polycarbonate filter with 0.4-µm pore size (Millipore, Bedford, MA, USA). Samples were filtered in 100-mL aliquots or to the point of the filter clogging (minimum volume 20-mL39). Six filter replicates were collected per sample that included two filters that were enriched for E. coli with modified mTEC agar following the incubation protocol in EPA Method 160340 to enhance detection of shiga-toxin producing E. coli STEC41,42 and four unenriched filters. Two unenriched filters and one enriched filter were processed for molecular analysis. The remaining filter replicates were retained as backups for use if needed. All filter replicates were stored at − 80 °C until DNA extraction. Sterile de-ionized water (sterile DI; autoclaved 15 PSI, 121 °C for 15 min per 1-L of sterile DI43) controls were filtered in conjunction with each sample event to determine presence of potential contamination during sample collection and/or laboratory processing and were treated identically to filtered samples in downstream processing.
Citizen science Escherichia coli analysis
Beginning in October 2018, select duplicate water samples were collected for E. coli analysis at the CONG Old-Growth Bottomland Forest Research and Education Center as part of a Citizen Science methods development program by CONG personnel as described in McCarthy24. Samples were processed within 6 h of collection and kept on wet ice during the time between collection and processing. Each sample was screened at three dilutions to ensure accurate and quantifiable results. Samples were tested for E. coli concentrations and enumerated using the most probable number (MPN) method with IDEXX Colilert-18 (IDEXX Laboratories, Inc. Westbrook, ME) for enumerating E. coli. A laboratory negative control (sterile DI) was analyzed alongside samples for each sample event to test for contamination in reagents or contamination potentially introduced during sample processing. Resulting data were used in a post-hoc analysis to investigate relationships between E. coli MPN and MST marker concentrations.
DNA extraction and quantitative PCR
DNA was extracted using the Qiagen DNeasy PowerLyzer PowerSoil DNA Isolation Kit (Qiagen; Germantown, MD)44 following manufacturer’s instructions. We did not explicitly test the efficiency of the DNA extractions; however, this isolation kit has been shown to efficiently extract Bacteroides and E. coli DNA from stool samples45. Samples were tested for PCR inhibition following Bradshaw, et al.46 using the Sketa22 assay47. Samples were screened with probe-based quantitative PCR (qPCR) MST markers for humans (HF18348, Table 2), ruminants (Rum2Bac49, Table 2), chickens (CL50, Table 2), and swine (Pig2Bac49, Table 2). Samples that tested positive for ruminants were also screened for cows (CowM351, Table 2). PCR efficiency ranged from 82 to 102%39. Enriched samples were screened for stx252 (Table 2). Pig2Bac has been found to be highly sensitive, but the MST marker has been found to occur in dog feces, human feces, and septage32. To test for possible cross amplification of Pig2Bac with human contamination, we calculated Spearman’s ρ for the correlation between concentrations of HF183 and Pig2Bac. A significant correlation between Pig2Bac and HF183 could indicate that Pig2Bac concentrations included detection of human contamination.
DNA from two filter replicates per sample were screened for each marker with two qPCR replicates for a total of four qPCR replicates per sample per marker. Reactions were carried out in 96-well qPCR plates in 20-µL reaction volumes. Final concentrations of reagents in the assays were 1-µM forward and reverse primers, 80-nM 6-carboxy-fluorescein FAM- labeled TaqMan probe, 0.02-mg/mL BSA (Life Technologies), 1 × DNA TaqMan Fast Universal PCR Master Mix (Life Technologies), and 4-µL of template (i.e., genomic DNA, nanopore water as a no template control, or MST marker standard concentrations of 10 to 106 copies). Synthetic gene fragments with the marker insert sequences were used to create the standard curves for all MST markers and as a qPCR positive template control for stx2. Microbial source tracking markers with low specificity can reduce confidence that a detection is indicative of contamination from the targeted host species. To reduce the likelihood of spurious detection of an MST marker from a non-target host species, samples were considered absent for a marker when amplification occurred in zero or one of the four qPCR replicates. Samples absent for an MST marker were assigned an estimated concentration of 0 gene copies. The stx2 assay was analyzed as presence/absence where amplification in one or both qPCR replicates was considered evidence of presence. With the exception of the standard curve qPCR replicates for MST markers and positive template control reactions for the stx2 qPCR assay, no procedural positive controls were included during sample processing. Therefore, the influence of variable enrichment efficiencies for E. coli or DNA extraction efficiencies among samples on MST marker concentration or stx2 presence/absence results cannot be ruled out. Across all MST markers, R2 values for standard curves ranged from 0.987 to 0.999, PCR amplification efficiencies ranged from 82 to 102% for all markers, and slopes ranged from -3.85 to -3.27. The limit of quantification (LOQ) for each MST marker was determined as the lowest standard curve copy number with a coefficient of variation (CV = 100*σ/µ; where σ is the standard deviation, and µ is the mean number of estimated gene copies) less than 35%. Back calculated gene copy estimates were based on the equation X0 = EAMP(b-CT) where X0 is the initial number of target copies in the qPCR; EAMP is the exponential amplification value, which is 1 + the amplification efficiency (e.g., if the amplification efficiency is 0.94, EAMP = 1.94); b is the y-intercept of the standard curve; and CT is the cycle number when the amplification curve crosses the threshold line that distinguishes fluorescence intensity of a reaction from background levels.
Statistical analysis
A statistical technique similar to McKee, et al.33 was used to account for error in qPCR estimates below LOQ. If a qPCR estimated gene copy number was greater than 0 and below LOQ, a random number was selected from a normal distribution centered at the qPCR estimated copy number. Randomly selected negative values were set to 0 and randomly selected values above LOQ were set to the LOQ. To determine the standard distribution of the error estimates for each marker, a trendline was created with the standard deviations of the back-calculated gene copy number estimates for the standard curve gene copy numbers at and below LOQ (Table A.1). An additional data point at the origin (0, 0) served as a value for the no template controls. The equation for that trendline was used to calculate the standard deviation for each randomly selected number, which was a function of the qPCR estimated gene copy number. Randomly selected numbers were treated the same as qPCR measured concentrations above the LOQ for statistical analysis. All analyses were conducted in JMP v 14.2.0 (SAS Institute Inc.). Recursive partitioning was conducted to determine the MST marker and concentration that best predicted stx2 detection and E. coli density splits. Recursive partitioning is a nonparametric data-mining analysis that splits the response variable into two categories based on an explanatory variable cutting value. The cutting value is determined with a partition algorithm that finds the explanatory variable split that maximizes the difference in the response frequencies between the two nodes of the split. Spearman’s ρ were used to assess correlative relationships between MST marker concentrations and E. coli MPN. To account for multiple comparisons of the same data, we used p < 0.01 to indicate statistical significance for all Spearman’s rank correlations.
Results
HF183 and Pig2Bac were detected at 14 sites, Rum2Bac at 2 sites, and stx2 at 7 sites. Pig2Bac was detected in 37 samples (46%), which was more frequent than HF183 (21 samples, 26%), Rum2Bac (4 samples, 5%), and stx2 (15 samples, 19%). Of the 324 qPCR replicates per MST marker, many qPCR replicate concentrations were less than LOQ and therefore estimated as described above (HF183: 74 qPCR replicates from 24 samples; Pig2Bac: 67 qPCR replicates from 25 samples; Rum2Bac: 16 qPCR replicates from four samples; Table A.2). Spearman’s rank correlation between HF183 and Pig2Bac was not significant (ρ = 0.445, p = 0.026, Table 3) and HF183 was detected in only 15 of 37 samples that were positive for Pig2Bac, suggesting limited influence of human contamination on Pig2Bac concentrations.
Spatial and temporal variability in sources throughout the Park and at Cedar Creek
HF183 and Pig2Bac were detected broadly across the study sites (Figs. 2, 3 and 4), with detections at sites inside, outside, and on the boundary of CONG (Figs. 3, 4). The highest concentrations of HF183 detected were on the Congaree River boundary waters (Site 1, Congaree R @ gage: 26 copies/mL; and Site 15, Congaree R @ SC601: 12 copies/mL, Fig. 2, 3). HF183 was the only marker detected at all four sites along Cedar Creek, with a maximum detected concentration of 6 copies/mL detected at both Cedar Cr @ gage and Cedar Cr abv Congaree R (Sites 6 and 12, respectively; Figs. 2, 3). The highest concentrations of Pig2Bac detected across all sites were from samples collected from two of the three study sites on the 2016 South Carolina 303(d) list: Site 6, Cedar Cr @ gage 398 copies/mL; and Site 10, Cedar Cr. @ Kingsnake 127 copies/mL (Figs. 2, 4). Rum2Bac was detected at two sites, one immediately upstream of the Park boundary (Site 4, Myers Cr abv Cedar Cr) and one within the Park on Cedar Cr @ gage (Site 6, Fig. 5). The stx2 marker was detected at boundary water sites, as well as sites within and upstream of CONG (Fig. 6). Of the internal sites, stx2 was detected most frequently in samples from Cedar Cr @ Kingsnake (Site 10, five of 11 samples, Fig. 6).

source tracking marker concentrations (gene copies per mL) for humans (HF183), swine (Pig2Bac), ruminants (Rum2Bac), and qPCR replicate detections of Shiga toxin gene marker (stx2) in water samples per site throughout Congaree National Park. Dots represent individual samples. Dashed blue lines indicate the limit of quantification (LOQ). Box indicates first and third quartiles. The upper and lower whiskers extend to the largest and smallest values within 150% of the inter-quartile range (non-detections for HF183 and Pig2Bac not shown on log scale).
Quantitative PCR (qPCR) microbial
HF183 concentrations throughout and surrounding Congaree National Park for samples collected between December 18, 2017 and June 11, 2019. Maps created in ArcMap v.10.5.1 (ESRI, https://www.esri.com/).
Pig2Bac concentrations throughout and surrounding Congaree National Park for samples collected between December 18, 2017 and June 11, 2019. Maps created in ArcMap v.10.5.1 (ESRI, https://www.esri.com/).
Rum2Bac concentrations within and around Congaree National Park for samples collected between December 18, 2017 and June 11, 2019. Maps created in ArcMap v.10.5.1 (ESRI, https://www.esri.com/).
stx2 marker detections within and surrounding Congaree National Park for samples collected between December 18, 2017 and June 11, 2019. Maps created in ArcMap v.10.5.1 (ESRI, https://www.esri.com/).
Temporally, Pig2Bac was the most consistent marker to be detected, because it was present in one or more samples for 10 of 11 sample events (Fig. 4). HF183 was detected at one or more sites for 6 of 11 sample events and Rum2Bac was only detected in samples from the last two sample events (Figs. 3 and 5). The stx2 marker was detected in samples from at least one site for 8 of 11 sample events (Fig. 6).
MST marker predictions of shiga-toxin producing E. coli presence and E. coli MPN
None of the three MST markers were strong predictors of stx2 detection. Recursive partitioning indicated that the best predictor of stx2 detection across all samples was Pig2Bac concentrations ≥ 43 copies/mL (R2 = 0.135). The Pig2Bac split correctly identified 5 of 15 samples with stx2 detections. Samples with Pig2Bac concentrations ≥ 43 copies/mL were associated with an average 51% greater probability of stx2 detection compared to only a 14% probability of stx2 detection in samples with Pig2Bac concentrations < 43 copies/mL. All samples with stx2 detections that were positive for HF183 were also positive for Pig2Bac, whereas seven samples with stx2 detections were positive for Pig2Bac but not HF183. Three of the four samples with stx2 detections that were not positive for either HF183 or Pig2Bac were from Myers Creek. stx2 was only detected in two samples for which Rum2Bac was also detected.
Posthoc analysis with culturable E. coli data from citizen science collected samples (Table A.2) also indicated that Pig2Bac concentrations ≥ 43 copies/mL were the best recursive partitioning split for E. coli MPN (R2 = 0.331, root mean square error = 137.6, AICc = 324.3). When Pig2Bac concentrations were ≥ 43 copies/mL, E. coli concentrations had a mean of 357 MPN (± 195 standard deviation). In contrast, mean E. coli concentration for samples with Pig2Bac concentrations < 43 copies/mL was 115 MPN (± 130), indicating an average increase in E. coli MPN of 242 in samples containing ≥ 43 copies/mL of Pig2Bac. No significant correlations were detected between MST markers and E. coli MPN (Table 3).
Discussion
Results of this study suggest that humans are an intermittent source of bacterial contamination and swine are primary contributors of fecal contamination in CONG. Infrequent detection of the ruminant marker and no detections of the cow or poultry marker suggest that wildlife, cattle, and poultry operations were not major contributors of contamination in the Park at the time of sample collection.
Internal versus external sources of contamination
Human MST marker data are consistent with upstream inputs at park boundaries as well as some backcountry locations, but show no evidence that the park’s primary visitor center septic system is a major source of fecal contamination in recreational waters within CONG. Feral swine markers, by contrast, are consistent with inputs from feral swine internal to CONG. Possible sources of contamination from outside the Park include failing septic systems and leaking wastewater conveyances. Sources of human contamination from within the park could include on-site septic systems as well as local backcountry contributions from hikers and campers. HF183 was detected at all four Cedar Creek sites, however, concentrations were never at levels above 10 copies/mL and all were below the LOQ. The highest concentrations of the human marker were observed from Congaree River samples, consistent with external municipal or residential wastewater conveyances as a potential source. Human contamination was not detected at the Muck Swamp site, located adjacent to the Park’s Visitor Center septic system, suggesting that the on-site septic system was likely not a major net source of bacterial contamination during this study. It is possible that lack of septic contamination here could be related to its filtering through organic rich soils in Muck Swamp, as these types of tannin-rich settings have been noted to have natural anti-bacterial properties54. Another previously suggested possible source of human waste contamination within the Park is direct local inputs of contamination associated with visitation in the backcountry. Contaminants indicative of anthropogenic contamination have been found in water samples from non-stream sites at CONG, including samples from Muck Swamp, Wise Lake, and Weston Lake, under non-flood conditions (i.e., in the absence of a surface-water connection to extra-Park contaminant sources on the Congaree and Wateree Rivers or tributaries)55. In contrast, under flood conditions, interior Park locations can be hydrologically connected to the Congaree and Wateree Rivers and the associated external human wastewater sources. Two U.S. Geological Survey real-time gaging stations on the Congaree River (Congaree R @ Gage, site 1; Fig. 1) and Cedar Creek (Cedar Cr @ Gage, site 6; Fig. 1) are used to inform the public when the Park is under flood conditions based on gage height (15 feet or greater at Congaree R @ Gage, 8 feet or greater for Cedar Cr @ Gage; www.nps.gov/cong/planyourvisit/conditions.htm, accessed Feb. 28, 2020). Data from April 1, 2019 (waterdata.usgs.gov/, accessed Feb. 28, 2020) indicate CONG was not under flood conditions when the human MST marker was detected at Stump Gut, Horseshoe Lake, and Cedar Creek @ Gage suggesting contamination may have come from internal inputs. Other detections of human contamination at the Cedar Cr. @ Gage site were concurrent with upstream detections at the Cedar Creek upstream Park boundary sites at Cedar Cr. @ Myers and Myers Cr., consistent with downstream transport from external sources during non-flood conditions. All other detections of human contamination were at boundary water sites, or sites proximal to the boundary and downstream from potential external sources.
Feral swine are well documented throughout the southeastern U.S. and particularly in the study region of the Coastal Plain in South Carolina5. The lack of hog farms within the Congaree River watershed indicates feral swine as the main source of the swine MST marker in CONG. Further, while a comparison of swine marker loads (concentration per unit time) would be necessary for verification, higher swine MST marker concentrations at Cedar Creek sites within the Park compared to sites at the upstream boundary of the Park suggests swine contamination is predominantly, but not exclusively, occurring within CONG.
The home ranges of feral swine follow similar patterns to other mammals and tend to be larger for males than females. Feral swine home ranges can vary annually and seasonally and are generally smaller when food sources are more abundant56,57. Previous studies on feral swine home ranges in South Carolina have indicated that in coastal marshes, the average home range sizes were 2.26 km2 for males and 1.81 km2 for females58 and 3.89 km2 on average for males and females in the Piedmont59. In-house data from Park files and partner research suggest that local swine can move up to four miles per day and swim the Congaree River multiple times per day, indicating that detections of the swine MST marker external to CONG could be from feral swine the home ranges of which include the Park.
There are a couple limitations to the results from this study. One limitation is that the methods did not test for markers from the American alligator (Alligator mississipianensis). Research suggests that crocodilian feces are exceptionally rich in FIB60, although limited research has been conducted on pathogenic E. coli in reptiles61. Research at the Park suggests that the local alligator populations are low density and transient, but they could be disproportionately represented in some samples. Most alligators, however, are concentrated along the Congaree and Wateree Rivers at the southeast end of the park and well away from the Cedar Creek locations central to this analysis. Additionally, analysis for specific markers for turkey and other avian sources were not conducted, and these animals can represent a significant portion of the non-swine and non-deer megafauna in the Park. A second limitation to the study is that we cannot rule out non-specific amplification of the swine marker as a factor in the frequency of detection and estimated concentrations of the swine marker over the course of the study. We did not detect a correlation between the human and swine marker concentrations, which we interpreted as evidence that non-specific amplification from the swine marker was not occurring when human fecal contamination was present. However, the lack of correlation may have been caused by differences in stability of the signal between the human and swine markers due to factors such as differences in decay rates among markers from different sources62. Differences in environmental persistence among MST markers and markers from different sources may also explain why we did not detect a significant association between any of the MST markers and the pathogenic E. coli marker.
Human health risk from exposure to water with fecal contamination
We did not find evidence of a strong association between sources of fecal contamination and presence of STEC. Possible methodological explanations for the lack of a strong association between stx2 presence and any of the MST markers include variable efficiencies in bacterial recovery from environmental samples, different DNA extraction efficiencies among samples, and suboptimal incubation conditions for STEC. Unlike most other fecal coliforms, which are characterized by their ability to grow at 44.5–45.5 °C, studies have suggested that E. coli O157:H7, the most common serotype of STEC to cause infection63,64, does not grow well at temperatures above 41 °C64,65. However, Duris, et al.42 detected STEC in more than 50% of river water samples from Michigan and Indiana using a similar incubation protocol as used in the study herein. Results from our study and Duris, et al.42 indicate that incubating enriched samples at 44.5 °C does not inherently prevent detection of STEC, although presence may be underestimated.
Recursive partitioning of E. coli indicated that swine MST marker levels above 43 copies/mL were associated with E. coli levels above the EPA beach action value (BAV) for recreational freshwaters of 235 colony forming units/100 mL (CFU/100 mL; conversion between CFU to MPN is 1:1). This suggests that when Pig2Bac levels are above a certain threshold, FIB levels are likely to exceed BAV levels. The E. coli BAV for recreational freshwaters is based on investigations of the frequency of gastro-intestinal (GI) illness in swimmers exposed to water downstream of a sewage treatment facility and thus primarily exposed to human contamination14. Research suggests that, in general, exposure to human fecal contamination may be a greater human-health risk than exposure to fecal contamination from non-human sources66,67,68. This is likely due in part to the host specificity of many viruses69 that are the etiologic agents most frequently responsible for human illness associated with exposure to recreational waters70. Therefore, the health risk from exposure to recreational freshwater contaminated by non-human sources may not be accurately represented by the BAV. Additionally, the BAV was developed to indicate the likelihood of illness from recreational exposure to contaminated waters. Feral swine zoonoses caused by exposure to contaminated freshwater may present increased variability in the severity of illness that is not represented by the BAV.
Studies suggest that exposure to fecal contamination from domestic pigs presents a lower health risk than exposure to human fecal contamination66,71. Health risks from exposure to recreational waters with FIB densities of 35 CFU/100 mL enterococci and 126 CFU/100 mL E. coli from domestic pig fecal contamination were estimated to be substantially lower than the risk from exposure to human sources, with the median risk from domestic pigs two orders of magnitude lower than the human-based benchmarks66. Based on these findings, a new water-quality benchmark for domestic pigs should not be ruled out66.
If health risks from exposure to domestic pig and feral swine fecal contamination are similar, the frequency of detection of the feral swine MST marker in CONG relative to the human MST marker may indicate the health risk is lower than would be predicted if humans were the predominant source of contamination. However, illness rates from exposure to feral swine fecal contamination may differ from illness rates due to exposure to domestic pig contamination. Feral swine and domestic pigs have been found to serve as reservoirs to different enteropathogenic bacterial strains72, suggesting that health risks from exposure to domestic pig fecal contamination may not accurately represent the risk from exposure to feral swine. Feral swine are known to be hosts for numerous zoonotic pathogens, including Mycobacterium avium complex73, which can cause respiratory illness and be transmitted through recreational waters74. The application of quantitative microbial risk assessments75 may be useful for predicting the likelihood of illness from zoonotic pathogens in recreational waters with feral swine fecal contamination. Further research is needed to determine the human health risks from exposure to feral swine contamination and to identify other locations where human exposure to feral swine fecal contamination is likely to occur from contact with contaminated recreational waters.
Regulatory implications of wildlife contamination in impaired recreational waters
Study data suggesting significant wildlife inputs (relative to humans) have potential regulatory implications. Current (2020) EPA Recreational Water Quality Criteria (RWQC)14 acknowledge the potential for differences in human health risks from exposure to recreational waters contaminated by human versus non-human sources but do not provide water-quality standard recommendations that take sources of fecal contamination into account. Instead, the 2012 RWQC describes methods that can be used to develop site-specific standards for states that desire to address the variability in human health risks associated with different sources of fecal contamination. In Florida, MST was used to assess sources of fecal contamination in a wildlife conservation-managed watershed that was included on the 2010 Florida Department of Environmental Protection 303(d) list of impaired waters for FIB76. Results from the MST analysis suggested that birds, not humans, were the main source of contamination76. A natural source exclusion status was obtained for the water body, which was subsequently removed from the Florida 303(d) list. At least 13 other states have natural resource—or “natural conditions”— exclusions in their administrative code, although the conditions for these exceptions and how they are applied can vary by state77. Intermittent detection of human contamination in South Carolina 303(d) listed impaired recreational waters in CONG may preclude these sites from a natural source exclusion. Further research would be necessary to determine if a natural resource exclusion would be appropriate or possible for the impaired streams in CONG.
Data availability
Quantitative PCR data for this study are available in the associated USGS data release39 in the USGS ScienceBase-Catalog, which is the authoritative repository of these data.
References
Eckert, K. D., Keiter, D. A. & Beasley, J. C. Animal visitation to wild pig (Sus scrofa) wallows and implications for disease transmission. J. Wildl. Dis. 55, 488–493. https://doi.org/10.7589/2018-05-143 (2019).
Fenaux, H. et al. Transmission of hepatitis E virus by water: an issue still pending in industrialized countries. Water Res. 151, 144–157 (2019).
Atwill, E. R. et al. Prevalence of and associated risk factors for shedding Cryptosporidium parvum oocysts and Giardia cysts within feral pig populations in California. Appl. Environ. Microbiol. 63, 3946–3949 (1997).
Hampton, J., Spencer, P. B. S., Elliot, A. D. & Thompson, R. C. A. Prevalence of zoonotic pathogens from feral pigs in major public drinking water catchments in Western Australia. EcoHealth 3, 103–108. https://doi.org/10.1007/s10393-006-0018-8 (2006).
McClure, M. L. et al. Modeling and mapping the probability of occurrence of invasive wild pigs across the contiguous United States. PLoS ONE 10, e0133771. https://doi.org/10.1371/journal.pone.0133771 (2015).
Mayer, J. J. & Beasley, J. C. Ecology and management of terrestrial vertebrate invasive species in the United States 221–250 (CRC Press, Boca Raton, 2017).
Boughton, E. H. & Boughton, R. K. Modification by an invasive ecosystem engineer shifts a wet prairie to a monotypic stand. Biol. Invas. 16, 2105–2114. https://doi.org/10.1007/s10530-014-0650-0 (2014).
Hone, J. Feral pig rooting in a mountain forest and woodland: distribution, abundance and relationships with environmental variables. Aust. J. Ecol. 13, 393–400 (1988).
Belden, R. C. & Pelton, M. R. Wallows of the European Wild Hog in the Mountains of East Tennessee. J. Tenn. Acad. Sci. 51, 91–93 (1976).
Kaller, M. & Kelso, W. in Proceedings of the Annual Conference of the southeastern Association of Fish and Wildlife Agencies. 291–298.
Kaller, M. D. & Kelso, W. E. Swine activity alters invertebrate and microbial communities in a coastal plain watershed. Am. Midl. Nat. 156, 163–178 (2006).
Kaller, M. D., Hudson, J. D. III., Achberger, E. C. & Kelso, W. E. Feral hog research in western Louisiana: expanding populations and unforeseen consequences. Hum. Wildl. Confl. 1, 168–177 (2007).
U.S. Environmental Protection Agency. National Summary of Impaired Waters and TMDL Information, accessed Nov. 20, 2020 at https://iaspub.epa.gov/waters10/attains_nation_cy.control?p_report_type=T (2020).
U.S. Environmental Protection Agency. Recreational Water Quality Criteria. (Health and Ecological Criteria Division, Office of Science and Technology, U.S. Environmental Protection Agency. EPA 820-F-12-058, 2012).
Hutton, T., DeLiberto, T. J., Owen, S. & Morrison, B. Disease risks associated with increasing feral swine numbers and distribution in the United States. Mich. Bovine Tubercul. Bibliogr. Database 59, 1–15 (2006).
Ruiz-Fons, F. A review of the current status of relevant zoonotic pathogens in wild swine (Sus scrofa) populations: changes modulating the risk of transmission to humans. Transb. Emerg. Dis. 64, 68–88 (2017).
Shwiff, S., Shwiff, S., Holderieath, J., Haden-Chomphosy, W. & Anderson, A. Ecology and management of terrestrial vertebrate invasive species in the United States 35–60 (CRC Press, Boca Raton, 2017).
Brown, V., Bowen, R. & Bosco-Lauth, A. Zoonotic pathogens from feral swine that pose a significant threat to public health. Transb. Emerg. Dis. 65, 649–659 (2018).
Jay, M. T. et al. Escherichia coli O157:H7 in feral swine near spinach fields and cattle, central California coast. Emerg. Infect. Dis. 13, 1908–1911. https://doi.org/10.3201/eid1312.070763 (2007).
Friebel, A. & Jodice, P. Home range and habitat use of feral hogs in Congaree National Park, South Carolina. Hum. Wildl. Confl. 3, 49–63 (2009).
Zengel, S. Wild pig habitat use, substrate disturbance, and understory vegetation at Congaree National Park Ph. D. thesis, Clemson University (2008).
U.S. National Park Service. Final management plan for non-native wild pigs within Congaree National Park (CONG), with Environmental Assessment. (2014).
U.S. National Park Service. National Park Service Visitor Use Statistics, 2020).
McCarthy, S. Bacterial Water Quality Monitoring as Citizen Science in Congaree National Park, South Carolina MSc thesis, University of South Carolina, (2020).
DHEC. The State of South Carolina's 2016 Integrated Report Part I: Listing of Impaired Waters, accessed Nov. 20, 2020 at https://scdhec.gov/sites/default/files/docs/HomeAndEnvironment/Docs/tmdl_16-303d.pdf (2016).
Meays, C., Broersma, K., Nordin, R. & Mazumder, A. Source tracking fecal bacteria in water: a critical review of current methods. J. Environ. Manag. 73, 71–79 (2004).
Bernhard, A. & Field, K. A PCR assay to discriminate human and ruminant feces on the basis of host differences in Bacteroides-Prevotella genes encoding 16S rRNA. Appl. Environ. Microbiol. 66, 4571–4574 (2000).
Beversdorf, L. J., Bornstein-Forst, S. M. & McLellan, S. L. The potential for beach sand to serve as a reservoir for Escherichia coli and the physical influences on cell die-off. J. Appl. Microbiol. 102, 1372–1381. https://doi.org/10.1111/j.1365-2672.2006.03177.x (2007).
Byappanahalli, M. N., Nevers, M. B., Korajkic, A., Staley, Z. R. & Harwood, V. J. Enterococci in the Environment. Microbiol. Mol. Biol. Rev. 76, 685–706. https://doi.org/10.1128/MMBR.00023-12 (2012).
Byappanahalli, M. N., Roll, B. M. & Fujioka, R. S. Evidence for occurrence, persistence, and growth potential of Escherichia coli and enterococci in Hawaii’s soil environments. Microbes Environ. 27, 164–170. https://doi.org/10.1264/jsme2.ME11305 (2012).
Bradshaw, J. K. et al. Characterizing relationships among fecal indicator bacteria, microbial source tracking markers, and associated waterborne pathogen occurrence in stream water and sediments in a mixed land use watershed. Water Res https://doi.org/10.1016/j.watres.2016.05.014 (2016).
Leray, M. et al. A new versatile primer set targeting a short fragment of the mitochondrial COI region for metabarcoding metazoan diversity: application for characterizing coral reef fish gut contents. Front. Zool. 10, 1 (2013).
McKee, B. A., Molina, M., Cyterski, M. & Couch, A. Microbial source tracking (MST) in chattahoochee river national recreation area: seasonal and precipitation trends in MST marker concentrations, and associations with E. coli levels, pathogenic marker presence, and land use. Water Res. 171, 115435. https://doi.org/10.1016/j.watres.2019.115435 (2020).
Allsop, K. & Stickler, J. An assessment fo Bacteroides fragilis group organisms as indicators of human faecal pollution. J. Appl. Bacteriol. 58, 95–99 (1985).
Kreader, C. A. Persistence of PCR-detectable Bacteroides distasonis from human feces in river water. Appl. Environ. Microbiol. 64, 4103–4105. https://doi.org/10.1128/aem.64.10.4103-4105.1998 (1998).
Ballesté, E. & Blanch, A. R. Persistence of Bacteroides Species populations in a river as measured by molecular and culture techniques. Appl. Environ. Microbiol. 76, 7608–7616. https://doi.org/10.1128/aem.00883-10 (2010).
USNPS. Foundation Document Congaree National Park South Carolina. 76 (U.S. National Park Servic (USNPS), 2014).
Congaree River Keeper. Lower Richland Sewer Project (2014).
McKee, A. M., Bradley, P. M. & Romanok, K. M. Microbial Source Tracking Marker Concentrations in Congaree National Park in 2017–2019, South Carolina, USA: U.S. Geological Survey data release. https://doi.org/10.5066/P9GFT8M7 (2020).
U.S. Environmental Protection Agency. Method 1603: Escherichia coli (E. coli) in Water by Membrane Filtration Using Modified membrane-Thermotolerant Escherichia coli Agar (Modified mTEC). 42 (Washington, D.C., 2009).
Bauer, L. & Alm, E. Escherichia coli toxin and attachment genes in sand at Great Lakes recreational beaches. J. Great Lakes Res. 38, 129–133. https://doi.org/10.1016/j.jglr.2011.10.004 (2012).
Duris, J. W., Haack, S. K. & Fogarty, L. R. Gene and antigen markers of shiga-toxin producing E. coli from Michigan and Indiana River water: occurrence and relation to recreational water quality criteria. J. Environ. Qual. 38, 1878–1886 (2009).
Myers, D. N., Stoeckel, D. M., Bushon, R. N., Francy, D. S. & Brady, A. M. G. in Book 9, chap. A7, section 7.1, May 2014 (2014).
Videnska, P. et al. Stool sampling and DNA isolation kits affect DNA quality and bacterial composition following 16S rRNA gene sequencing using MiSeq Illumina platform. Sci. Rep. 9, 1–14 (2019).
Claassen, S. et al. A comparison of the efficiency of five different commercial DNA extraction kits for extraction of DNA from faecal samples. J. Microbiol. Methods 94, 103–110. https://doi.org/10.1016/j.mimet.2013.05.008 (2013).
Bradshaw, J. K. et al. Characterizing relationships among fecal indicator bacteria, microbial source tracking markers, and associated waterborne pathogen occurrence in stream water and sediments in a mixed land use watershed. Water Res. 101, 498–509 (2016).
Haugland, R. A. et al. Evaluation of genetic markers from the 16S rRNA gene V2 region for use in quantitative detection of selected Bacteroidales species and human fecal waste by qPCR. Syst. Appl. Microbiol. 33, 348–357. https://doi.org/10.1016/j.syapm.2010.06.001 (2010).
Green, H. C. et al. Improved HF183 quantitative real-time PCR assay for characterization of human fecal pollution in ambient surface water samples. Appl. Environ. Microbiol. 80, 3086–3094 (2014).
Mieszkin, S., Yala, J. F., Joubrel, R. & Gourmelon, M. Phylogenetic analysis of Bacteroidales 16S rRNA gene sequences from human and animal effluents and assessment of ruminant faecal pollution by real-time PCR. J. Appl. Microbiol. 108, 974–984. https://doi.org/10.1111/j.1365-2672.2009.04499.x (2010).
Ryu, H. et al. Comparison of two poultry litter qPCR assays targeting the 16S rRNA gene of Brevibacterium sp. Water Res. 48, 613–621 (2014).
Shanks, O. C. et al. Quantitative PCR for detection and enumeration of genetic markers of bovine fecal pollution. Appl. Environ. Microbiol. 74, 745–752 (2008).
Imamovic, L., Serra-Moreno, R., Jofre, J. & Muniesa, M. Quantification of Shiga toxin 2-encoding bacteriophages, by real-time PCR and correlation with phage infectivity. J. Appl. Microbiol. 108, 1105–1114 (2010).
Mieszkin, S., Furet, J.-P., Corthier, G. & Gourmelon, M. Estimation of pig fecal contamination in a river catchment by real-time PCR using two pig-specific Bacteroidales 16S rRNA genetic markers. Appl. Environ. Microbiol. 75, 3045–3054 (2009).
Field, J. A. & Lettinga, G. in Plant Polyphenols. Basic Life Sciences, vol 59 (eds Hemingway R.W. & Laks P.E.) (Springer, 1992).
Bradley, P. M. et al. Widespread occurrence and potential for biodegradation of bioactive contaminants in Congaree National Park, USA. Environ. Toxicol. Chem. 36, 3045–3056 (2017).
Harestad, A. S. & Bunnel, F. Home range and body weight—a reevaluation. Ecology 60, 389–402 (1979).
Sanderson, G. C. The study of mammal movements: a review. J. Wildl. Manag. 30, 215–235 (1966).
Wood, G. W. & Brenneman, R. E. Feral hog movements and habitat use in coastal South Carolina. J. Wildl. Manag. 44, 420–427. https://doi.org/10.2307/3807973 (1980).
Kurz, J. C. & Marchinton, R. L. Radiotelemetry studies of feral hogs in South Carolina. J. Wildl. Manag. 36, 1240–1248 (1972).
Johnston, M. A., Porter, D. E., Scott, G. I., Rhodes, W. E. & Webster, L. F. Isolation of faecal coliform bacteria from the American alligator (Alligator mississippiensis). J. Appl. Microbiol. 108, 965–973. https://doi.org/10.1111/j.1365-2672.2009.04498.x (2010).
Ramos, C. P. et al. Identification and characterization of Escherichia coli, Salmonella spp., Clostridium perfringens, and C. difficile Isolates from reptiles in Brazil. BioMed Res. Int. 2019, 9530732. https://doi.org/10.1155/2019/9530732 (2019).
Ballesté, E., García-Aljaro, C. & Blanch, A. R. Assessment of the decay rates of microbial source tracking molecular markers and faecal indicator bacteria from different sources. J. Appl. Microbiol. 125, 1938–1949. https://doi.org/10.1111/jam.14058 (2018).
Hadler, J. L. et al. Ten-year trends and risk factors for non-O157 Shiga Toxin-producing Escherichia coli found through Shiga Toxin testing, Connecticut, 2000–2009. Clin. Infect. Dis. 53, 269–276. https://doi.org/10.1093/cid/cir377 (2011).
Doyle, M. P. & Schoeni, J. L. Survival and growth characteristics of Escherichia coli associated with hemorrhagic colitis. Appl. Environ. Microbiol. 48, 855–856 (1984).
Raghubeer, E. V. & Matches, J. R. Temperature range for growth of Escherichia coli serotype O157:H7 and selected coliforms in E. coli medium. J. Clin. Microbiol. 28, 803–805 (1990).
Soller, J. A., Schoen, M. E., Bartrand, T., Ravenscroft, J. E. & Ashbolt, N. J. Estimated human health risks from exposure to recreational waters impacted by human and non-human sources of faecal contamination. Water Res. 44, 4674–4691 (2010).
Schoen, M. E. & Ashbolt, N. J. Assessing pathogen risk to swimmers at non-sewage impacted recreational beaches. Environ. Sci. Technol. 44, 2286–2291 (2010).
Sedmak, G., Bina, D., MacDonald, J. & Couillard, L. Nine-year study of the occurrence of culturable viruses in source water for two drinking water treatment plants and the influent and effluent of a wastewater treatment plant in Milwaukee, Wisconsin (August 1994 through July 2003). Appl. Environ. Microbiol. 71, 1042–1050 (2005).
Medina, R. A. & García-Sastre, A. Influenza A viruses: new research developments. Nat. Rev. Microbiol. 9, 590–603. https://doi.org/10.1038/nrmicro2613 (2011).
Soller, J. A., Bartrand, T., Ashbolt, N. J., Ravenscroft, J. & Wade, T. J. Estimating the primary etiologic agents in recreational freshwaters impacted by human sources of faecal contamination. Water Res. 44, 4736–4747 (2010).
U.S. Environmental Protection Agency. in EPA 822-R-10-005. (US EPA Office of Water, Washington, DC, 2010).
Fredriksson-Ahomaa, M., Wacheck, S., Bonke, R. & Stephan, R. Different enteropathogenic Yersinia strains found in wild boars and domestic pigs. Foodborne Pathogens Dis. 8, 733–737 (2011).
Lara, G. H. B. et al. Occurrence of Mycobacterium spp. and other pathogens in lymph nodes of slaughtered swine and wild boars (Sus scrofa). Res. Vet. Sci. 90, 185–188. https://doi.org/10.1016/j.rvsc.2010.06.009 (2011).
Pond, K. Water recreation and disease, plausibility of associated infections: acute effects, sequelae, and mortality (World Health Organization, London, 2005).
Soller, J. et al. Estimated human health risks from recreational exposures to stormwater runoff containing animal faecal material. Environ. Model. Softw. 72, 21–32. https://doi.org/10.1016/j.envsoft.2015.05.018 (2015).
Nguyen, K. et al. Determination of wild animal sources of fecal indicator bacteria by microbial source tracking (MST) influences regulatory decisions. Water Res. 144, 424–434 (2018).
Balanson, S. Holding nature responsible: the natural conditions exception to water quality standards of the clean water act. Clev. St. L. Rev. 56, 1057 (2008).
Acknowledgements
This study was supported by the U.S. Geological Survey/National Park Service (USGS-NPS) Water Quality Partnership Program and the USGS Toxic Substances Hydrology Program. The authors thank Congaree National Park personnel and Marcella Cruz (USGS) who assisted with sample collection and transportation. Reviews by Eric Vallegas (EPA) and Marcella Cruz (USGS) greatly improved the manuscript. This document has been reviewed in accordance with U.S Environmental Protection Agency policy and approved for publication. Any use of trade, firm, or product names is for descriptive purposes only and does not imply endorsement by the U.S. Government.
Author information
Authors and Affiliations
Contributions
A.M.M., P.M.B., and D.S. contributed to the conception and design of the study. P.M.B. and D.S. acquired water samples. A.M.M. processed samples for molecular data. S.M. processed samples for E. coli data. A.M.M., P.M.B. and. M.M. analyzed and interpreted the data. All authors contributed to drafting the manuscript and gave approval for its publication.
Corresponding author
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
McKee, A.M., Bradley, P.M., Shelley, D. et al. Feral swine as sources of fecal contamination in recreational waters. Sci Rep 11, 4212 (2021). https://doi.org/10.1038/s41598-021-83798-6
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-021-83798-6
- Springer Nature Limited