Abstract
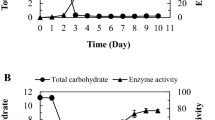
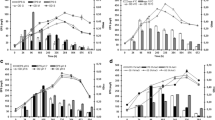
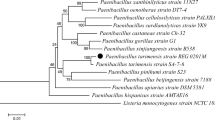
A psychrotolerant dextranase-producing bacterium was isolated from the Gaogong island seacoast near Jiangsu, China. The bacterium, denoted as DP03, was identified as Catenovulum sp. based on its phenotype, biochemical characteristics, and 16S rRNA gene comparison. The optimal enzyme production time, initial pH, temperature, and aeration conditions of strain DP03 were found to be 28 h, 8.0, 30 °C, and 25 % volume of liquid in 100-ml Erlenmeyer flasks, respectively. The ability of 1 % dextran T20 to induce dextranase was investigated. Dextranase from strain DP03 displayed its maximum activity at pH 8.0 and 40 °C and was found to be stable at 30 °C and over a broad range of pH values (pH 6–11). Scanning electron microscopy showed that dextranase from the isolate DP03 could at least partially prevent Streptococcus mutans from forming biofilms on glass coverslips.








Similar content being viewed by others
References
Bowler G, Wones S (2011) Application of dextranase in UK sugar beet factories. Zuckerindustrie 136(12):780–783
Cieslinski H, Kur J, Bialkowska A, Baran I, Makowski K, Turkiewicz M (2005) Cloning, expression, and purification of a recombinant cold-adapted beta-galactosidase from antarctic bacterium Pseudoalteromonas sp 22b. Protein Expres Purif 39(1):27–34
Esawy MA, Mansour SH, Ahmed EF, Hassanein NM, El Enshasy HA (2012) Characterization of extracellular dextranase from a novel Halophilic bacillus subtilis NRC-B233b a mutagenic honey isolate under solid state fermentation. E-J Chem 9(3):1494–1510
Galvez-Mariscal A, Lopez-Munguia A (1991) Production and characterization of a dextranase from an isolated Paecilomyces-lilacinus strain. Appl Microbiol Biotechnol 36(3):327–331
Gulder TM, Moore BS (2009) Chasing the treasures of the sea—bacterial marine natural products. Curr Opin Microbiol 12(3):252–260
Hild E, Brumbley SM, O’Shea MG, Nevalainen H, Bergquist PL (2007) A Paenibacillus sp dextranase mutant pool with improved thermostability and activity. Appl Microbiol Biotechnol 75(5):1071–1078
Kalpana BJ, Aarthy S, Pandian SK (2012) Antibiofilm activity of α-amylase from Bacillus subtilis S8-18 against biofilm forming human bacterial pathogens. Biotechnol Appl Biochem 167(6):1778–1794
Keyes PH, Hicks MA, Goldman M, McCabe RM, Fitzgerald RJ (1971) 3. Dispersion of dextranous bacterial plaques on human teeth with dextranase. J Am Dent Assoc 82(1):136–141
Khalikova E, Susi P, Korpela T (2005) Microbial dextran-hydrolyzing enzymes: fundamentals and applications. Microbiol Mol Biol R 69(2):306–325
Lu MS, Fang Y, Li H, Liu H, Wang S (2010) Isolation of a novel cold-adapted amylase-producing bacterium and study of its enzyme production conditions. Ann Microbiol 60(3):557–563
Majeed A, Grobler SR, Moola MH (2011) The pH of various tooth-whitening products on the South African market. SADJ 66(6):278–281
Margesin R, Fauster V, Fonteyne PA (2005) Characterization of cold-active pectate lyases from psychrophilic Mrakia frigida. Lett Appl Microbiol 40(6):453–459
Marotta M, Martino A, De Rosa A, Farina E, Carteni M, De Rosa M (2002) Degradation of dental plaque glucans and prevention of glucan formation using commercial enzymes. Process Biochem 38(1):101–108
Mehvar R (2000) Dextrans for targeted and sustained delivery of therapeutic and imaging agents. J Control Release 69(1):1–25
Ota F, Fukui K (1982) Scanning electron microscopic studies of the extracellular polysaccharides (EP) synthesized in colonies of Streptococcus mutans: development of EP and the effect of dextranase on them. Microbiol Immunol 26(7):623–628
Papaleo E, Tiberti M, Invernizzi G, Pasi M, Ranzani V (2011) Molecular determinants of enzyme cold adaptation: comparative structural and computational studies of cold- and warm-adapted enzymes. Curr Protein Pept Sci 12(7):657–683
Park T-S, Jeong HJ, Ko J-A, Ryu YB, Park S-J, Kim D, Kim Y-M, Lee WS (2012) Biochemical characterization of thermophilic dextranase from a thermophilic bacterium, Thermoanaerobacter pseudethanolicus. J Microbiol Biotechnol 22(5):637–641
Purushe S, Prakash D, Nawani NN, Dhakephalkar P, Kapadnis B (2012) Biocatalytic potential of an alkalophilic and thermophilic dextranase as a remedial measure for dextran removal during sugar manufacture. Bioresour Technol 115:2–7
Rahim ZHA, Thurairajah N (2011) Scanning electron microscopic study of Piper betle L. leaves extract effect against Streptococcus mutans ATCC 25175. J Appl Oral Sci 19(2):137–146
Schilling KM, Bowen WH (1992) Glucans synthesized in situ in experimental salivary pellicle function as specific binding-sites for streptococcus-mutans. Infect Immun 60(1):284–295
Staat RH, Schachtele CF (1976) Analysis of the dextranase activity produced by an oral strain of Bacteroides ochraceus. J Dent Res 55(6):1103–1110
Tao R, Tong Z, Lin Y, Xue Y, Wang W, Kuang R, Wang P, Tian Y, Ni L (2011) Antimicrobial and antibiofilm activity of pleurocidin against cariogenic microorganisms. Peptides 32(8):1748–1754
Thompson JD, Higgins DG, Gibson TJ (1994) Clustal-W—improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position-specific gap penalties and weight matrix choice. Nucleic Acids Res 22(22):4673–4680
Wu D-T, Zhang H-B, Huang L-J, Hu X-Q (2011) Purification and characterization of extracellular dextranase from a novel producer, Hypocrea lixii F1002, and its use in oligodextran production. Process Biochem 46(10):1942–1950
Wynter CVA, Chang M, DeJersey J, Patel B, Inkerman PA, Hamilton S (1997) Isolation and characterization of a thermostable dextranase. Enzyme Microb Tech 20(4):242–247
Yan S, Yu M, Wang Y, Shen C, Zhang XH (2011) Catenovulum agarivorans gen. nov., sp nov., a peritrichously flagellated, chain-forming, agar-hydrolysing gammaproteobacterium from seawater. Int J Syst Evol Microbiol 61(12):2866–2873
Acknowledgments
This work was financially supported by National 863 Program Foundation of China (2011AA09070302) and national funding project for local universities’ development.
Author information
Authors and Affiliations
Corresponding authors
Electronic supplementary material
Below is the link to the electronic supplementary material.
ESM 1
(DOC 160 kb)
Rights and permissions
About this article
Cite this article
Cai, R., Lu, M., Fang, Y. et al. Screening, production, and characterization of dextranase from Catenovulum sp.. Ann Microbiol 64, 147–155 (2014). https://doi.org/10.1007/s13213-013-0644-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13213-013-0644-7