Abstract
Rhizobia that nodulate peas comprise a heterogeneous group of bacteria. The aim of this study was to investigate the relationship between phylogeny and electrophoretic and hydroxy fatty acid lipopolysaccharide (LPS) profiles of pea microsymbionts. Based on amplified fragment length polymorphism (AFLP) fingerprinting data, the pea microsymbionts were grouped into two clusters distinguished at 58% similarity level. Based on the concatenated 16S rRNA, recA, and atpD housekeeping gene data, the microsymbionts appeared to be most closely related to Rhizobium leguminosarum biovars viciae and trifolii. Applying cluster analysis to their LPS electrophoretic profiles, the strains were assigned to two major groups with different banding patterns. All hydroxy fatty acids common to R. leguminosarum and R. etli were detected in each examined strain. Differences in the proportions of 3- to ω-1 hydroxy fatty acids allowed us to distinguish two groups of strains. This classification did not overlap with one based on LPS electrophoretic profiles. No clear correlation was apparent between the genetic traits and LPS profiles of the pea nodule isolates.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Biological nitrogen fixation is one of the most important processes whereby reduced nitrogen compounds are introduced into the biosphere. Bacteria from the order Rhizobiales, both free-living and those forming a symbiotic relationship with legume plants, are capable of fixing atmospheric nitrogen (Zahran 1999). One of the striking features of symbiosis is the relatively high specificity of the interaction between the microsymbiont and the host plant. Most rhizobia have a narrow host range and form nodules on well-defined plant species, e.g., Rhizobium leguminosarum bv. viciae is a symbiont of the legumes of the tribe Vicieae, which includes the genera Vicia, Pisum, Lathyrus, and Lens, while the symbiosis of Rhizobium bv. trifolii is confined to Trifolium plants (Tian et al. 2010; Kumar et al. 2015).
Nodule development requires the exchange of molecular signals between the two partners, flavonoids (produced by plants) and Nod factors (produced by bacteria) that are recognized by plant receptors. This leads to the expression of plant genes, cell de-differentiation, organogenesis, and infection of root nodules (Skorupska et al. 2006; Zgadzaj et al. 2015).
In addition, partner recognition and effective symbiosis require an appropriate structure of surface polysaccharides, such as lipopolysaccharides (LPS), exopolysaccharides (EPS), external capsular polysaccharides (CPS or K-antigen polysaccharides, KPS), as well as periplasmic cyclic β-glucans, high molecular weight neutral polysaccharide (glucomannan), and gel-forming polysaccharide (GPS) (Laus et al. 2006; Janczarek 2011; Kawaharada et al. 2015).
The importance of the different types of polysaccharides in the nodulation process varies depending on the type of nodules (determinate or indeterminate). For instance, acidic EPS secreted into the extracellular environment is especially significant in the establishment of effective symbiosis with host plants that form indeterminate nodules (Hotter and Scott 1991). On the other hand, the presence of the O-chain portion of LPS is required for effective symbiosis in both determinate and indeterminate (Priefer 1989) nodule-forming hosts (Noel et al. 1986).
Metabolism-related traits and physiological characteristics are quite often used to study the diversity of rhizobia (Dresler-Nurmi et al. 2009). Native rhizobial populations are diverse and contain strains differing in their physiological features, genomic structure, and the efficiency of nitrogen fixation (Wielbo et al. 2010, 2011; Kumar et al. 2015). Accordingly, some LPS traits, such as their electrophoretic profile and fatty acid composition, may be used for taxonomic classification (Santamaria et al. 1997). The composition of both total cellular fatty acids and LPS fatty acids has been used for bacterial identification and taxonomy (Yokota and Sakane 1991; Dresler-Nurmi et al. 2009; Choma and Komaniecka 2011).
Recent studies have revealed particularly high diversity in the genome organization and metabolic versatility of R. leguminosarum isolates. Kumar et al. (2015) demonstrated that the diversity of R. leguminosarum within a local population of nodule isolates was 10 times higher than that found in Ensifer medicae.
The molecular methods used for the classification of bacterial species include 16S rRNA gene sequencing and phylogenetic analysis of housekeeping genes (Ramírez-Bahena et al. 2008). Amplified fragment length polymorphism (AFLP) analysis has also been used to evaluate the genomic diversity of nodule bacteria (Wolde-Meskel et al. 2004; Wdowiak-Wróbel and Małek 2005).
The aim of the current study was to investigate the diversity of strains nodulating Pisum sativum according to genetic markers and phenotypic properties. 16S rRNA, recA, and atpD gene sequences were used as genetic markers, and were also profiled by AFLP. The phenotypical assessment was based on electrophoretic patterns and fatty acid composition of LPS, and on EPS production. Additionally, such physiological criteria as sensitivity to salt, detergents, pH, and elevated temperature were studied.
Materials and methods
Bacterial strains and growth conditions
All strains used in this study (15 strains) were obtained from surface-sterilized nodules of pea (P. sativum) grown in arable soil in the region of Lublin, Poland, as described earlier (Wielbo et al. 2010, 2011). The isolated strains were reinoculated onto P. sativum cv. Ramrod to confirm their nodulation ability in sterile conditions. For further studies, the bacteria were grown either on yeast mannitol agar (79CA) plates (Vincent 1970) or in liquid 79CA medium, with 1% glycerol as a carbon source, at 28 °C with shaking (160 rpm).
AFLP analyses
The genome diversity of rhizobial strains was studied using a modified AFLP method, as described by Wdowiak-Wróbel and Małek (2005). Briefly, genomic DNA was digested with PstI and ligated to adaptors Pst_AR (5′-TGTACGCAGTCTACG-3′) and Pst_AF (5′-CTCGTAGACTGCGTACATGCA-3′) for 4 h at 37 °C. Next, it was subjected to a selective PCR using a PstI-GCG primer containing additional GCG nucleotides at the 3′-terminus (5′-GACTGCGTACATGCAGGCG-3′). The amplified products were separated by electrophoresis in 2% (w/v) agarose gel in Tris–borate-EDTA (TBE) buffer. AFLP profiles were used for the construction of a dendrogram, employing the unweighted pair-group method with arithmetic mean (UPGMA) clustering method in STATISTICA v. 10 for Windows (StatSoft Inc., Tulsa, OK).
PCR amplification and gene sequencing
Genomic DNA was extracted and purified as described previously (Marek-Kozaczuk et al. 2013). The 16S rRNA was amplified using 16S63f (5′-CAGGCCTAACACATGCAAGTC-3′) and 16S1387r (5′-GGGCGGWGTGTACAAGGC-3′) primers (Marchesi et al. 1998). The primers and protocols used for the amplification and sequencing of chromosomal housekeeping genes atpD, encoding the beta subunit of ATP-synthase (Bailly et al. 2007), and recA, encoding a DNA recombinase (Gaunt et al. 2001), were described earlier (Marek-Kozaczuk et al. 2013).
Sequencing was performed using the BigDye® Terminator Cycle Sequencing Kit (Applied Biosystems, Carlsbad, CA, USA) and an ABI Prism 3730 XL Genetic Analyzer (Applied Biosystems). The sequences were aligned with those deposited in the GenBank using the MEGA5.05 software package (Tamura et al. 2011). Distances were calculated according to Kimura’s two-parameter model of substitutions (Kimura 1980). Phylogenetic trees were constructed using the neighbor-joining (NJ) method. Bootstrap analysis was based on 1000 replications (Felsenstein 1985).
Nucleotide sequence accession numbers
The sequences have been submitted to the GenBank database under accession numbers KJ528929–KJ528943 for 16S rRNA genes; KJ528903–KJ528911, KJ493937–KJ493941, and KJ481208 for atpD; and KJ528913–KJ528927 for recA genes.
Sodium dodecyl sulfate (SDS)-tricine-polyacrylamide gel electrophoresis (PAGE)
LPS was prepared by whole-cell microextraction using proteinase K digestion. Bacterial cells were washed three times with 0.5 M NaCl, and wet cell mass samples (50 mg) were solubilized in 200 μL of lysing buffer according to Apicella (2008). Samples (5 μL) were loaded onto 12.5% SDS-PAGE gel. Electrophoresis was carried out using a tricine buffer system. LPS profiles were visualized as described by Tsai and Frasch (1982). The UPGMA method was used to construct a dendrogram of LPS profiles. The presence of a band was scored as 1 and its absence as 0. Bands with the same mobility, regardless of their intensity, were considered to be identical.
Analysis of LPS-derived fatty acids
Fatty acids were released (4 M HCl, 100 °C, 4.5 h) from LPS samples obtained from whole cell lysates. Free fatty acids were esterified by methanolysis (1 M methanolic HCl, 80 °C, 1.5 h). The samples were trimethylsilylated (TMS) with a Sylon HTP kit (Supelco, Bellefonte, PA, USA) for 30 min at room temperature. Methyl esters of nonpolar fatty acids and TMS ethers of hydroxy fatty acids were identified by gas chromatography–mass spectrometry (GC–MS) on an Agilent gas chromatograph (7890A, Santa Clara, CA, USA) equipped with a capillary column (HP-5MS, 30 m × 0.25 mm, Supelco, Bellefonte, PA, USA) connected to a mass selective detector (MSD5975, Agilent Technologies, Santa Clara, CA, USA).
Fatty acid weight percentage of total methyl esters was analyzed with the UPGMA method using unweighted pair-group average Euclidean distances, in the STATISTICA package.
Quantitation of EPS production
Three-day 79CA medium cultures of rhizobia were centrifuged at 14,000×g for 30 min, and EPS was precipitated from supernatants with three volumes of cold ethanol. The quantification of total carbohydrates, expressed as glucose equivalents, was performed by the anthrone method (Yasar 2005). The EPS yield was expressed as mg of EPS produced per mg of bacterial protein. Total protein was determined by the Lowry method after overnight solubilization of bacteria in 0.1 M NaOH and 5.0% SDS at room temperature.
Sensitivity to salt, detergents, pH, and elevated temperature
The ability of strains to grow in the presence of saline was determined in liquid 79CA medium supplemented with 0.01, 0.1, 1.0, and 3.0% (w/v) of NaCl. The bacterial growth was measured as optical density at 550 nm (OD550). The growth of each strain was tested at pH 4.5, 5, 7.2, 7.5, and 8.0, in liquid 79CA medium. Temperature ranges and growth optima of the isolates were determined by incubating the inoculated 79CA liquid medium cultures at 20, 28, 37, and 42 °C. Cell viability was tested by spreading 100 µL of undiluted cultures on 79CA agar plates.
Sensitivity to SDS and sodium deoxycholate (DOC) was determined using a dilution method in 96-well microtiter plates with serially diluted detergents at concentrations from 0.001 to 0.25% (w/v) in 79CA medium. The minimum inhibitory concentration (MIC) was defined as the lowest concentration of the detergents at which no bacterial growth was visible.
All samples were incubated for 3 days. The experiments were performed in triplicate.
Phenotypic data (sensitivity to salt, SDS, DOC, and pH) were analyzed by the UPGMA method using weighted pair-group average Euclidean distances.
Results
Analysis of the genetic diversity of pea nodule isolates
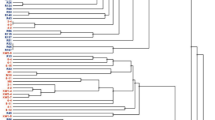
Based on cluster analysis of AFLP fingerprints with the unweighted pair-group method, all strains were grouped into two clusters at 58% similarity level (Fig. 1). Only two strains, GC 5.8 and GC 7.4, had identical DNA fingerprints. Strains GC 1.3 and GD 4 shared the highest similarity with the reference strain R. leguminosarum bv. viciae 3841 with 70 and 61% similarity, respectively. Strain GD 4 was most distant from all isolates. At the cut off value of 85%, the strains could be subdivided into three subgroups: group I—strains GB 42, GC 5.8, GC 7.4, P 1.37, and P 1.42, displaying 92% similarity; group II—strains GB 53, GD 31, GD 29, and P 2.24; and group III—formed by strains GC 5.5 and P 1.47, with 88% similarity.
The 16S rRNA gene-based phylogenetic tree was less discriminative and showed that the analyzed 15 pea microsymbionts clustered in a well-resolved branch with R. leguminosarum bv. viciae strains USDA 237 and 3841, and R. leguminosarum bv. trifolii ATCC 14480 with bootstrap support of 98% (Supplemental Fig. 1).
To further classify the isolates, the recA and atpD core genes were sequenced. The similarity values for the recA and atpD genes were high and ranged from 95.1 to 100% and from 92.7 to 100%, respectively (Supplemental Figs. 2, 3). The phylogenetic tree constructed on the basis of the concatenated genes 16S rRNA, recA, and atpD showed that all strains except GD 4 formed a cluster with the reference strains R. leguminosarum bv. trifolii ATCC 14480 and R. leguminosarum bv. viciae USDA 237 (Fig. 2) and confirmed that the P. sativum isolates were phylogenetically related to the biovars viciae and trifolii of R. leguminosarum. Isolate GD 4 showed the highest similarity, i.e., 98% sequence identity of combined gene sequences to R. leguminosarum bv. viciae 3841. A distinct branch at 99.5–99.6% sequence identity of the concatenated genes was formed by the three microsymbionts GD31, P1.42, and P2.24.
Electrophoretic analysis of LPS
Based on LPS electrophoretic profiles, the strains isolated from pea nodules were grouped into two major clusters at a similarity level of 58%, as determined by the UPGMA method (Fig. 3a, b). The largest cluster (cluster A) contained 10 isolates, including the reference strain 3841 (Table 1), with the LPS profile consisting of two major bands representing high molecular weight LPS I, i.e., LPS containing the O-polysaccharide, and LPS II, a low molecular weight LPS that contains lipid A and a core oligosaccharide (D’Heaze et al. 2007) (Fig. 3a). Four isolates, P 1.37, P 1.42, P 1.47, and GC 7.4, had the same LPS profile as strain 3841. Strains GB 42 and GC 1.3 formed a subcluster with 90% similarity, characterized by a less prominent LPS I region.
In the other cluster (cluster B), LPS of strains GC 5.5, GC 5.8, and GD 31 (Table 1) was characterized by a much higher heterogeneity of slow- and fast-migrating bands, whereas the LPS of strain P 1.12 migrated as two major bands. The LPS of strain P 2.24 had a distinct pattern and was categorized as type C (Table 1).
Four isolates (P1.37, P1.42, P1.47, and GC7.4) with LPS patterns similar to that of 3841 (Fig. 3) were more closely related to strains R. leguminosarum bv. viciae USDA 2370 and R. leguminosarum bv. trifolii ATCC 14480 than to strain 3841, while isolate GD 4, with the highest similarity to strain 3841, had a different LPS profile.
Fatty acid analysis
Strong acidic hydrolysis of LPS preparations was used to release both ester- and amide-bound fatty acids. The analysis revealed the presence of the following hydroxy fatty acids in all strains isolated from pea nodules and R. leguminosarum bv. viciae 3841: 3-hydroxymyristic (C14:0 3-OH), 3-hydroxypentadecanoic (C15:0 3-OH) 3-hydroxypalmitic (C16:0 3-OH), 3-hydroxystearic (C18:0 3-OH), and (ω-1) very long chain 27-hydroxyoctacosanoic (C28:0 27-OH) acids, which are specific to R. leguminosarum (Vedam et al. 2003; Brown et al. 2011) and R. etli lipid A (Que et al. 2000) (Table 2). In addition, 29-hydroxytriacontanoic acid (C30:0 29-OH) was detected. The presence of this fatty acid in lipid A of strain 3841 was also reported by Vedam et al. (2003). The fatty acid ratio varied among the strains; different ratios of primary acyl chains (ester- and amide-linked to the lipid A backbone) and secondary fatty acids (substituting hydroxyl groups of the primary acyl chains) were found. Based on fatty acid composition, two discrete profile groups could be distinguished (Table 2).
A dendrogram based on the mean fatty acid content (% of total fatty acids) confirmed the grouping of fatty acid profiles into two major clusters (Fig. 4; Table 2). In the first cluster, designated as 1 and comprising strains P 1.47, GC 5.8, GD 4, and, GC 1.3, the ratio of fatty acids was similar to that of R. leguminosarum 3841; in cluster 2, comprised by the remaining 11 strains, the content of C15:0 3-OH and C16:0 3-OH and ω-1 fatty acids was variable (Supplemental Fig. 4).
In addition, variable amounts of nonhydroxy fatty acids, such as hexadecanoic (C16:0), octadecanoic (C18:0), and octadecenoic (C18:1) fatty acids, were detected in all strains. Low levels of palmitic and stearic acids have been also found in lipid A preparations of R. leguminosarum and R. etli (Muszyński et al. 2011).
EPS production
All isolates except strain GC 5.5 produced large amounts of EPS, ranging from 216.7 ± 15.1 µg Glc mg−1 protein (isolate GC 1.6) to 453 ± 40.2 µg Glc mg−1 protein (isolate P 1.42), with an average of 280 µg Glc mg−1 protein. Isolate GC 5.5, which exhibited a moderately mucoid colony morphology, produced 2.5 times less EPS (113.7 ± 16.3 µg Glc mg−1 protein) than average (Table 1).
Sensitivity to salt, detergents, pH, and elevated temperature
Features such as sensitivity to NaCl and detergents are usually analyzed to determine whether some environmental conditions might impact the natural selection of strains nodulating legume plants (Zahran 1999; Bolanos et al. 2003).
The analysis of NaCl tolerance revealed moderate resistance to salinity (0.01–1.0% NaCl) of most strains. Four strains, GD 29, GB 53, GC 1.3, and GC 1.6, with group A LPS pattern, were more tolerant and grew at 3.0% NaCl (Table 1).
Half of the evaluated pea microsymbionts, including strain 3841, grew over a relatively wide range of pH values (pH 4.5–7.5), with an optimum at pH 7.2. The remaining isolates grew well at higher pH values, i.e., pH 4.5–8.0 (Table 1). Strain 5.5 that produced the lowest amount of EPS grew at a narrow pH range, pH 7.2–8.0.
All the isolates grew at temperatures from 20 to 28 °C, with an optimum at 28 °C. Although all strains survived at 37 and 42 °C, no strain grew at these two temperatures (data not shown).
Most strains isolated from pea nodules were more sensitive to SDS than to DOC. The concentration of SDS that inhibited the growth of the isolates varied from 0.015 to 0.12% (w/v), whereas the inhibitory concentrations of DOC were two times higher, i.e., 0.015–0.25% (Table 1). Strains P 2.24, P 1.12, and 3841 were able to grow only at the lowest tested concentration of SDS and DOC (0.06%).
A correlation between the ability to grow in the presence of higher salt concentrations and the LPS type was apparent in strains with type A LPS. Strains with type A LPS could all grow at similar concentrations of DOC and SDS; detergent sensitivity of strains with type B LPS was more variable. On the other hand, high sensitivity to detergents observed for strains 3841, P 2.24, and GC 5.5 (which all produce lower amounts of EPS) was in accordance with earlier data that indicated that EPS biosynthesis influences strain sensitivity to surface-active detergents (Janczarek et al. 2010). Similarly, according to Vanderlinde et al. (2010), the differences in strain sensitivity to anionic detergents can be linked to the composition of fatty acyl chains in LPS.
A dendrogram showing phenetic relationships between the studied strains, based on their sensitivity to salt, SDS, DOC, and pH, is presented in Fig. 5. At least three main strain groups can be distinguished. Group a includes seven isolates, and groups b and c comprise five and three strains, respectively (Fig. 5). Isolate GC 5.5 was unrelated to other strains, probably because of weak pH tolerance. Interestingly, five out of seven strains from phenetic group a also belonged to AFLP I clade, whereas the isolates from groups b and c were dispersed between AFLP groups II and III or remained ungrouped after AFLP profiling (Figs. 1, 5).
Euclidean distance-based dendrogram presenting phenetic relationships (Table 1) between pea nodule isolates
Discussion
A previous investigation of a local population of pea microsymbionts based on PCR–RFLP analysis of 16–23S rRNA ITS and nodA–F regions, plasmid patterns, and metabolic capabilities showed that individual plants were infected by numerous and diverse strains (Wielbo et al. 2011). The isolates from pea nodules were most closely related to R. leguminosarum bv. viciae 3841 and Rhizobium fabae CCBAU33202 (Wielbo et al. 2011), whereas P. sativum isolate GB 30 was most closely related to R. leguminosarum bv. trifolii WSM1689 (Mazur et al. 2015). In this paper, the strains nodulating P. sativum showed greatest similarity to R. leguminosarum biovars viciae and trifolii. Genotypic characteristics of the strains were compared with their electrophoretic and fatty acid LPS profiles. The effect of environmental stress conditions, such as salt, detergents, pH, and elevated temperature, on strain growth was quantitatively analyzed and compared with strain classifications based on the genetic and LPS profiles.
The diversity of LPS is generally associated with the variation in the O-antigen gene cluster. It has been well documented that horizontally acquired genes play a key role in defining LPS structure (Lerouge and Vanderleyden 2001; Muszyński et al. 2011). The differences in the electrophoretic patterns of LPS from strains analyzed herein indicated that the strains occupying the nodules of the pea cultivar significantly differ from one another. No similarity was observed between the dendrogram constructed based on the LPS electrophoretic pattern and the phylogenetic tree based on AFLP analysis. This finding is in agreement with the study of Santamaria et al. (1997, 1998), who demonstrated that LPS profiling ensures good discrimination of strains with a ladder-like pattern of LPS, e.g., Bradyrhizobium strains, but it is less useful for Rhizobium species since their LPS migrates as less distinctive bands.
The profiles of LPS-derived hydroxy fatty acids were reported to be a valuable chemotaxonomic marker for the systematics of the family Rhizobiaceae and distinguishing the genus (Yokota and Sakane 1991; Jarvis et al. 1996; Tighe et al. 2000; Zahran et al. 2003; Vanderlinde et al. 2010). The results of the current analysis of LPS-derived fatty acids suggested that the isolates are highly similar to R. leguminosarum and R. etli. The cladogram constructed based on the weight percentage of total fatty acids (UPGMA) of the isolates did not reflect the similarities between their LPS electrophoretic profiles.
It was reported that the differences in LPS fatty acid composition between the biovars viciae, phaseoli, and trifolii of R. leguminosarum were small (Jarvis et al. 1996; Dunfield et al. 1999). Although the variation in fatty acid profiles within a particular biovar of R. leguminosarum is not pronounced, it might nonetheless be used to establish the identity of pea nodule occupants (Dunfield et al. 1999). Our results revealed an agreement between fatty acid composition and phylogenetic classification at the genus level. A previous polyphasic taxonomic study of rhizobia from wild legumes (20 strains) demonstrated both an agreement and lack thereof between data clustered based on phenotypic analysis and total cellular fatty acid profiles (Zahran et al. 2003).
Despite close genetic similarity of strains, their LPS and fatty acid profiles are different. Strain classification based on these differences does not correlate with the genetic relationship, although it allows discrimination between the strains. The results are in agreement with the study of Dunfield et al. (1999). Our observation that there were differences in the proportion of 3-hydroxy to ω-1 hydroxy fatty acids suggests that additional studies are required to determine whether this ratio is dependent on the intrinsic heterogeneity of lipid A, is influenced by external conditions, or a combination of the two possibilities.
Based on a previous study (Wielbo et al. 2011), we can assume that strains producing higher amounts of EPS (GB 42, GD 31, and P 1.42) are also more efficient symbionts. The differences in LPS and fatty acid pattern and EPS production among pea isolates could be reflected in their sensitivity to environmental factors, e.g., salinity, pH range, or temperature. Besides these compounds, the tolerance of strains to environmental conditions, e.g., to acidic pH, is regulated by the adaptive response (Graham et al. 1994). The physiological properties of the strains were slightly discriminative (salt tolerance and growth pH range) or not discriminative (temperature growth range) in diversification of the studied isolates.
The relationship between phenotypic and genetic correlations in symbiotic bacteria is not entirely clear. For example, no apparent relationship between the genotypic and phenotypic diversity of rhizobial strains was noted by Xavier et al. (1998). Similar results were reported by Elboutahiri et al. (2010) for Sinorhizobium strains nodulating Medicago sativa. In contrast, a correlation between the phenotypic and genotypic characteristics of rhizobial strains isolated in situ from Accacia senegal nodules was found (Fall et al. 2008; Bakhoum et al. 2014).
Together with the results of our earlier study revealing a large diversity of pea microsymbionts (Wielbo et al. 2010), the current analysis suggests that the genome-level relationship between rhizobial strains in a localized area does not reflect their metabolic profiles but also their surface characteristics. We propose that the correlation seen between the largest AFLP group I and the physiological properties of ca. 30% of strains could be associated with isolate adaptation to the local soil conditions. Genomic relationships are indispensable for microsymbiont classification in light of high heterogeneity of surface compounds, LPS and EPS, observed in rhizobia infecting the same host plant. The LPS and fatty acid patterns, although not directly related to the genetic relationship, can nonetheless be used for strain differentiation.
Conclusion
The aim of this work was to evaluate whether LPS profiles and LPS-derived fatty acid patterns of rhizobia nodulating P. sativum are related to strain phylogeny. The phylogenetic tree constructed on the basis of the concatenated genes—16S rRNA, recA, and atpD—revealed that all strains belong to R. leguminosarum biovars viciae and trifolii. The pea isolates were divided into two major groups by AFLP; however, no evident correlation with LPS profiles and LPS-derived fatty acid composition was found. A relationship between LPS profiling and phenotypic characteristics, including sensitivity to anionic detergents and salinity, was not apparent.
References
Apicella MA (2008) Isolation and characterization of lipopolysaccharides. Methods Mol Biol 431:3–13
Bailly X, Olivieri I, Brunel B, Cleyet-Marel JC, Bena G (2007) Horizontal gene transfer and homologous recombination drive the evolution of the nitrogen-fixing symbionts of Medicago species. J Bacteriol 189:5223–5236
Bakhoum N, Le Roux C, Diouf D, Kane A, Ndoye F, Fall D, Duponnois R, Noba K, Sylla SN, Galiana A (2014) Distribution and diversity of rhizobial populations associated with Acacia senegal (L.) Willd. provenances in senegalese arid and semiarid regions. Open J For 4:136–143
Bolanos L, El-Hamdaoui A, Bonilla L (2003) Recovery of development and functionality of nodules and plant growth in salt-stressed Pisum sativum-Rhizobium leguminosarum symbiosis by boron and calcium. J Plant Physiol 160:1493–1497
Brown DB, Huang YC, Kannenberg EL, Sherrier DJ, Carlson RW (2011) An acpXL mutant of Rhizobium leguminosarum bv. phaseoli lacks 27-hydroxyoctacosanoic acid in its lipid A and is developmentally delayed during symbiotic infection of the determinate nodulating host plant Phaseolus vulgaris. J Bacteriol 193:4766–4778
Choma A, Komaniecka I (2011) Straight and branched (ω-1)-hydroxylated very long chain fatty acids are components of Bradyrhizobium lipid A. Acta Biochim Pol 58:51–57
D’Heaze W, Leoff C, Freshour G, Noel KD, Carlson RW (2007) Rhizobium etli CE3 bacteroid lipopolysaccharides are structurally similar but not identical to those produced by cultured CE3 bacteria. J Biol Chem 282:17101–17113
Dresler-Nurmi A, Fewer DP, Räsänen LA, Lindström K (2009) The diversity and evolution of Rhizobia. Microbiol Monogr 8:3–41
Dunfield KE, Xavier LJC, Germida JJ (1999) Identification of Rhizobium leguminosarum and Rhizobium sp. (Cicer) strains using a custom fatty acid methyl ester (FAME) profile library. J Appl Microbiol 86:78–86
Elboutahiri N, Thami-Alami I, Udupa SM (2010) Phenotypic and genetic diversity in Sinorhizobium meliloti and S. medicae from drought and salt affected regions of Morocco. BMC Microbiol 10:15. doi:10.1186/1471-2180-10-15
Fall D, Diouf D, Ourarhi M, Faye A, Abdelmounen H, Neyra M, Sylla SN, Missbah El Idrissi M (2008) Phenotypic and genotypic characteristics of Acacia senegal (L.) Willd. root-nodulating bacteria isolated from soils in the dryland part of Senegal. Lett Appl Microbiol 47:85–97
Felsenstein J (1985) Phylogenies and the comparative method. Am Nat 125:1–15
Gaunt MW, Turner SL, Rigottier-Gois L, Lloyd-Macgilp SA, Young JPW (2001) Phylogenies of atpD and recA support the small subunit rRNA-based classification of rhizobia. Int J Syst Evol Microbiol 51:2037–2048
Graham PH, Draeger KJ, Ferrey ML, Conroy MJ, Hammer BE, Martinez E, Aarons SR, Quinto C (1994) Acid pH tolerance in strains of Rhizobium and Bradyrhizobium, and initial studies on the basis for acid tolerance of Rhizobium tropici Umr 1899. Can J Microbiol 40:198–207
Hotter GS, Scott B (1991) Exopolysaccharide mutants of Rhizobium loti are fully effective on a determinate nodulating host but are ineffective on an indeterminate nodulating host. J Bacteriol 173:851–859
Janczarek M (2011) Environmental signals and regulatory pathways that influence exopolysaccharide production in rhizobia. Int J Mol Sci 12:7898–7933
Janczarek M, Kutkowska J, Piersiak T, Skorupska A (2010) Rhizobium leguminosarum bv. trifolii rosR is required for interaction with clover, biofilm formation and adaptation to the environment. BMC Microbiol 10:1–23
Jarvis BDW, Sivakumaran S, Tighe SW, Gillis M (1996) Identification of Agrobacterium and Rhizobium species based on cellular fatty acid composition. Plant Soil 184:143–158
Kawaharada Y, Kelly S, Wibroe Nielsen M, Hjuler CT, Gysel K, Muszyński A, Carlson RW, Thygesen MB, Sandal N, Asmussen MH, Vinther M, Andersen SU, Krusell L, Thirup S, Jensen KJ, Ronson CW, Blaise M, Radutoiu S, Stougaard J (2015) Receptor-mediated exopolysaccharide perception controls bacterial infection. Nature 523:308–312
Kimura M (1980) A simple method for estimating evolutionary rates of base substitutions through comparative studies of nucleotide sequences. J Mol Evol 16:111–120
Kumar N, Lad G, Giuntini E, Kaye ME, Udomwong P, Shamsani NJ, Young JPW, Bailly X (2015) Bacterial genospecies that are not ecologically coherent: population genomics of Rhizobium leguminosarum. Open Biol. 5:140133. doi:10.1098/rsob.140133
Laus MC, Logman TJ, Lamers GE, van Brussel AA, Carlson RW, Kijne JW (2006) A novel polar surface polysaccharide from Rhizobium leguminosarum binds host plant lectin. Mol Microbiol 59:1704–1713
Lerouge I, Vanderleyden J (2001) O-antigen structural variation: mechanisms and possible roles in animal/plant-microbe interactions. FEMS Microbiol Rev 26:17–47
Marchesi JR, Sato T, Weightman AJ, Martin TA, Fry JC, Hiom SJ, Wade WG (1998) Design and evaluation of useful bacterium-specific PCR primers that amplify genes coding for bacterial 16S rRNA. Appl Environ Microbiol 64:795–799
Marek-Kozaczuk M, Leszcz A, Wielbo J, Wdowiak-Wróbel S, Skorupska A (2013) Rhizobium pisi sv. trifolii K3.22 harboring nod genes of the Rhizobium leguminosarum sv. trifolii cluster. Syst Appl Microbiol 36:252–258
Mazur A, De Meyer SE, Tian R, Wielbo J, Zebracki K, Seshadri R, Reddy T, Markowitz V, Ivanova NN, Pati A, Woyke T, Kyrpides NC, Reeve W (2015) High-quality permanent draft genome sequence of Rhizobium leguminosarum bv. viciae strain GB30; an effective microsymbiont of Pisum sativum growing in Poland. Stand Genomic Sci 10:36
Muszyński A, Laus MC, Kijne JW, Carlson RW (2011) Structures of the lipopolysaccharides from Rhizobium leguminosarum RBL5523 and its UDP-glucose dehydrogenase mutant (exo5). Glycobiology 21:55–68
Noel KD, Vandenbosch KA, Kulpaca B (1986) Mutations in Rhizobium phaseoli that lead to arrested development of infection threads. J Bacteriol 168:1392–1401
Priefer UB (1989) Genes involved in lipopolysaccharide production and symbiosis are clustered on the chromosome of Rhizobium leguminosarum biovar viciae VF39. J Bacteriol 171:6161–6168
Que NLS, Lin S, Cotter RJ, Raetz CRH (2000) Purification and mass spectrometry of six lipid A species from the bacterial endosymbiont Rhizobium etli. Demonstration of a conserved distal unit and a variable proximal portion. J Biol Chem 275:28006–28016
Ramírez-Bahena MH, García-Fraile P, Peix A, Valverde A, Rivas R, Igual JM, Mateos PF, Martínez-Molina E, Velázquez E (2008) Revision of the taxonomic status of the species Rhizobium leguminosarum (Frank 1879) Frank 1889AL, Rhizobium phaseoli Dangeard 1926AL and Rhizobium trifolii Dangeard 1926AL. R. trifolii is a later synonym of R. leguminosarum. Reclassification of the strain R. leguminosarum DSM 30132 (5NCIMB 11478) as Rhizobium pisi sp. nov. Int J Syst Evol Microbiol 58:2484–2490
Santamaria M, Corzo J, León-Barrios M, Gutierrez-Navarro AM (1997) Characterization and differentiation of indigenous rhizobia isolated from Canarian shrub legumes of agricultural and ecological interest. Plant Soil 190:143–152
Santamaria M, Gutierrez-Navarro AM, Corzo J (1998) Lipopolysaccharide profiles from nodules as markers of Bradyrhizobium strains nodulating wild legumes. Appl Environ Microbiol 64:902–906
Skorupska A, Janczarek M, Marczak M, Mazur A, Król J (2006) Rhizobial exopolysaccharides: genetic control and symbiotic functions. Microb Cell Fact 5:7
Tamura K, Peterson D, Peterson N, Stecher G, Nei M, Kumar S (2011) MEGA5: molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods. Mol Biol Evol 28:2731–2739
Tian CF, Young JP, Wang ET, Tamimi SM, Chen WX (2010) Population mixing of Rhizobium leguminosarum bv. viciae nodulating Vicia faba: the role of recombination and lateral gene transfer. FEMS Microbiol Ecol 73:563–576
Tighe SW, de Lajudie P, Dipietro K, Lindstrom K, Nick G, Jarvis BDW (2000) Analysis of cellular fatty acids and phenotypic relationship of Agrobacterium, Bradyrhizobium, Mesorhizobium, Rhizobium and Sinorhizobium species using the Sherlock Microbial Identification System. Int J Syst Evol Microbiol 50:787–801
Tsai CM, Frasch CE (1982) A sensitive silver stain for detecting lipopolysaccharides in polyacrylamide gels. Anal Biochem 119:115–119
Vanderlinde EM, Harrison JJ, Muszyński A, Carlson RW, Turner RJ, Yostet CK (2010) Identification of a novel ABC transporter required for desiccation tolerance, and biofilm formation in Rhizobium leguminosarum bv. viciae 3841. FEMS Microbiol Ecol 71:327–340
Vedam V, Kannenberg EL, Haynes JG, Sherrier DJ, Datta A, Carlson RW (2003) A Rhizobium leguminosarum AcpXL mutant produces lipopolysaccharide lacking 27- hydroxyoctacosanoic acid. J Bacteriol 185:1841–1850
Vincent JM (1970) A manual for the practical study of root-nodule bacteria. Blackwell Scientific Publications Ltd, Oxford
Wdowiak-Wróbel S, Małek W (2005) Genomic diversity of Astragalus cicer microsymbionts revealed by AFLP fingerprinting. J Gen Appl Microbiol 51:369–378
Wielbo J, Kuske J, Marek-Kozaczuk M, Skorupska A (2010) The competition between Rhizobium leguminosarum bv. viciae strains progresses until late stages of symbiosis. Plant Soil 337:125–135
Wielbo J, Marek-Kozaczuk M, Mazur A, Kubik-Komar A, Skorupska A (2011) The structure and metabolic diversity of population of pea microsymbionts isolated from root nodules. BMRJ 1:55–69
Wolde-Meskel E, Terefework Z, Lindström K, Frostegard A (2004) Metabolic and genomic diversity of Rhizobia isolated from field standing native and exotic woody legumes in Southern Ethiopia. Syst Appl Microbiol 27:603–611
Xavier GR, Martins LMV, Neves MCP, Rumjanek NG (1998) Edaphic factors as determinants for the distribution of intrinsic antibiotic resistance in a cowpea rhizobia population. Biol Fertil Soils 27:386–392
Yasar S (2005) Spectrophotometric determination of hexose and pentose amounts by artificial neural network calibration and its using in wood analysis. Acta Chim Slov 52:435–439
Yokota A, Sakane T (1991) Taxonomic significance of fatty acid compositions in whole cells and lipopolysaccharides in Rhizobiaceae. IFO Res Commun 15:57–75
Zahran HH (1999) Rhizobium–legume symbiosis and nitrogen fixation under severe conditions and in arid climate. Microbiol Mol Biol Rev 63:968–989
Zahran HH, Abdel-Fattah M, Ahmad MS, Zaky AY (2003) Polyphasic taxonomy of symbiotic rhizobia from wild leguminous plants growing in Egypt. Folia Microbiol (Praha) 48:510–520
Zgadzaj R, James EK, Kelly S, Kawaharada Y, de Jonge N, Jensen DB, Madsen LH, Radutoiu S (2015) A legume genetic framework controls infection of nodules by symbiotic and endophytic bacteria. PLoS Genet 11(6):e1005280. doi:10.1371/journal.pgen.1005280
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by Jorge Membrillo-Hernández.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Supp Fig. 1
Phylogenetic tree generated using the neighbor-joining method, constructed based on partial sequences of the 16S rRNA gene of pea nodule isolates and closely related Rhizobium species. The distances were calculated according to Kimura’s two-parameter correction. Bootstrap analysis was based on 1000 resamplings. Bar, nucleotide substitution per site (TIFF 12534 kb)
Supp Fig. 2
Neighbor-joining tree showing the phylogenetic relationship between the test strains and related species based on atpD gene sequences. Bootstrap values (expressed as percentages of 1000 replications) are given at the nodes. Bar, nucleotide substitution per site (TIFF 12446 kb)
Supp Fig. 3
Neighbor-joining tree showing the phylogenetic relationship between the test strains and related species based on recA gene sequences. Bootstrap values (expressed as percentages of 1000 replications) are given at the nodes. Bar, nucleotide substitution per site (TIFF 11957 kb)
Supp Fig. 4
Gas chromatograms of methyl ester TMS ethers of fatty acids derived from the LPS of pea isolate GC 5.8, representing a fatty acid profile similar to that of the LPS from strain 3841 (A); GC 1.6, representing the second profile, with different C16:0 3-OH and C18:0 3-OH to ω-1 hydroxy fatty acid ratios (B). *, palmitic acid (C16:0); **, TMS derivative of sugar constituents of LPS; ○, octadecenoic acid (C18:1); ●, C19:0 cyclo, derived from membrane phospholipids; ■, artifacts formed during hydrolysis of ω-1 hydroxy fatty acid (TIFF 17589 kb)
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
About this article
Cite this article
Kutkowska, J., Marek-Kozaczuk, M., Wielbo, J. et al. Electrophoretic profiles of lipopolysaccharides from Rhizobium strains nodulating Pisum sativum do not reflect phylogenetic relationships between these strains. Arch Microbiol 199, 1011–1021 (2017). https://doi.org/10.1007/s00203-017-1374-1
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00203-017-1374-1