Abstract
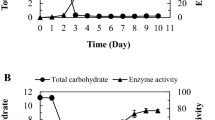
Dextranase can hydrolyze dextran to low-molecular-weight polysaccharides, which have important medical applications. In the study, dextranase-producing strains were screened from various soil sources. The strain H6 was identified as Talaromyces pinophilus by a standard ITS rDNA analysis. Crude dextranase was purified by ammonium sulfate fractionation and Sepharose 6B chromatography, which resulted in a 6.69-fold increase in the specific activity and an 11.27% recovery. The enzyme was 58 kDa, lower than most dextranase, with an optimum temperature of 45 °C and an optimum pH of 6.0, and identified as an endodextranase. It was steady over a pH range from 3.0 to 10.0 and had reasonable thermal stability. The dextranase activity was increased by urea, which enhanced its activity to 115.35% and was conducive to clinical dextran production. Therefore, T. pinophilus H6 dextranase could show its superiority in practical applications.
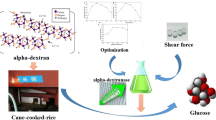
Graphical Abstract





Similar content being viewed by others
References
Abdel-Naby MA, Ismail AS, Abdel-Fattah AM, Abdel-Fattah AF (1999) Preparation and some properties of immobilized Penicillium funiculosum 258 dextranase. Process Biochem 34(4):391–398
Barret JF, Barret TA, Curtiss R (1987) Purification and partial characterization of the multicomponent dextranase complex of Streptococcus sobrinus and cloning of the dextranase gene. Infect Immun 55(3):792–802
Bradford MM (1976) A rapid and sensitive method for the quantitation of microgram quantities of protein utilizing the principle of protein-dye binding. Anal Biochem 72:248–254
Cai RH, Lu MS, Fang YW, Jiao YL, Zhu Q, Liu ZP, Wang SJ (2014) Ann Microbiol 64:147–155
Chen L, Zhou X, Fa W, Zhang Y (2008) Expression, purification and characterization of a recombinant Lipomyces starkey dextranase in Pichia pastoris. Protein Expres Purif 58(1):87–93
Das DK, Dutta SK (1996) Purification, biochemical characterization and mode of action of an extracellular endo-dextranase from the culture filtrate of Penicillium lilacinum. Int J Biochem Cell Biol 28:107–113
Finnegan PM, Brumbley SM, O’Shea MG, Nevalainen H, Bergquist PL (2004) Isolation and characterization of genes encoding thermoactive and thermostable dextranases from two thermo tolerant soil bacteria. Curr Microbiol 49(5):327–333
Gan WW, Zhang HB, Zhang YQ, Hu XQ (2014) Biosynthesis of oligodextrans with different Mw by synergistic catalysis of dextransucrase and dextranase. Carbohydrate Polym 112:387–395
Goulas AK, Fisher DA, Grimble GK, Grandison AS, Rastall RA (2004) Synthesis of isomaltooligosaccharides and oligodextrans by the combined use of dextransucrase and dextranase. Enzyme Microb Technol 35:327–338
Hayacibara MF, Koo F, Smith AMV, Kopec LK, Scott-Anne K, Cury JA, Bowen WH (2004) The influence of mutanase and dextranase on the production and structure of glucans synthesized by streptococcal glucosyltransferases. Carbohyd Res 339(12):2127–2137
Hussain I, Bhoyroo J, Butcher A, Koch TA, He A, Bregman DB (2013) Direct comparison of the safety and efficacy of ferric carboxymaltose versus iron dextran in patients with iron deficiency anemia. Anemia 169107:1–11
Jiao YL, Wang SJ, Lv MS, Jiao BH, Li WJ, Fang YW, Liu S (2014) Characterization of a marine-derived dextranase and its application to the prevention of dental caries. J Ind Microbiol Biotechnol 41:17–26
Khalikova E, Susi P, Korpela T (2005) Microbial dextran-hydrolyzing enzymes: fundamentals and applications. Microbiol Mol Biol R 69:306–325
Kim D, Day DF (1994) A new process for the production of clinical dextran by mixed-culture fermentation of Lipomyces starkeyi and Leuconostoc mesenteroides. Enzyme Microb Tech 16:844–848
Kim D, Robyt JF, Lee SY, Lee JH, Kim YM (2003) Dextran molecular size and degree of branching as a function of sucrose concentration, pH, and temperature of reaction of Leuconostoc mesenteroides B-512FMCM dextransucrase. Carbohyd Res 338:1183–1189
Kim YM, Kimura A, Kim D (2011) Novel quantitative method for the degree of branching in dextran. Food Sci Biotechnol 20(2):537–541
Kim YM, Kim D (2010) Characterization of novel thermostable dextranase from Thermotoga lettingae TMO. Appl Microbiol Biotechnol 85(3):581–587
Kulthida K, Pranee I, Emmanuelle M, Alain D (2012) Enzymatically degradable nanoparticles of dextran esters as potential drug delivery systems. Carbohyd Polym 88:875–881
Kang C, Yu XW, Xu Y (2014) Purification and characterization of a prolyl endopeptidase isolated from Aspergillus oryzae. J Ind Microbiol Biotechnol 41:49–55
Laemmli UK (1970) Cleavage of structural proteins during the assembly of the head of bacteriophage T4. Nature 227(5259):680–685
Larsson AM, Andersson R, Stahlberg J, Kenne L, Jones TA (2003) Dextranase from Penicillium minioluteum: reaction course, crystal structure, and product complex. Structure 11(9):1111–1121
Lineweaver H, Burk D (1934) The determination of enzyme dissociation constants. J Am Chem Soc 56:658–666
Mehvar R (2000) Dextrans for targeted and sustained delivery of therapeutic and imaging agents. J Control Release 69:1–25
Nelson N (1944) A photometric adaptation of the Somogyi method for the determination of glucose. J Biol Chem 153:375–380
Olcer Z, Tanriseven A (2010) Co-immobilization of dextransucrase and dextranase in alginate. Process Biochem 45:1645–1651
Purushe S, Prakash D, Nawani N, Dhakephalkar P, Kapadnis B (2012) Biocatalytic potential of an alkalophilic and thermophilic dextranase as a remedial measure for dextran removal during sugar manufacture. Bioresource Technol 115:2–7
Li RH, Zhang HB, Hu XQ, Gan WW, Li QP (2016) An efficiently sustainable dextran-based flocculant: synthesis, characterization and flocculation. Chemosphere 159:342–350
Singleton V, Horn J, Bucke C, Adlard M (2001) A new polarimetric method for the analysis of dextran and sucrose. Int Sugar J 103(1230):251–254
Sugiura M, Ito A, Ogiso T, Kato K, Asano H (1973) Purification of dextranase from Penicillium funiculosum and its enzymatic properties. B B A 309(2):357–362
Thitaram SN, Chung CH, Day DF, Hinton A, Bailey JS, Siragusa GR (2005) Isomaltooligosaccharide increases cecal Bifidobacterium population in young broiler chickens. Poultry Sci 84(7):998–1003
Virgen-Ortíza JJ, Ibarra-Junqueraa V, Escalante-Minakataa P, Ornelas-Pazb J de J,Osuna-Castroc JA, González-Potes A (2015) Kinetics and thermodynamic of the purified dextranase from Chaetomium erraticum. J Mol Catal B-Enzym 122:80–86
Walker GJ (1978) Dextrans. Biochem Carbohyd 11(16):75–126
Webb E, Spencer-Martins I (1983) Extracellular endodextranase from the yeast Lipomyces starkeyi. Can J Microbiol 29(9):1092–1095
Wu DT, Zhang HB, Huang LJ, Hu XQ (2011) Purification and characterization of extracellular dextranase from a novel producer, Hypocrea lixii F1002, and its use in oligodextran production. Process Biochem 46:1942–1950
Zdolsek HJ, Vecfors M, Lindahl TL, Tornquist T, Bortnik P, Hahn RG (2011) Hydroxyethyl starches and dextran during hip replacement surgery: effects on blood volume and coagulation. Acta Anaesth Scand 55(6):677–685
Zhang HB, Hu YJ, Zhu CB (2008) Cloning, sequencing and expression of a dextransucrase gene (dexYG) from Leuconostoc mesenteroides. Biotechnol Lett 30(8):1441–1446
Zhang HB, Wu DT, Huang LJ, Hu XQ, Wang X (2011) Purification, characterization of an extracellular dextranase from an isolated Penicillium sp. Acta Microbiol Sin 51(4):495–503
Zohra RR, Aman A, Ansari A, Haider MS, Qader SAU (2015) Purification, characterization and end product analysis of dextran degrading endodextranase from Bacillus licheniformis KIBGE-IB25. Int J Biol Macromol 78:243–248
Acknowledgements
The study was supported by the National Natural Science Foundation of China (Grant No. 81573399).
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Zhang, YQ., Li, RH., Zhang, HB. et al. Purification, characterization, and application of a thermostable dextranase from Talaromyces pinophilus . J Ind Microbiol Biotechnol 44, 317–327 (2017). https://doi.org/10.1007/s10295-016-1886-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10295-016-1886-8