Abstract
Main conclusion
MpBHY codes for a carotene β-ring 3(,3′)-hydroxylase responsible for both zeaxanthin and lutein biosynthesis in liverwort. MpCYP97C functions as an ε-ring hydroxylase (zeinoxanthin 3′-hydroxylase) to produce lutein in liverwort.
Abstract
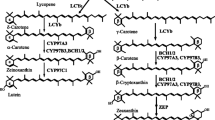
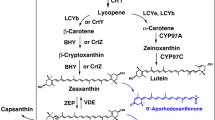
Xanthophylls are oxygenated or hydroxylated carotenes that are most abundant in the light-harvesting complexes of plants. The plant-type xanthophylls consist of α-xanthophyll (lutein) and β-xanthophylls (zeaxanthin, antheraxanthin, violaxanthin and neoxanthin). The α-xanthophyll and β-xanthophylls are derived from α-carotene and β-carotene by carotene hydroxylase activities, respectively. β-Ring 3,3′-hydroxylase that mediates the route of zeaxanthin from β-carotene via β-cryptoxanthin is present in higher plants and is encoded by the BHY (BCH) gene. On the other hand, CYP97A (or BHY) and CYP97C genes are responsible for β-ring 3-hydroxylation and ε-ring 3′-hydroxylation, respectively, in routes from α-carotene to lutein. To elucidate the evolution of the biosynthetic routes of such hydroxylated carotenoids from carotenes in land plants, we identified and functionally analyzed carotenoid hydroxylase genes of liverwort Marchantia polymorpha L. Three genes homologous to higher plants, BHY, CYP97A, and CYP97C, were isolated and named MpBHY, MpCYP97A, and MpCYP97C, respectively. MpBHY was found to code for β-ring hydroxylase, which is responsible for both routes starting from β-carotene and α-carotene. MpCYP97C functioned as an ε-ring hydroxylase not for α-carotene but for zeinoxanthin, while MpCYP97A showed no hydroxylation activity for β-carotene or α-carotene. These findings suggest the original functions of the hydroxylation enzymes of carotenes in land plants, which are thought to diversify in higher plants. In addition, we generated recombinant Escherichia coli cells, which produced rare and novel carotenoids such as α-echinenone and 4-ketozeinoxanthin, through pathway engineering using bacterial carotenogenic genes that include crtW, in addition to the liverwort MpLCYb, MpLCYe and MpBHY genes.







Similar content being viewed by others
Abbreviations
- BCH:
-
β-Carotene 3,3′-hydroxylase
- BHY:
-
β-Carotene 3,3′-hydroxylase
- CD:
-
Circular dichroism
- CYP:
-
Cytochrome P450
- LCYb:
-
Lycopene β-cyclase
- LCYe:
-
Lycopene ε-cyclase
- NMR:
-
Nuclear magnetic resonance
References
Albrecht M, Takaichi S, Misawa N, Schnurr G, Böger P, Sandmann G (1997) Synthesis of atypical cyclic and acyclic hydroxy carotenoids in Escherichia coli transformants. J Biotechnol 58:177–185
Bak S, Beisson F, Bishop G, Hamberger B, Höfer R, Paquette S, Werck-Reichhart D (2011) Cytochromes P450. Arabidopsis Book. doi:10.1199/tab.0144
Breitenbach J, Bai C, Rivera SM, Canela R, Capell T, Christou P, Zhu C, Sandmann G (2014) A novel carotenoid, 4-keto-α-carotene, as an unexpected by-product during genetic engineering of carotenogenesis in rice callus. Phytochemistry 98:85–91
Britton G, Liaaen-Jensen S, Pfander H (2004) Carotenoids handbook. Birkhäuser Verlag, Basel
Castenmiller JJ, West CE (1998) Bioavailability and bioconversion of carotenoids. Annu Rev Nutr 18:19–38
Chapple C (1998) Molecular-genetic analysis of plant cytochrome P450-dependent monooxygenases. Annu Rev Plant Physiol Plant Mol Biol 49:311–343
Christopher MJ, Waterman MR (1998) NADPH-flavodoxin reductase and flavodoxin from Escherichia coli: characteristics as a soluble microsomal P450 reductase. Biochemistry 37:6106–6113
Cui H, Yu X, Wang Y, Cui Y, Li X, Liu Z, Qin S (2013) Evolutionary origins, molecular cloning and expression of carotenoid hydroxylases in eukaryotic photosynthetic algae. BMC Genom 14:457–476
Cunningham FX Jr, Chamovitz D, Misawa N, Gantt E, Hirschberg J (1993) Cloning and functional expression in Escherichia coli of a cyanobacterial gene for lycopene cyclase, the enzyme that catalyzes the biosynthesis of β-carotene. FEBS Lett 328:130–138
Dall’Osto L, Fiore A, Cazzaniga S, Giuliano G, Bassi R (2007) Different roles of α- and β-branch xanthophylls in photosystem assembly and photoprotection. J Biol Chem 282:35056–35068
Das A, Yoon S-H, Lee S-H, Kim J-Y, Oh D-K, Kim S-W (2007) An update on microbial carotenoid production: application of recent metabolic engineering tools. Appl Microbiol Biotechnol 77:505–512
Fraser PD, Miura Y, Misawa N (1997) In vitro characterization of astaxanthin biosynthetic enzymes. J Biol Chem 272:6128–6135
Fraser PD, Elisabete M, Pinto S, Holloway DE, Bramley PM (2000) Application of high-performance liquid chromatography with photodiode array detection to the metabolic profiling of plant isoprenoids. Plant J 24:551–558
Galpaz N, Ronen G, Khalfa Z, Zamir D, Hirschberg J (2006) A chromoplast-specific carotenoid biosynthesis pathway is revealed by cloning of the tomato white-flower locus. Plant Cell 18:1947–1960
Hannemann F, Bichet A, Ewen KM, Bernhardt R (2007) Cytochrome P450 systems-biological variations of electron transport chains. Biochim Biophys Acta 1770:330–344
Harada H, Shindo K, Iki K, Teraoka A, Okamoto S, Yu F, Hattan J, Utsumi R, Misawa N (2011) Efficient functional analysis system for cyanobacterial or plant cytochromes P450 involved in sesquiterpene biosynthesis. Appl Microbiol Biotechnol 90:467–476
Harada H, Maoka T, Osawa A, Hattan J, Kanamoto H, Shindo K, Otomatsu T, Misawa N (2014) Construction of transplastomic lettuce (Lactuca sativa) dominantly producing astaxanthin fatty acid esters and detailed chemical analysis of generated carotenoids. Transgenic Res 23:303–315
Hasunuma T, Miyazawa S, Yoshimura S, Shinzaki Y, Tomizawa K, Shindo K, Choi SK, Misawa N, Miyake C (2008) Biosynthesis of astaxanthin in tobacco leaves by transplastomic engineering. Plant J 55:857–868
Hull AK, Celenza JL (2000) Bacterial expression and purification of the Arabidopsis NADPH-cytochrome P450 reductase ATR2. Protein Expr Purif 18:310–315
Iwamoto T, Hosoda K, Hirano R, Kurata H, Matsumoto A, Miki W, Kamiyama M, Itakura H, Yamamoto S, Kondo K (2000) Inhibition of low-density lipoprotein oxidation by astaxanthin. J Atheoscler Thomb 7:216–222
Kajiwara S, Fraser PD, Kondo K, Misawa N (1997) Expression of an exogenous isopentenyl diphosphate isomerase gene enhances isoprenoid biosynthesis in Escherichia coli. Biochem J 324:421–426
Kim J, DellaPenna D (2006) Defining the primary route for lutein synthesis in plants: the role of Arabidopsis carotenoid β-ring hydroxylase CYP97A3. Proc Natl Acad Sci 103:3474–3479
Kim I-J, Ko K-C, Kim C-S, Chung W-I (2001) Isolation and characterization of cDNAs encoding β–carotene hydroxylase in Citrus. Plant Sci 161:1005–1010
Kim J, Smith JJ, Tian L, DellaPenna D (2009) The evolution and function of carotenoid hydroxylases in Arabidopsis. Plant Cell Physiol 50:463–479
Kim J-E, Cheng KM, Craft NE, Hamberger B, Douglas CJ (2010) Over-expression of Arabidopsis thaliana carotenoid hydroxylases individually and in combination with a β-carotene ketolase provides insight into in vivo function. Phytochemistry 71:168–178
Krinsky NI, Landrum JT, Bone RA (2003) Biological mechanisms of the protective role of lutein and zeaxanthin in the eye. Annu Rev Nutr 23:171–201
Lee PC, Schmidt-Danner C (2002) Metabolic engineering towards biotechnological production of carotenoids in microorganisms. Appl Microbiol Biotechnol 60:1–11
Li Q, Farre G, Naqvi S, Breitenbach J, Sanahuja G, Bai C, Sandmann G, Capell T, Christou P, Zhu C (2010) Cloning and functional characterization of the maize carotenoid isomerase and β–carotene hydroxylase genes and their regulation during endosperm maturation. Transgenic Res 19:1053–1068
Lv M-Z, Chao D-Y, Shan J-X, Zhu M-Z, Shi M, Gao J-P, Lin H-X (2012) Rice carotenoid β-ring hydroxylase CYP97A4 is involved in lutein biosynthesis. Plant Cell Physiol 53:987–1002
Maoka T, Takemura M, Tokuda H, Suzuki N, Misawa N (2014) 4-Ketozeinoxanthin, a novel carotenoid produced in Escherichia coli through metabolic engineering using carotenogenic genes of bacterium and liverwort. Tetrahedron Lett 55:6708–6710
Mayne ST (1996) Beta-carotene, carotenoids, and disease prevention in humans. FASEB J 10:690–701
Misawa N (2011) Pathway engineering for functional isoprenoids. Curr Opin Biotechnol 22:627–633
Misawa N, Nakagawa M, Kobayashi K, Yamano S, Izawa Y, Nakamura K, Harashima K (1990) Elucidation of the Erwinia uredovora carotenoid biosynthetic pathway by functional analysis of gene products expressed in Escherichia coli. J Bacteriol 172:6704–6712
Misawa N, Satomi Y, Kondo K, Yokoyama A, Kajiwara S, Saito T, Ohtani T, Miki W (1995) Structure and functional analysis of a marine bacterial carotenoid biosynthesis gene cluster and astaxanthin biosynthetic pathway proposed at the gene level. J Bacteriol 177:6575–6584
Moore RC, Purugganan MD (2005) The evolutionary dynamics of plant duplicate genes. Curr Opin Plant Biol 8:122–128
Nishida Y, Adachi K, Kasai H, Shizuri Y, Shindo K, Sawabe A, Komemushi S, Miki W, Misawa N (2005) Elucidation of a carotenoid biosynthesis gene cluster encoding a novel enzyme, 2,2′-β-hydroxylase, from Brevundimonas sp. strain SD212 and combinatorial biosynthesis of new or rare xanthophylls. Appl Environ Microbiol 71:4286–4296
Nishino H, Tokuda H, Murakoshi M, Satomi Y, Masuda M, Onozuka M, Yamaguchi S, Takayasu J, Tsuruta J, Okuda M, Khachik F, Narisawa T, Takasuka N, Yano M (2000) Cancer prevention by natural carotenoids. BioFactors 13:89–94
Niyogi KK, Shih C, Soon Chow W, Pogson BJ, DellaPenna D, Bjökman O (2001) Photoprotection in a zeaxanthin- and lutein- deficient double mutant of Arabidopsis. Photosynth Res 67:139–145
Qin X, Zhang W, Dubcovsky J, Tian L (2012) Cloning and comparative analysis of carotenoid β-hydroxylase genes provides new insights into carotenoid metabolism in tetraploid (Triticum turgidum ssp. durum) and hexaploid (Triticum aestivum) wheat grains. Plant Mol Biol 80:631–646
Quinlan RF, Jaradat TT, Wurtzel ET (2007) Escherichia coli as a platform for functional expression of plant P450 carotene hyrdoxylases. Arch Biochem Biophys 458:146–157
Quinlan RF, Shumskaya M, Bradbury LMT, Beltrán J, Ma C, Kennelly EJ, Wurtzel ET (2012) Synergistic interactions between carotene ring hydroxylases drive lutein formation in plant carotenoid biosynthesis. Plant Physiol 160:204–214
Ruther A, Misawa N, Böger P, Sandman G (1997) Production of zeaxanthin in Escherichia coli transformed with different carotenogenic plasmids. Appl Microbiol Biotechnol 48:162–167
Schückel J, Rylott EL, Grogan G, Bruce NC (2012) A gene-fusion approach to enabling plant cytochromes p450 for biocatalysis. Chem BioChem 13:2758–2763
Shindo K, Hasunuma T, Asagi E, Sano A, Hotta E, Minemura N, Miyake C, Maoka T, Misawa N (2008) 4-Ketoantheraxanthin, a novel carotenoid produced by the combination of the bacterial enzyme β-carotene ketolase CrtW and endogenous carotenoid biosynthetic enzymes in higher plants. Tetrahedron Lett 49:3294–3296
Stigliani AL, Giorio G, D’Ambrosio C (2011) Characterization of P450 carotenoid β- and ε-hydroxylases of tomato and transcriptional regulation of xanthophyll biosynthesis in root, leaf, petal and fruit. Plant Cell Physiol 52:851–865
Sugiura M, Nakamura M, Ogawa K, Ikoma M, Yano N (2012) High serum carotenoids associated with lower risk for bone loss and osteoporosis in post-menopausal Japanese female subjects: prospective cohort study. PLoS One 7:e52643
Sun Z, Gantt E, Cunningham FX Jr (1996) Cloning and functional analysis of the β-carotene hydroxylase of Arabidopsis thaliana. J Biol Chem 271:24349–24352
Takaichi S (2011) Carotenoids in algae: distributions, biosyntheses and functions. Mar Drugs 9:1101–1118
Takemura M, Maoka T, Misawa N (2014) Carotenoid analysis of a liverwort Marchantia polymorpha and functional identification of its lycopene β- and ε-cyclase genes. Plant Cell Physiol 55:194–200
Talegawkar SA, Johnson EJ, Carithers TC, Taylor HA Jr, Bogle ML, Tucker KL (2008) Carotenoid intakes, assessed by food-frequency questionnaires (FFQs), are associated with serum carotenoid concentrations in the Jackson Heart Study: validation of the Jackson Heart Study Delta NIRI Adult FFQs. Public Health Nutr 11:989–997
Tamura K, Stecher G, Peterson D, Filipski A, Kumar S (2013) MEGA6: molecular evolutionary genetics analysis version 6.0. Mol Biol Evol 30:2725–2729
Tian L, DellaPenna D (2001) Characterization of a second carotenoid β-hydroxylase gene from Arabidopsis and its relationship to the LUT1 locus. Plant Mol Biol 47:379–388
Tian L, Magallanes-Lundback M, Musetti V, DellaPenna D (2003) Functional analysis of β- and ε-ring carotenoid hydroxylases in Arabidopsis. Plant Cell 15:1320–1332
Tian L, Musetti V, Kim J, Magallanes-Lundback M, DellaPenna D (2004) The Arabidopsis LUT1 locus encodes a member of the cytochrome P450 family that is required for carotenoid ε-ring hydroxylation activity. Proc Natl Acad Sci 101:402–407
Urban P, Mignotte C, Kazmaier M, Delorme F, Pompon D (1997) Cloning, yeast expression, and characterization of the coupling of two distantly related Arabidopsis thaliana NADPH-cytochrome P450 reductases with P450 CYP73A5. J Biol Chem 272:19176–19186
Vallabhaneni R, Gallagher CE, Licciardello N, Cuttriss AJ, Quinlan RF, Wurtzel ET (2009) Metabolite sorting of a germplasm collection reveals the Hydroxylase3 locus as a new target for maize provitamin A biofortification. Plant Physiol 151:1635–1645
Yang L-E, Huang X-Q, Hang Y, Deng Y-Y, Lu Q-Q, Lu S (2014) The P450-type carotene hydroxylase PuCHY1 from Porphyra suggests the evolution of carotenoid metabolism in red algae. J Integr Plant Biol 56:902–915
Ye VM, Bhatia SK (2012) Pathway engineering strategies for production of beneficial carotenoids in microbial hosts. Biotechnol Lett 34:1405–1414
Yokoyama A, Miki W (1995) Composition and presumed biosynthetic pathway of carotenoids in the astaxanthin-producing bacterium Agrobacterium aurantiacum. FEMS Microbiol Lett 128:139–144
Yokoyama A, Shizuri Y, Misawa N (1998) Production of new carotenoids, astaxanthin glucosides, by Escherichia coli transformants carrying carotenoid biosynthetic genes. Tetrahedron Lett 39:3709–3712
Acknowledgments
The authors thank Dr. Hiroshi Shimada, Hiroshima University, for the construction of plasmid pACHP-Beta. We also thank Dr. Takayuki Kohchi, Kyoto University; Dr. Katsuyuki T. Yamato, Kinki University and Dr. Kimitsune Ishizaki, Kobe University for their assistance in searching liverwort sequences.
Conflict of interest
The authors declare that they have no conflict of interest.
Author information
Authors and Affiliations
Corresponding author
Additional information
Gene accession numbers
The nucleotide sequence reported in this paper has been submitted to DDBJ under accession numbers, AB981062 (MpBHY), AB981063 (MpCYP97A), AB981064 (MpCYP97B) and AB981065 (MpCYP97C).
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Takemura, M., Maoka, T. & Misawa, N. Biosynthetic routes of hydroxylated carotenoids (xanthophylls) in Marchantia polymorpha, and production of novel and rare xanthophylls through pathway engineering in Escherichia coli . Planta 241, 699–710 (2015). https://doi.org/10.1007/s00425-014-2213-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00425-014-2213-0