Abstract
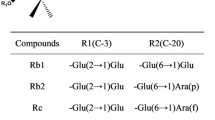
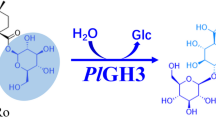
An eco-friendly and convenient preparation method for notoginsenoside ST-4 has been established by completely transforming vina-ginsenoside R7 using a recombinant glycosidase hydrolyzing enzyme (HaGH03) from Herpetosiphon aurantiacus. This enzyme specifically hydrolyzed the glucose at the C-20 position but not the external xylose or two inner glucoses at position C-3. Protein sequence BLAST revealed that HaGH03, composed of 749 amino acids and presumptively listed as a member of the family 3 glycoside hydrolases, has highest identity (48 %) identity with a thermostable β-glucosidase B, which was not known of any functions for ginsenoside transformation. The steady state kinetic parameters for purified HaGH03 measured against p-nitrophenyl β-D-glucopyranoside and vina-ginsenoside R7 were K M = 5.67 ± 0.24 μM and 0.59 ± 0.23 mM, and k cat = 69.2 ± 0.31/s and 2.15 ± 0.46/min, respectively. HaGH03 converted 2.5 mg/mL of vina-ginsenoside R7 to ST-4 with a molar yield of 100 % and a space-time yield of 104 mg/L/h in optimized conditions. These results underscore that HaGH03 has much potential for the effective preparation of target ginsenosides possessing valuable pharmacological activities. This is the first report identifying an enzyme that has the ability to transform vina-ginsenoside R7 and provides an approach to preparing rare notoginsenoside ST-4.






Similar content being viewed by others
References
An DS, Cui CH, Lee HG, Wang L, Kim SC, Lee ST, Jin FX, Yu HS, Chin YW, Lee HK, Im WT, Kim SG (2010) Identification and characterization of a novel Terrabacter ginsenosidimutans sp nov β-Glucosidase that transforms Ginsenoside Rb1 into the rare Gypenosides XVII and LXXV. Appl Environ Microbiol 76:5827–5836
Attele AS, Wu JA, Yuan CS (1999) Ginseng pharmacology: multiple constituents and multiple actions. Biochem Pharmacol 58:1685–1693
Castle LA, Smith KD, Morris RO (1992) Cloning and sequencing of an Agrobacterium tumefaciens beta-glucosidase gene involved in modifying a vir-inducing plant signal molecule. J Bacteriol 174:1478–1486
Chang KH, Jee HS, Lee NK, Park SH, Lee NW, Paik HD (2009) Optimization of the enzymatic production of 20(S)-ginsenoside Rg3 from white ginseng extract using response surface methodology. New Biotechnol 26:181–186
Cui CH, Kim SC, Im WT (2013a) Characterization of the ginsenoside-transforming recombinant β-glucosidase from Actinosynnema mirum and bioconversion of major ginsenosides into minor ginsenosides. Appl Microbiol Biotechnol 97:649–659
Cui CH, Liu QM, Kim JK, Sung BH, Kim SG, Kim SC, Im WT (2013b) Identification and characterization of a Mucilaginibacter sp Strain QM49 β-glucosidase and its use in the production of the pharmaceutically active minor Ginsenosides (S)-Rh1 and (S)-Rg2. Appl Environ Microbiol 79:5788–5798
Dan M, Su M, Gao X, Zhao T, Zhao A, Xie G, Qiu Y, Zhou M, Liu Z, Jia W (2008) Metabolite profiling of Panax notoginseng using UPLC–ESI-MS. Phytochemistry 69:2237–2244
Gao B, Huang L, Liu H, Wu H, Zhang E, Yang L, Wu X, Wang Z (2014a) Platelet P2Y12 receptors are involved in the haemostatic effect of notoginsenoside Ft1, a saponin isolated from Panax notoginseng. Br J Pharmacol 171:214–223
Gao B, Shi HL, Li X, Qiu SP, Wu H, Zhang BB, Wu XJ, Wang ZT (2014b) p38 MAPK and ERK1/2 pathways are involved in the pro-apoptotic effect of notoginsenoside Ft1 on human neuroblastoma SH-SY5Y cells. Life Sci 108:63–70
Groussin AL, Antoniotti S (2012) Valuable chemicals by the enzymatic modification of molecules of natural origin: terpenoids, steroids, phenolics and related compounds. Bioresour Technol 115:237–243
Jin S, Luo M, Wang W, Zhao CJ, Gu CB, Li CY, Zu YG, Fu YJ, Guan Y (2013) Biotransformation of polydatin to resveratrol in Polygonum cuspidatum roots by highly immobilized edible Aspergillus niger and Yeast. Bioresour Technol 136:766–770
Kim JK, Cui CH, Yoon MH, Kim SC, Im WT (2012) Bioconversion of major ginsenosides Rg1 to minor ginsenoside F1 using novel recombinant ginsenoside hydrolyzing glycosidase cloned from Sanguibacter keddieii and enzyme characterization. J Biotechnol 161:294–301
Kim JK, Cui CH, Liu Q, Yoon MH, Kim SC, Im WT (2013) Mass production of the ginsenoside Rg3(S) through the combinative use of two glycoside hydrolases. Food Chem 141:1369–1377
Larsbrink J, Rogers TE, Hemsworth GR, McKee LS, Tauzin AS, Spadiut O, Klinter S, Pudlo NA, Urs K, Koropatkin NM, Creagh AL, Haynes CA, Kelly AG, Cederholm SN, Davies GJ, Martens EC, Brumer H (2014) A discrete genetic locus confers xyloglucan metabolism in select human gut Bacteroidetes. Nature 506:498–502
Lee CH, Kim JH (2014) A review on the medicinal potentials of ginseng and ginsenosides on cardiovascular diseases. J Ginseng Res 38:161–166
Lee GW, Kim KR, Oh DK (2012) Production of rare ginsenosides (compound Mc, compound Y and aglycon protopanaxadiol) by β-glucosidase from Dictyoglomus turgidum that hydrolyzes β-linked, but not α-linked, sugars in ginsenosides. Biotechnol Lett 34:1679–1686
Lee GW, Yoo MH, Shin KC, Kim KR, Kim YS, Lee KW, Oh DK (2013) β-Glucosidase from Penicillium aculeatum hydrolyzes exo-, 3-O-, and 6-O-β-glucosides but not 20-O-β-glucoside and other glycosides of ginsenosides. Appl Microbiol Biotechnol 97:6601
Lee HJ, Shin KC, Lee GW, Oh DK (2014) Production of aglycone protopanaxatriol from ginseng root extract using Dictyoglomus turgidum β-glycosidase that specifically hydrolyzes the xylose at the C-6 position and the glucose in protopanaxatriol-type ginsenosides. Appl Microbiol Biotechnol 98:3659–3667
Liu ZQ (2012) Chemical insights into ginseng as a resource for natural antioxidants. Chem Rev 112:3329–3355
Liu C, Jin Y, Yu H, Sun C, Gao P, Xiao Y, Zhang T, Xu L, Im WT, Jin F (2014) Biotransformation pathway and kinetics of the hydrolysis of the 3-O- and 20-O-multi-glucosides of PPD-type ginsenosides by ginsenosidase type I. Process Biochem 49:813–820
Minh DN, Kasai R, Ohtani K, Ito A, Thoi Nham N, Yamasaki K, Tanaka O (1994) Saponins from Vietnamese ginseng, Panax vietnamensis Ha et Grushv. Collected in central Vietnam. II. Chem Pharm Bull 42:115–122
Ng TB (2006) Pharmacological activity of sanchi ginseng (Panax notoginseng). J Pharm Pharmacol 58:1007–1019
Park CS, Yoo MH, Noh KH, Oh DK (2010) Biotransformation of ginsenosides by hydrolyzing the sugar moieties of ginsenosides using microbial glycosidases. Appl Microbiol Biotechnol 87:9–19
Pei Y, Du Q, Liao PY, Chen ZP, Wang D, Yang CR, Kitazato K, Wang YF, Zhang YJ (2011) Notoginsenoside ST-4 inhibits virus penetration of herpes simplex virus in vitro. J Asian Nat Prod Res 13:498–504
Pozzo T, Pasten JL, Karlsson EN, Logan DT (2010) Structural and functional analyses of β-glucosidase 3B from Thermotoga neapolitana: a thermostable three-domain representative of glycoside hydrolase 3. J Mol Biol 397:724–739
Quan LH, Min JW, Yang DU, Kim YJ, Yang DC (2012) Enzymatic biotransformation of ginsenoside Rb1 to 20(S)-Rg3 by recombinant β-glucosidase from Microbacterium esteraromaticum. Appl Microbiol Biotechnol 94:377–384
Robert X, Gouet P (2014) Deciphering key features in protein structures with the new ENDscript server. Nucleic Acids Res. doi:10.1093/nar/gku316
Shen KK, Ji LL, Gong CY, Ma YB, Yang L, Fan Y, Hou MQ, Wang ZT (2012) Notoginsenoside Ft1 promotes angiogenesis via HIF-1α mediated VEGF secretion and the regulation of PI3K/AKT and Raf/MEK/ERK signaling pathways. Biochem Pharmacol 84:784–792
Shen KK, Leung SWS, Ji LL, Huang Y, Hou MQ, Xu AM, Wang ZT, Vanhoutte PM (2014) Notoginsenoside Ft1 activates both glucocorticoid and estrogen receptors to induce endothelium-dependent, nitric oxide-mediated relaxations in rat mesenteric arteries. Biochem Pharmacol 88:66–74
Shin KC, Oh HJ, Kim BJ, Oh DK (2013) Complete conversion of major protopanaxadiol ginsenosides to compound K by the combined use of α-l-arabinofuranosidase and β-galactosidase from Caldicellulosiruptor saccharolyticus and β-glucosidase from Sulfolobus acidocaldarius. J Biotechnol 167:33–40
Shin KC, Seo MJ, Oh HJ, Oh DK (2014) Highly selective hydrolysis for the outer glucose at the C-20 position in ginsenosides by β-glucosidase from Thermus thermophilus and its application to the production of ginsenoside F2 from gypenoside XVII. Biotechnol Lett 36:1287–1293
Su JH, Xu JH, Lu WY, Lin GQ (2006) Enzymatic transformation of ginsenoside Rg3 to Rh2 using newly isolated Fusarium proliferatum ECU2042. J Mol Catal B Enzym 38:113–118
Su JH, Xu JH, Yu HL, He YC, Lu WY, Lin GQ (2009) Properties of a novel glucosidase from Fusarium proliferatum ECU2042 that converts ginsenoside Rg3 into Rh2. J Mol Catal B Enzym 57:278–283
Tawab MA, Bahr U, Karas M, Wurglics M, Schubert ZM (2003) Degradation of ginsenosides in humans after oral administration. Drug Metab Dispos 31:1065–1071
Wang CZ, McEntee E, Wicks S, Wu JA, Yuan CS (2006) Phytochemical and analytical studies of Panax notoginseng (Burk.) F.H. Chen. J Nat Med 60:97–106
Wang XY, Wang D, Ma XX, Zhang YJ, Yang CR (2008) Two new dammarane-type bisdesmosides from the fruit pedicels of Panax notoginseng. Helv Chim Acta 91:60–66
Ye L, Zhou CQ, Zhou W, Zhou P, Chen DF, Liu XH, Shi XL, Feng MQ (2010) Biotransformation of ginsenoside Rb1 to ginsenoside Rd by highly substrate-tolerant Paecilomyces bainier 229-7. Bioresour Technol 101:7872–7876
Yoshida E, Hidaka M, Fushinobu S, Koyanagi T, Minami H, Tamaki H, Kitaoka M, Katayama T, Kumagai H (2010) Role of a PA14 domain in determining substrate specificity of a glycoside hydrolase family 3 β-glucosidase from Kluyveromyces marxianus. Biochem J 431:39–49
Acknowledgments
This work was financially supported by the National Natural Science Foundation of China (U1032604) and the Program for Changjiang Scholars and Innovative Research Team in University (IRT1071).
Author information
Authors and Affiliations
Corresponding authors
Electronic supplementary material
Below is the link to the electronic supplementary material.
ESM 1
(PDF 409 kb)
Rights and permissions
About this article
Cite this article
Wang, RF., Zheng, MM., Cao, YD. et al. Enzymatic transformation of vina-ginsenoside R7 to rare notoginsenoside ST-4 using a new recombinant glycoside hydrolase from Herpetosiphon aurantiacus . Appl Microbiol Biotechnol 99, 3433–3442 (2015). https://doi.org/10.1007/s00253-015-6446-z
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-015-6446-z