Abstract
The competition potential of 14 Rhizobium leguminosarum bv. viciae isolates originating from nodules of Pisum sativum was estimated. Genotypic analyses of the isolates revealed a high level of chromosomal and plasmid content diversity. The isolates tagged with a plasmid-bearing constitutively expressed gusA gene were used to inoculate vetch (Vicia villosa) in competition experiments carried out under laboratory conditions. Soil extract containing autochthonous rhizobial population was used as competitor for gus-tagged strains, and the competition was studied by: (i) estimation of Gus+ root nodules on whole root systems, (ii) the pattern of individual nodule colonization by Gus+/Gus− rhizobia, and (iii) the number of Gus+/Gus− bacteria recovered from individual nodules. Several patterns of nodule colonization by Gus+/Gus− bacteria were found. Some nodules identified as Gus+ contained gus-tagged bacteria only in the young and saprophytic zones, while the symbiotic zone was occupied by unmarked soil rhizobia. In other Gus+ nodules, despite the visible colonization of the entire nodule by gus-marked bacteria, a high number of Gus− soil-derived rhizobia were recovered. The results suggest that rhizobial strains compete with each other also in the late stage of nodule development. Therefore, they may use different strategies to reach the late saprophytic zone of the nodule, which serves as an optimal environment for massive proliferation.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Bacteria of the genera Azorhizobium, Bradyrhizobium, Mesorhizobium, Rhizobium and Sinorhizobium, collectively known as rhizobia, enter into N2-fixing symbioses with leguminous plants (Sprent 2001; Willems 2006). Despite genetic and physiological dissimilarities between rhizobial genera, common stages of symbiosis were identified, such as the recognition of symbiotic partners by exchange of molecular signals (usually plant flavonoids and bacterial Nod factors), nodule formation on plant roots or shoots, and finally, conversion of dinitrogen to ammonium (reviewed by Perret et al. 2000; Jones et al. 2007; Masson-Boivin et al. 2009). The sequential stages of symbiosis are controlled by a set of bacterial nod-nol-noe genes responsible for nodulation and nif-fix genes for nitrogen fixation, and several plant host genes. The bacterial genes are clustered on the symbiotic plasmid (pSym) or the chromosome in a region called a symbiotic island. In some rhizobia the symbiotic genes are scattered throughout chromosomes and plasmids (Palacios and Newton 2005).
Diazotrophic symbioses are beneficial for plants since bacteria supply the plant host with reduced nitrogenous compounds derived from nitrogen fixation, the major source of nitrogen in legumes (Graham and Vance 2003; Franche et al. 2009). About half of all the biological nitrogen fixation is carried out by rhizobia that form nitrogen-fixing symbioses with compatible leguminous species (Graham and Vance 2003). Symbiosis is also beneficial to rhizobia because a new environment, i.e., the plant tissues inside root nodules, becomes accessible to rhizobia. Since this ecological niche is not available to the majority of soil microorganisms, it is highly probable that rhizobia invest a lot of resources in adapting to the specific plant tissue environment and the symbiotic interaction may play a considerable role in evolution of rhizobia (Martinez-Romero 2009).
Rhizobium-legume interaction is specific and in most cases defined species of legumes may enter into symbiosis with only one (or few) bacterial species (Perret et al. 2000; Willems 2006). The exception is Rhizobium sp. NGR234 that is extremely promiscuous and able to nodulate different host plants (over 112 hosts) (Pueppke and Broughton 1999). On the other hand, the local rhizobial soil population may be composed of numerous strains differing in genomic structure, physiological properties and the efficiency of nitrogen fixation. Rhizobia compete with each other for plant root colonization and domination in the nodules of a specific legume host (Rangin et al. 2008; Silva et al. 2007; Beyene et al. 2003; Duodu et al. 2007). Numerous studies concerning competitiveness of rhizobia have been conducted, and different traits affecting bacterial competitiveness were indicated, such as bacteriocin production (Robleto et al. 1998; Oresnik et al. 1999), vitamin production (Streit et al. 1996), response to plant-derived flavonoid induction (Maj et al. 2008), utilization of specific carbon and energy sources (Hynes and O’Connel 1990; Oresnik et al. 1998), as well as utilization of a subset of carbon and energy substrates (Wielbo et al. 2007). All these traits might be advantageous in acquiring the domination in the rhizosphere, which is considered as the main factor leading to successful colonization of a host plant.
Recent studies of symbiosis showed that interactions between the plant host and a rhizobium colonizing the plant roots are more complex. The presence of different bacterial strains inside a single infection thread (Stuurman et al. 2000; Gage 2004), as well as active competition between strains during growth of infection thread, has been evidenced (Duodu et al. 2009). Moreover, it has been shown that in indeterminate nodules forming different developmental zones (Vasse et al. 1990; Timmers et al. 2000), the plant host controls bacterial cell cycle and enforces the differentiation of rhizobia into nitrogen-fixing bacteroids that are not able to divide after release from nodules due to irreversible changes in the genome and outer membrane structures (Mergaert et al. 2006). Only a small fraction of rhizobial cells in the nodules remains in a viable, vegetative form that can be recovered from the nodules (Mergaert et al. 2006). However, during nodule senescence a saprophytic zone is formed in which rhizobia can multiply massively (Timmers et al. 2000; Wielbo et al. 2009). Thus, a possibility exists that active competition between rhizobial strains takes place throughout the entire course of the symbiosis, from recruitment of the microsymbionts from rhizosphere to nodule decay. The question of rhizobial competition, which takes place inside plant nodules, is considerably much less explored relative to the competition in the rhizosphere.
Here, we report that competition between Rhizobium leguminosarum bv. viciae isolates could be also evidenced inside mature indeterminate nodules formed on vetch. For purpose of this study, we used a model reflecting the soil environment in which a gus-tagged inoculant strain in a mixture with the soil extract containing autochthonous rhizobia was used to inoculate vetch grown under laboratory conditions. As a result of competition between rhizobia inside the nodules different patterns of nodule colonization were found and different proportions of Gus+/Gus− rhizobia were recovered from the nodules.
Materials and methods
Characterization of rhizobial strains
Fourteen R. leguminosarum bv. viciae strains were isolated from nodules of pea (Pisum sativum cv. Ramrod) cultivated in arable sandy loam soil in the region of Lublin, Poland. Nodules were surface-sterilized, crushed and content of the nodules was cultured in 79CA medium (Vincent 1970). Strains isolated from the nodules were purified by successive isolation of single colonies and single clones were used in further experiments.
For growth assays, the bacteria grown 72 h at 28°C on TY plates (Sambrook et al. 1989) were harvested and suspended in sterile water to the optical density of cultures OD550 of 0.1 (approximately 107 cells mL−1). 150 μL liquid 79CA or TY media in 96-well microplate was inoculated with 10 μL bacterial suspension and incubated 24 h at 28°C with shaking (800 rpm) (Microtec AK120, Infors AG), following which OD550 was measured. Each experiment was conducted in triplicate.
Analysis of plasmid content in the isolates was performed as described by Eckhardt (1978) and then compared with plasmid standard of R. leguminosarum bv. viciae strain 3841 (Young et al. 2006).
PCR assays of 16S-23S rDNA internal transcribed spacer (ITS) were carried out using genomic DNA isolated from R. leguminosarum bv. viciae strains as templates and primers FGPS1490-5′-TGCGGCTGGATCACCTCCTT-3′ and FGPL132′-5′-CCGGGTTTCCC CATTCGG-3′ (Laguerre et al. 1996). PCR products were digested with BsuRI (HaeIII) and TaqI restriction enzymes (FERMENTAS) and restriction fragments were separated by 3% agarose gel electrophoresis.
Tagging of rhizobia
To study nodule occupancy, 14 sampled rhizobial strains were tagged by a stable plasmid vector pJBA21Km carrying a constitutively expressed gusA gene (Wielbo and Skorupska 2001). The pJBA21Km plasmid was introduced into bacteria by electroporation as described earlier (Wielbo et al. 2007).
Preparation of soil extract
Sandy loam soil was suspended in sterile water (200 g soil in 400 mL water) and the extraction carried out at 28°C for 20 min with shaking (120 rpm). Next, three successive sedimentation cycles (15 min/cycle) were applied, which allowed removal of soil particles. The final supernatant contained ~3 × 104 mL−1 of soil rhizobia. To determine the number of nodulating vetch rhizobia in the soil extract, most probable number (MNP) technique was used (Brockwell 1982).
Plant test for bacterial competitiveness
Vetch seeds (Vicia villosa cv. Wista) were surface-sterilized, germinated and grown on nitrogen-free Fahraeus medium (Vincent 1970), supplemented with commercial fungicide Funaben T (Organika-Azot S.A., Jaworzno, Poland) at final concentration 1 μg mL−1. 5-day-old vetch seedlings growing on agar slants were inoculated with 200 μL mixture of soil extract and R. leguminosarum bv. viciae gus-tagged isolate (OD550 0.1, ~107 cells mL−1) in water (1:1 v/v). Vetch plants were grown in a greenhouse under natural light supplemented with artificial light (14 h day/10 h night, at 24/19°C). After 5 weeks, eight plants were harvested, the roots were stained for 18 h for β-glucuronidase (GUS) activity (Wilson et al. 1995; Wielbo et al. 2007) and the number of blue and white nodules was calculated. The competitive potential of the tested isolates was estimated by comparing the number of blue nodules colonized by the gusA-tagged strains with that of white nodules colonized by indigenous strains from the soil extract. Five randomly chosen Gus+ nodules were taken for microscopic observation from each treatment. For isolation of total bacteria, before staining, five additional nodules were randomly chosen from each treatment. To estimate the number of Gus+ and Gus− rhizobia colonizing the individual nodules, they were surface-sterilized, crushed in 1 mL of sterile water, and sequential dilutions of bacteria plated on 79CA medium supplemented with 5-bromo-4-chloro-3-indoxyl-β-D-glucuronide (X-GlcA) at final concentration of 50 μg mL−1. For microscopic observation of Gus+/Gus− colonization pattern, Gus+ nodules were longitudinally hand sectioned in halves with a razor blade, and microscopic observations were performed using Olympus BX-41 light microscope.
Two control groups in nodule colonization assays were used: (I) comprised plants inoculated solely with 200 μL of a water suspension of individual gus-tagged rhizobial isolates (OD550 of 0.1), and (II) comprised plants inoculated with 200 μL of a water suspension (OD550 of 0.1) of mixture (1:1 v/v) of two isogenic strains: gus-tagged rhizobial isolates and wild-type, Gus− strains.
Results
Characterization of R. leguminosarum bv. viciae isolates and their nodulation competitiveness
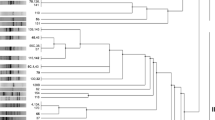
Fourteen R. leguminosarum bv. viciae nodule isolates representing an indigenous rhizosphere soil population originating from the same soil site were genetically and physiologically characterized. The plasmid profiles of the isolates showed great variability and each isolate contained from 2 to 4 plasmids (Fig. 1). RFLP analysis of PCR-amplified 16S-23S rDNA ITS region with BsuRI and TaqI restriction enzymes allowed identification of three distinct PCR-RFLP types (Fig. 2). Differences in growth of individual isolates on TY and 79CA liquid media as assessed by OD550 estimation were observed, especially in the case of growth on 79CA medium in which the differences between the isolates were as high as two-fold (Table 1). Strains belonging to PCR-RFLP group B showed higher OD550 mean value than strains belonging to groups A and C growing both on 79CA and TY, however, these differences were not significant at p < 0.05 (Table 3).
Plasmid profiles of Rhizobium leguminosarum bv. viciae isolates obtained by Eckhardt’s method. Lane1—GB25; lane 2—GB47; lane 3—P11; lane 4—GB36; lane 5—P272; lane 6—P111; lane 7—P147; lane 8—P270; lane 9—P229; lane 10—GC16; lane 11—GC31; lane 12—GC613; lane 13—GC76; lane 14—GD22; lane M—plasmids of R. leguminosarum bv. viciae strain 3841 used as molecular weight standard
PCR-RFLP analysis of 16S-23S rRNA ITS region of R. leguminosarum bv. viciae. PCR amplification products were digested with BsuRI and TaqI. Lanes 1 and 5- molecular size standard 100–300 bp (O’Gene Ruler™ 100 bp DNA ladder, Fermentas); lane 2—ITS type B/BsuRI (strains GB36, GC16, GC613, P229, P272); lane 3—ITS type A/BsuRI (strains GB25, GB47, GC31, P11); lane 4—ITS type C/BsuRI (strains GC76, GD22, P111, P147, P270); lane 6—ITS type B/TaqI (strains GB36, GC16, GC613, P229, P272); lane 7—ITS type A/TaqI (strains GB25, GB47, GC31, P11); lane 8—ITS type C/TaqI (strains GC76, GD22, P111, P147, P270)
In the competition experiment, the results of vetch coinoculation with gusA-tagged isolates and the rhizobia from soil extract were variable and strain-dependent (Table 2). The number of Gus+ nodules in whole root systems after mixed inoculation varied over a broad range, from 44.9% (P147) to 100% (P111), showing differences in competitive ability of the studied isolates. When bacteria from Gus+ nodules were plated on 79CA medium supplemented with X-GlcA, in most cases both Gus+ and Gus− colonies were recovered, although in various Gus+/Gus− cell ratios. In the cases of GB25, P272, GC16, GC613, GD22 and P111 isolates, only Gus+ colonies were recovered (Table 2).
Nodule occupancy patterns of R. leguminosarum bv. viciae Gus+ isolates
Nodule colonization patterns of gusA-marked rhizobia inoculated together with the soil extract were studied under a light microscope in longitudinally sectioned nodules.
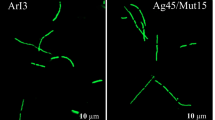
Considerable differences in nodule occupancy between individual isolates were observed (Fig. 3). GB25 and P272 isolates in mixed inoculation occupied the young, infection zone and the senescent, or saprophytic, zone of the nodules, but Gus+ bacteria were not detected in the symbiotic zone (Fig. 3a, c). The same isolates colonized the entire nodule space after a single strain inoculation (Fig. 3b, d) and after isogenic two-strains (GUS+ and Gus−) inoculation (details not shown). Similarly, GB36, GB47, P147 and P229 isolates were not able to colonize the entire nodule in a mixed plant inoculation (Fig. 3e, g, i, k), whereas they occupied all the nodule zones when they were used in a single inoculation (Fig. 3f, h, j, l) and in isogenic two-strains inoculation (details not shown). In contrast, GC31, GC76, P270, P11, GC16, GC613, P111 and GD22 isolates occupied the entire or almost entire nodules, both when competing with the soil indigenous rhizobia in mixed inoculations (Fig. 3m, n, p, q, s, u, w, x) and in single inoculation assays (Fig. 3o, r, t, y).
Patterns of vetch nodules occupancy by R. leguminosarum bv. viciae gusA-marked isolates in single (s) and mixed inoculation (m) with soil extract (photos a-y). (a) GB25m; (b) GB25s; (c) P272m; (d) P272s; (e) GB36m; (f) GB36s, (g) GB47m; (h) GB47s; (i) P147m; (j) P147s; (k) P229m; (l) P229s; (m) GC31m; (n) GC76m; (o) GC76s; (p) P11m; (q) P270m; (r) P270s; (s) P111m; (t) P111s; (u) GC16m; (w) GC613m; (x) GD22m; (y) GD22s. Reference nodules presenting the localization of typical indeterminate nodule zones: I—meristem, II—infection zone, int.—intermediate (II–III) zone, III—symbiotic zone, IV—senescence zone (photos 1–3). (1) unstained nodule (color of zone III cells results from leghaemoglobin); (2) toluidine blue-stained nodule (purple staining of cytoplasm and nuclei of plant cells in zones I, II and III); (3) Lugol’s solution-stained nodule (brown-colored starch grains accumulated preferentially in II–III interzone)
Considering the number of Gus+ nodules detected on root systems, the number of Gus+ colonies recovered from individual nodules (Tables 2 and 3) and the pattern of nodule colonization (Fig. 3), the R. leguminosarum bv. viciae isolates can be divided into four groups. Group I (GB25, P272) was characterized by a moderate number of Gus+ nodules that were entirely populated by Gus+ bacteria (Tables 2 and 3). These nodules had a regular shape. On the other hand, Gus+ rhizobia were not found in the symbiotic zone after coinoculation with the soil extract (Fig. 3a, c), even though they were found in all nodule zones after a single strain inoculation (Fig. 3b, d). One can assume that nodules elicited by GB25 or P272 were actually coinfected with soil extract–derived rhizobia that exclusively colonized the symbiotic zone but these rhizobia were not recovered from nodules because they were terminally differentiated as bacteroids. On the other hand, GB25 or P272 present in the saprophytic zones (Fig. 3a, c) were the only bacteria, which could be recovered from the nodules (Tables 2 and 3). In conclusion, the competition between GB25 or P272 gusA-tagged isolates and autochthonous strains seems to be the not only in the infection threads, but also in the late saprophytic stage of symbiosis.
Gus+ isolates of group II (GB36, GB47, P147 and P229) were being recovered in a variable number (from 16.7% to 87.5%) (Tables 2 and 3) from individual nodules elicited in mixed inoculation, and their pattern of nodule colonization was to some extent similar to group I. They colonized whole nodules only when no competitor strains were present (Fig. 3f, h, j, and l). In mixed inoculations, they showed various infection successes, but in each case visibly shared the nodule space with other rhizobia (Fig. 3e, g, i and k). It may be concluded that in contrast to group I, these isolates did not out-compete other strains in the saprophytic zone as a substantial number of Gus− rhizobial colonies was isolated from these nodules (Tables 2 and 3).
Isolates of group III elicited a relatively high number of Gus+ nodules (from 70.6% to 95.6%) on vetch roots (Table 2). Microscopic observations of nodule sections revealed that the infection success rate of these isolates was higher than of isolates of group II and in some nodules they appeared as the only inhabitants (Fig. 3m, n, p and q). The sections of nodules in single and mixed infections were similar (Fig. 3o, r). However, after isolation of rhizobia from the individual nodules, in some cases the “invisible” Gus− strains derived from the soil extract constituted nearly 50% of the entire population recovered from the nodule (Table 2). Mean percentage of Gus+ rhizobia isolated from the nodules was also relatively high for strains of this group and reached 81 ± 19% (Table 3).
Finally, isolates of group IV competing with the soil extract-derived strains colonized a relatively high number of nodules similarly to group III isolates (Tables 2 and 3), and occupied all zones of nodules (Fig. 3s, u, w and x). Nodule sections were similar in single and mixed infection (Fig. 3t, y). Except for a very similar colonization pattern when compared to group III isolates, no Gus− colonies were recovered from these nodules (Table 2). In conclusion, the results of competition were similar to group I isolates showing the out-competing of soil extract rhizobia by gusA-strains in the saprophytic zone.
Discussion
Rhizobial strains that inhabit the rhizosphere and enter symbioses with legumes vary in effectiveness, with ineffective bacteria being widespread (Simms and Taylor 2002). The competitive potential of rhizobia, besides great environmental and ecological importance, is economically significant since it is crucial for a successful application of commercial rhizobial inoculants (Streeter 1994; Triplett and Sadowsky 1992; Toro 1996). Numerous molecular techniques are available for the evaluation of competitive abilities of rhizobia, which may facilitate the assessment of usefulness of a prospective inoculant strain (Wilson et al. 1995; Sessitsch et al. 1998; Gage 2004). These techniques (with β-glucuronidase activity assay amongst them) are easily accessible and allow examining large numbers of host plants. Unfortunately, they are often based on the assumption that detection of a specific trait (e.g. β-glucuronidase activity) in a nodule is synonymous with the nodule being colonized exclusively by the one tagged strain, and that all or almost all the competition between rhizobial strains takes place in the rhizosphere.
More recent studies on rhizobia competitiveness revealed that competition between strains takes place not only in the rhizosphere, but also extends to the process of infection thread initiation and the growth of rhizobia in the infection threads (Stuurman et al. 2000; Gage 2004; Duodu et al. 2009). Greater competitive ability of a rhizobial strain was expressed as a high number of infections in root hairs, high rate of bacterial growth inside the infection threads and, subsequently, higher nodule occupancy (Gage 2004; Duodu et al. 2009).
In this study, we compared the competitive potential of R. leguminosarum bv. viciae gusA-tagged isolates in relation to autochthonous soil rhizobia at the level of whole root systems and individual nodules. In individual nodules, preferences for colonization of defined developmental zones were studied, and also the numbers of gusA-marked strain cells (inoculant) and of the unmarked strains (soil extract-derived rhizobia) in the population recovered from nodules. We wanted to investigate whether nodules identified as Gus+ were colonized exclusively by the introduced gus-marked strain or whether they were coinhabited by the marker strain and the unmarked autochthonous rhizobia representing the soil population.
Mixed occupancy of legume nodules was described years ago (McLoughlin and Dunican 1985; Ames-Gottfred and Christie 1989; McDermott and Graham 1990), but these experiments were conducted with two-strain mixtures, the nodules were assessed only as infected by mixed or pure cultures, and the patterns of nodule colonization by the individual strains have not been taken into consideration. The results of this study confirmed that indeed mixed colonization of particular nodules is a common phenomenon, also in the case of plant inoculation with more than two rhizobial strains. Moreover, mixed colonization can manifest itself in different patterns of developmental nodule zones occupancy (group I and II isolates) that may be essential for plant benefitting from the interaction.
Rhizobia, which enter into symbioses with plants may use mutualistic or parasitic strategy during nodule colonization (Denison and Kiers 2004a). The parasitic one offers low-cost and the most efficient multiplication of bacteria but is not beneficial for plant host; therefore legumes impose sanctions and reduce the reproductive success of rhizobia that do not fix enough N2 (Denison and Kiers 2004b). Different rates of rhizobial proliferation inside the saprophytic zone observed in this study may be a result of such microsymbiont “life strategies”, and may have important ecological implication for the rhizobia that are afterwards released into the soil.
In this work we assessed three different parameters concerning the competitive abilities of R. leguminosarum strains: (a) the number of Gus+ nodules in relation to all the nodules, (b) the number of gusA-marked cells in the rhizobial population recovered from the nodules, and (c) the nodule colonization pattern, and we investigated possible connections between them.
Since the pattern of nodule occupancy was not a measurable trait, no correlations with other traits were possible to draw. There were no significant differences in the mean percentage of Gus+ nodules when comparing groups I–IV of isolates (Table 3) suggesting that these values do not correlate with nodule colonization patterns. The mean percentage of Gus+ nodules was high as a result of the experimental conditions, since the amount of gusA-marked cells used for plant inoculation exceeded the number of rhizobia in the soil extract. We decided to employ such excess of the tagged strain so as to counter the potentially high competitiveness of autochthonous rhizobia (Wielbo et al. 2007; Maj et al. 2009). On the other hand, despite the initial numerical dominance of the marked strains, some differences in the number of Gus+ nodules were noticed between the individual isolates (Table 2) that might reflect their various competition abilities.
The percentage of Gus+ rhizobia recovered from the nodules varied considerably between isolates and between groups (Tables 2 and 3). Grouping of isolates on the basis of nodule occupancy pattern revealed greater differences in the percentage of Gus+ rhizobia than grouping the same isolates based on polymorphism of 16S-23S ITS region (Table 3). The correlation coefficients between total nodule numbers, the percentage of Gus+ and Gus− nodules as well as the number of Gus+ and Gus− rhizobia recovered from the nodules were very low (data not shown). Only the number of Gus+ nodules and total nodule count were positively and significantly correlated (correlation coefficient equal of 0.868), which stemmed from the prevalence of gusA-marked bacteria in the inocula as explained earlier. On the other hand, the number of Gus+ and Gus− nodules was not correlated with the number of Gus+ and Gus− rhizobia recovered from the nodules, suggesting that the competition during nodule colonization and the competition during bacteria multiplication in the saprophytic zone are independent of each other. Taken together, we propose that strain competition progresses until late stages of symbiosis. Strain success or failure during the initial plant tissue colonization, resulting in greater or lesser nodule occupancy, is not the sole determinant of bacterial yield, defined as the amount of viable rhizobia which are released into the soil from decaying nodules at the end of symbiosis.
In our previous work we have shown that the metabolic potential of rhizobia is essential for rhizobial competition (Wielbo et al. 2007). Here, we attempted to relate the metabolic capabilities of rhizobia assessed in growth experiments on rich media (TY and 79CA) to strain competiveness measured as a number of bacteria recovered from nodules and the pattern of nodule colonization. However, despite considerable differences in rhizobial growth on both media, no significant correlations with the number of Gus+ bacteria recovered from the nodules were found.
Nevertheless, the differences in colonization of specific nodule zones are intriguing. Some of the studied isolates colonized the entire nodule space and most probably actively participated in symbiotic nitrogen fixation (e.g. group IV isolates), but others (e.g. group I) infected only the young and saprophytic zones of nodules and probably did not take part in N fixation and were only stowaways in the nodules. Based on the well-known facts that various zones of indeterminate nodules contain different types of rhizobia, i.e., bacteria or bacteroids (Vasse et al. 1990; Timmers et al. 2000), and recent finding that differentiation of bacteria into bacteroids is enforced by the plant host (Mergaert et al. 2006), it is tempting to speculate that rhizobia may differ in their individual susceptibility to plant-derived factors, and the most susceptible strains from several occupying nodule space differentiate into bacteroids more easily, resulting in a great occupancy of the symbiotic zone and subsequently affecting the level of nitrogen fixation. The presented results demonstrate that some rhizobial strains could evade the host control during symbiosis. The use of standard histochemical methods (Wilson et al. 1995; Sessitsch et al. 1998) to classify rhizobia as “highly competitive” and the subsequent isolation of tagged bacteria from nodules lead to the same conclusion. However, in the case of rhizobia that only weakly colonize the symbiotic zone of nodules, their effect on plant growth is doubtful. From the bacterial point of view, this may be an advantageous strategy, since only those bacteria, which avoided the differentiation into bacteroids constitute the reproducible part of a population (Mergaert et al. 2006). In conclusion, the rhizobial soil population is composed of strains with different competition abilities. Some strains do not colonize the symbiotic zone in the presence of other competitors and thus are not well suited to become commercial inoculants even if they possess advantageous symbiotic properties when applied as the sole inoculant under laboratory conditions.
References
Ames-Gottfred NP, Christie BR (1989) Competition among strains of Rhizobium leguminosarum biovar trifolii and use of diallel analysis in assessing competition. Appl Environ Microbiol 55:1599–1604
Beyene D, Kassa S, Ampy F, Asseffa A, Gebremedhin T, van Berkum P (2003) Ethiopian soils harbor natural populations of rhizobia that form symbioses with common bean (Phaseolus vulgaris L.). Arch Microbiol 181:129–136
Brockwell J (1982) Plant-infection counts of rhizobia in soils. In: Vincent JM (ed) Nitrogen fixation in legumes. Academic, Sydney, pp 41–57
Denison RF, Kiers ET (2004a) Lifestyle alternatives for rhizobia: mutualism, parasitism, and forgoing symbiosis. FEMS Microbiol Lett 237:187–193
Denison RF, Kiers ET (2004b) Why are most rhizobia beneficial for their plant hosts, rather than parasitic? Microbes Infect 6:1235–1239
Duodu S, Carlsson G, Huss-Danell K, Svenning MM (2007) Large genotypic variation but small variation in N2 fixation among rhizobia nodulating red clover in soils of northern Scandinavia. J Appl Microbiol 102:1625–1635
Duodu S, Brophy C, Connolly J, Svenning MM (2009) Competitiveness of native Rhizobium leguminosarum bv. trifolii strain for nodule occupancy is manifested during infection. Plant Soil 318:117–126
Eckhardt T (1978) A rapid method for the identification of plasmid deoxyribonucleic acid in bacteria. Plasmid 1:584–588
Franche C, Lindström K, Elmerich C (2009) Nitrogen-fixing bacteria associated with leguminous and non-leguminous plants. Plant Soil 321:35–59
Gage DJ (2004) Infection and invasion of roots by symbiotic, nitrogen-fixing bacteria during nodulation of temperate legumes. Microbiol Mol Biol Rev 68:280–300
Graham PH, Vance CP (2003) Legumes: Importance and constraints to greater utilization. Plant Physiol 131:872–877
Hynes MF, O’Connel MP (1990) Host plant effect on competition among strains of Rhizobium leguminosarum. Can J Microbiol 36:864–869
Jones KM, Kobayashi H, Davies BW, Taga ME, Walker GC (2007) How symbionts invade plants: the Sinorhizobium-Medicago model. Nature 5:619–633
Laguerre G, Mavingui P, Allard MR, Charnay MP, Louvrier P, Mazurier SI, Rigottier-Gois L, Amarger N (1996) Typing of rhizobia by PCR DNA fingerprinting and PCR-restriction fragment length polymorphism analysis of chromosomal and symbiotic gene regions: application to Rhizobium leguminosarum and its different biovars. Appl Environ Microbiol 62:2029–2036
Maj D, Wielbo J, Marek-Kozaczuk M, Skorupska A (2008) Response to flavonoids as a factor influencing competitiveness and symbiotic activity of Rhizobium leguminosarum. Microbiol Res 165:50–60
Maj D, Wielbo J, Marek-Kozaczuk M, Martyniuk S, Skorupska A (2009) Pretreatment of clover seeds with Nod factors improves growth and nodulation of Trifolium pratense. J Chem Ecol 35:479–487
Martinez-Romero E (2009) Coevolution in Rhizobium-legume symbiosis? DNA Cell Biol 28:361–370
Masson-Boivin C, Giraud E, Perret X, Batut J (2009) Establishing nitrogen-fixing symbiosis with legumes: how many rhizobium recipes? Trends Microbiol 17:458–466
McDermott TR, Graham PH (1990) Competitive ability and efficiency in nodule formation of strains of Bradyrhizobium japonicum. Appl Environ Microbiol 56:3035–3039
McLoughlin TJ, Dunican LK (1985) Competition studies with Rhizobium trifolii in laboratory experiments. Plant Soil 88:139–143
Mergaert P, Uchiumi T, Alunni B, Evanno G, Cheron A, Catrice O, Mausset AE, Barloy-Hubler F, Galibert F, Kondorosi A, Kondorosi E (2006) Eukaryotic control on bacterial cell cycle and differentiation in the Rhizobium-legume symbiosis. Proc Natl Acad Sci USA 103:5230–5235
Oresnik IJ, Pacarynuk LA, O’Brien SHP, Yost C, Hynes MF (1998) Plasmid-encoded catabolic genes in Rhizobium leguminosarum bv. trifolii: evidence for a plant-inducible rhamnose locus involved in competition for nodulation. Mol Plant Microbe Interact 11:1175–1185
Oresnik IJ, Twelker S, Hynes MF (1999) Cloning and characterization of a Rhizobium leguminosarum gene encoding a bacteriocin with similarities to RTX toxins. Appl Environ Microbiol 65:2833–2840
Palacios R, Newton WE (2005) In: Palacios R, Newton WE (eds) Genomes and genomics of nitrogen-fixing organisms. Springer, Dordrecht
Perret X, Staehelin C, Broughton WJ (2000) Molecular basis of symbiotic promiscuity. Microbiol Mol Biol Rev 64:180–201
Pueppke SG, Broughton WJ (1999) Rhizobium sp. strain NGR234 and R. fredii USDA257 share exceptionally broad, nested host ranges. Mol Plant Microbe Interact 12:293–318
Rangin C, Brunel B, Cleyet-Marel JC, Perrineau MM, Bena G (2008) Effect of Medicago truncatula genetic diversity, rhizobial competition and strain effectiveness on the diversity of a natural Sinorhizobium species community. Appl Environ Microbiol 74:5653–5661
Robleto EA, Kmiecik K, Oplinger ES, Nienhuis J, Triplett EW (1998) Trifolitoxin production increases nodulation competitiveness of Rhizobium etli CE3 under agricultural conditions. Appl Environ Microbiol 64:2630–2633
Sambrook J, Fritsch EF, Maniatis T (1989) Molecular cloning: a laboratory manual, 2nd edn. Cold Spring Harbor Laboratory Press, Cold Spring Harbor
Sessitsch A, Hardarson G, de Vos WM, Wilson KJ (1998) Use of marker genes in competition studies of Rhizobium. Plant Soil 204:35–45
Silva C, Kan FL, Martinez-Romero E (2007) Population genetic structure of Sinorhizobium meliloti and S. medicae isolated from nodules Medicago spp. in Mexico. FEMS Microbiol Ecol 60:477–489
Simms EL, Taylor DL (2002) Partner choice in nitrogen-fixation mutualisms of legumes and rhizobia. Integr Comp Biol 42:369–380
Sprent JI (2001) Nodulation in legumes. Royal Botanic Gardens, Kew
Streeter JG (1994) Failure of inoculant rhizobia to overcome the dominance of indigenous strains for nodule formation. Can J Microbiol 40:513–522
Streit WR, Joseph CM, Philips DA (1996) Biotin and other water-soluble vitamins are key growth factors for alfalfa root colonization by Rhizobium meliloti 1021. Mol Plant Microbe Interact 9:330–338
Stuurman N, Pacios Bras C, Schlaman HRM, Wijfjes AHM, Bloemberg G, Spaink HP (2000) Use of green fluorescent protein color variants expressed on stable broad-host-range vectors to visualize rhizobia interacting with plants. Mol Plant Microbe Interact 13:1163–11699
Timmers AC, Soupène E, Auriac MC, de Billy F, Vasse J, Boistard P, Truchet G (2000) Saprophytic intracellular rhizobia in alfalfa nodules. Mol Plant Microbe Interact 13:1204–1213
Toro N (1996) Nodulation competitiveness in the Rhizobium-legume symbiosis. World J Microbiol Biotechnol 12:157–162
Triplett EW, Sadowsky MJ (1992) Genetics of competition for nodulation of legumes. Annu Rev Microbiol 46:399–428
Vasse J, de Billy F, Camut S, Truchet G (1990) Correlation between ultrastructural differentiation of bacteroids and nitrogen fixation in alfalfa nodules. J Bacteriol 172:4295–4306
Vincent JM (1970) A manual for the practical study of root nodule bacteria. International biological program handbook no.15. Blackwell Scientific Publications Ltd, Oxford
Wielbo J, Skorupska A (2001) Construction of improved vectors and cassettes containing gusA and antibiotic resistance genes for studies of transcriptional activity and bacterial localization. J Microbiol Methods 45:197–205
Wielbo J, Marek-Kozaczuk M, Kubik-Komar A, Skorupska A (2007) Increased metabolic potential of Rhizobium spp. is associated with bacterial competitiveness. Can J Microbiol 53:957–967
Wielbo J, Golus J, Marek-Kozaczuk M, Skorupska A (2009) Symbiosis stage-associated alterations in quorum sensing autoinducer molecules biosynthesis in Sinorhizobium meliloti. Plant Soil 329:399–410
Willems A (2006) The taxonomy of rhizobia: an overview. Plant Soil 287:3–14
Wilson KJ, Sessitsch A, Corbo JC, Giller KE, Akkermans AD, Jefferson RA (1995) beta-Glucuronidase (GUS) transposons for ecological and genetic studies of rhizobia and other gram-negative bacteria. Microbiology 141:1691–1705
Young JP, Crossman LC, Johnston AW, Thomson NR, Ghazoui ZF, Hull KH, Wexler M, Curson AR, Todd JD, Poole PS, Mauchline TH, East AK, Quail MA, Churcher C, Arrowsmith C, Cherevach I, Chillingworth T, Clarke K, Cronin A, Davis P, Fraser A, Hance Z, Hauser H, Jagels K, Moule S, Mungall K, Norbertczak H, Rabbinowitsch E, Sanders M, Simmonds M, Whitehead S, Parkhill J (2006) The genome of Rhizobium leguminosarum has recognizable core and accessory components. Genome Biol 7:R34
Acknowledgement
The research reported in this paper was supported by a grant NN304 026734 of Ministry of Sciences and Higher Education of Poland.
Open Access
This article is distributed under the terms of the Creative Commons Attribution Noncommercial License which permits any noncommercial use, distribution, and reproduction in any medium, provided the original author(s) and source are credited.
Author information
Authors and Affiliations
Corresponding author
Additional information
Responsible Editor: Petra Marschner.
Rights and permissions
Open Access This is an open access article distributed under the terms of the Creative Commons Attribution Noncommercial License (https://creativecommons.org/licenses/by-nc/2.0), which permits any noncommercial use, distribution, and reproduction in any medium, provided the original author(s) and source are credited.
About this article
Cite this article
Wielbo, J., Kuske, J., Marek-Kozaczuk, M. et al. The competition between Rhizobium leguminosarum bv. viciae strains progresses until late stages of symbiosis. Plant Soil 337, 125–135 (2010). https://doi.org/10.1007/s11104-010-0510-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11104-010-0510-3