Abstract
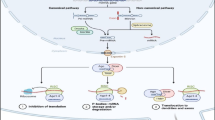
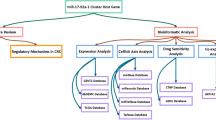
The reprogramming of glucose metabolism from oxidative to glycolytic metabolism, known as the Warburg effect, is an anomalous characteristic of cancer cell metabolism. Recent studies have revealed a subset of microRNAs (miRNAs) that play critical roles in regulating the reprogramming of glucose metabolism in cancer cells. These miRNAs regulate cellular glucose metabolism by directly targeting multiple metabolic genes, including those encoding key glycolytic enzymes. In the first part of this review, we summarized the recent knowledge of miRNA regulation in the reprogramming of glucose metabolism in cancer cells and discussed the potential utilization of the key miRNA regulators as metabolic targets for developing new antitumor agents. Then, we summarized recent advances in methods and techniques for studying miRNA regulation in cancer cell metabolism.


Similar content being viewed by others
Abbreviations
- MiRNA:
-
MicroRNA
- 3′ UTR:
-
3′ Untranslated region
- KO:
-
Knockout
- GLUT:
-
Glucose transporter
- SGLT:
-
Na+-coupled glucose transporter
- HK:
-
Hexokinase
- G-6-P:
-
Glucose-6-phosphate
- F-6-P:
-
Fructose-6-phosphate
- F-1,6-P:
-
Fructose-1,6-bisphosphate
- F-2,6-P:
-
Fructose-2,6-bisphosphate
- G-3-P:
-
Glyceraldehyde 3-phosphate
- R-5-P:
-
Ribulose 5-phosphate
- PKM1/M2:
-
Pyruvate kinase isozymes M1/M2
- HCC:
-
Hepatocellular carcinoma cell
- PTB1 or PTBP1:
-
Polypyrimidine tract-binding protein 1
- GC:
-
Gastric cancer
- LDH:
-
Lactate dehydrogenase
- NAD:
-
Nicotinamide adenine dinucleotide
- PDC:
-
Pyruvate dehydrogenase complex
- PDH:
-
Pyruvate dehydrogenase
- PDK:
-
Pyruvate dehydrogenase kinase
- EMT:
-
Epithelial–mesenchymal transition
- PFK:
-
Phosphofructokinase
- 1,3-BPG:
-
1,3-Bisphosphoglyceric acid
- DHAP:
-
Dihydroxyacetone phosphate
- PEP:
-
Phosphoenolpyruvate
- GT:
-
Glutamine transporter
- AldoA:
-
Aldolase A
- GAPDH:
-
Glyceraldehyde-3-phosphate dehydrogenase
- MCTs:
-
Monocarboxylate transporters
- MS:
-
Mass spectrometry
- NMR:
-
Nuclear magnetic resonance
- EC:
-
Esophageal cancer
- HNSCC:
-
Head and neck squamous cell carcinoma
- UHPLC:
-
Ultra-high-performance liquid chromatography
- TCA:
-
Tricarboxylic acid
- LC/MS:
-
Liquid chromatography/mass spectrometry
- CE/MS:
-
Capillary electrophoresis/mass spectrometry
- GC/MS:
-
Gas chromatography/mass spectrometry
- 2DG:
-
2-Deoxy-d-glucose
- 2-NBDG:
-
2-(N-(7-nitrobenz-2-oxa-1,3-diazol-4-yl) amino)-2-deoxyglucose
- 3-OPG:
-
3-O-propargyl-d-glucose
- NIR:
-
Near infrared fluorescence
- CTC:
-
Circulating tumor cells
- FRET:
-
Förster resonance energy transfer
- TPM:
-
Two-photon microscopy
- PET:
-
Positron emission tomography
- FLT:
-
Fluorothymidine
- SUVs:
-
Standardized uptake values
- MRS:
-
Magnetic resonance spectroscopy
- AML:
-
Acute myeloid leukemia
- 5-FU:
-
5-Fluorouracil
- α-KG:
-
α-Ketoglutarate
- lncRNAs:
-
Long noncoding RNAs
- OCR:
-
Oxygen consumption rate
- ECAR:
-
Extracellular acidification rate
References
Hanahan D, Weinberg RA (2011) Hallmarks of cancer: the next generation. Cell 144(5):646–674. doi:10.1016/j.cell.2011.02.013
Seyfried TN, Shelton LM (2010) Cancer as a metabolic disease. Nutr Metab (Lond) 7:7. doi:10.1186/1743-7075-7-7
Cairns RA, Harris IS, Mak TW (2011) Regulation of cancer cell metabolism. Nat Rev Cancer 11(2):85–95. doi:10.1038/nrc2981
Ahn CS, Metallo CM (2015) Mitochondria as biosynthetic factories for cancer proliferation. Cancer Metab 3(1):1. doi:10.1186/s40170-015-0128-2
Santos CR, Schulze A (2012) Lipid metabolism in cancer. FEBS J 279(15):2610–2623. doi:10.1111/j.1742-4658.2012.08644.x
Dang CV (2012) Links between metabolism and cancer. Genes Dev 26(9):877–890. doi:10.1101/gad.189365.112
Warburg O (1956) On the origin of cancer cells. Science 123(3191):309–314
Kroemer G, Pouyssegur J (2008) Tumor cell metabolism: cancer’s Achilles’ heel. Cancer Cell 13(6):472–482. doi:10.1016/j.ccr.2008.05.005
Vander Heiden MG, Cantley LC, Thompson CB (2009) Understanding the Warburg effect: the metabolic requirements of cell proliferation. Science 324(5930):1029–1033. doi:10.1126/science.1160809
Swietach P, Vaughan-Jones RD, Harris AL (2007) Regulation of tumor pH and the role of carbonic anhydrase 9. Cancer Metastasis Rev 26(2):299–310. doi:10.1007/s10555-007-9064-0
Fischer K, Hoffmann P, Voelkl S, Meidenbauer N, Ammer J, Edinger M, Gottfried E, Schwarz S, Rothe G, Hoves S, Renner K, Timischl B, Mackensen A, Kunz-Schughart L, Andreesen R, Krause SW, Kreutz M (2007) Inhibitory effect of tumor cell-derived lactic acid on human T cells. Blood 109(9):3812–3819. doi:10.1182/blood-2006-07-035972
He L, Hannon GJ (2004) MicroRNAs: small RNAs with a big role in gene regulation. Nat Rev Genet 5(7):522–531. doi:10.1038/nrg1379
Li Y, Kowdley KV (2012) MicroRNAs in common human diseases. Genomics Proteomics Bioinform 10(5):246–253. doi:10.1016/j.gpb.2012.07.005
O’Connell RM, Rao DS, Baltimore D (2012) microRNA regulation of inflammatory responses. Annu Rev Immunol 30:295–312. doi:10.1146/annurev-immunol-020711-075013
Hayes J, Peruzzi PP, Lawler S (2014) MicroRNAs in cancer: biomarkers, functions and therapy. Trends Mol Med 20(8):460–469. doi:10.1016/j.molmed.2014.06.005
Rottiers V, Naar AM (2012) MicroRNAs in metabolism and metabolic disorders. Nat Rev Mol Cell Biol 13(4):239–250. doi:10.1038/nrm3313
Lunt SY, Vander Heiden MG (2011) Aerobic glycolysis: meeting the metabolic requirements of cell proliferation. Annu Rev Cell Dev Biol 27:441–464. doi:10.1146/annurev-cellbio-092910-154237
Chen LQ, Cheung LS, Feng L, Tanner W, Frommer WB (2015) Transport of sugars. Annu Rev Biochem 84:865–894. doi:10.1146/annurev-biochem-060614-033904
Mueckler M, Thorens B (2013) The SLC2 (GLUT) family of membrane transporters. Mol Aspects Med 34(2–3):121–138. doi:10.1016/j.mam.2012.07.001
Singh PK, Brand RE, Mehla K (2012) MicroRNAs in pancreatic cancer metabolism. Nat Rev Gastroenterol Hepatol 9(6):334–344. doi:10.1038/nrgastro.2012.63
Fei X, Qi M, Wu B, Song Y, Wang Y, Li T (2012) MicroRNA-195-5p suppresses glucose uptake and proliferation of human bladder cancer T24 cells by regulating GLUT3 expression. FEBS Lett 586(4):392–397. doi:10.1016/j.febslet.2012.01.006
Lu H, Buchan RJ, Cook SA (2010) MicroRNA-223 regulates Glut4 expression and cardiomyocyte glucose metabolism. Cardiovasc Res 86(3):410–420. doi:10.1093/cvr/cvq010
Horie T, Ono K, Nishi H, Iwanaga Y, Nagao K, Kinoshita M, Kuwabara Y, Takanabe R, Hasegawa K, Kita T, Kimura T (2009) MicroRNA-133 regulates the expression of GLUT4 by targeting KLF15 and is involved in metabolic control in cardiac myocytes. Biochem Biophys Res Commun 389(2):315–320. doi:10.1016/j.bbrc.2009.08.136
Chen YH, Heneidi S, Lee JM, Layman LC, Stepp DW, Gamboa GM, Chen BS, Chazenbalk G, Azziz R (2013) miRNA-93 inhibits GLUT4 and is overexpressed in adipose tissue of polycystic ovary syndrome patients and women with insulin resistance. Diabetes 62(7):2278–2286. doi:10.2337/db12-0963
Singh PK, Mehla K, Hollingsworth MA, Johnson KR (2011) Regulation of aerobic glycolysis by microRNAs in cancer. Mol Cell Pharmacol 3(3):125–134
Mathupala SP, Ko YH, Pedersen PL (2006) Hexokinase II: cancer’s double-edged sword acting as both facilitator and gatekeeper of malignancy when bound to mitochondria. Oncogene 25(34):4777–4786. doi:10.1038/sj.onc.1209603
Cardenas ML, Cornish-Bowden A, Ureta T (1998) Evolution and regulatory role of the hexokinases. Biochim Biophys Acta 1401(3):242–264
Jiang S, Zhang LF, Zhang HW, Hu S, Lu MH, Liang S, Li B, Li Y, Li D, Wang ED, Liu MF (2012) A novel miR-155/miR-143 cascade controls glycolysis by regulating hexokinase 2 in breast cancer cells. EMBO J 31(8):1985–1998. doi:10.1038/emboj.2012.45
Jiang S, Zhang HW, Lu MH, He XH, Li Y, Gu H, Liu MF, Wang ED (2010) MicroRNA-155 functions as an OncomiR in breast cancer by targeting the suppressor of cytokine signaling 1 gene. Cancer Res 70(8):3119–3127. doi:10.1158/0008-5472.can-09-4250
Fang R, Xiao T, Fang Z, Sun Y, Li F, Gao Y, Feng Y, Li L, Wang Y, Liu X, Chen H, Liu XY, Ji H (2012) MicroRNA-143 (miR-143) regulates cancer glycolysis via targeting hexokinase 2 gene. J Biol Chem 287(27):23227–23235. doi:10.1074/jbc.M112.373084
Peschiaroli A, Giacobbe A, Formosa A, Markert EK, Bongiorno-Borbone L, Levine AJ, Candi E, D’Alessandro A, Zolla L, Finazzi Agro A, Melino G (2013) miR-143 regulates hexokinase 2 expression in cancer cells. Oncogene 32(6):797–802. doi:10.1038/onc.2012.100
Gregersen LH, Jacobsen A, Frankel LB, Wen J, Krogh A, Lund AH (2012) MicroRNA-143 down-regulates hexokinase 2 in colon cancer cells. BMC Cancer 12:232. doi:10.1186/1471-2407-12-232
Yoshino H, Enokida H, Itesako T, Kojima S, Kinoshita T, Tatarano S, Chiyomaru T, Nakagawa M, Seki N (2013) Tumor-suppressive microRNA-143/145 cluster targets hexokinase-2 in renal cell carcinoma. Cancer Sci 104(12):1567–1574. doi:10.1111/cas.12280
Chen X, Guo X, Zhang H, Xiang Y, Chen J, Yin Y, Cai X, Wang K, Wang G, Ba Y, Zhu L, Wang J, Yang R, Zhang Y, Ren Z, Zen K, Zhang J, Zhang CY (2009) Role of miR-143 targeting KRAS in colorectal tumorigenesis. Oncogene 28(10):1385–1392. doi:10.1038/onc.2008.474
Kent OA, Chivukula RR, Mullendore M, Wentzel EA, Feldmann G, Lee KH, Liu S, Leach SD, Maitra A, Mendell JT (2010) Repression of the miR-143/145 cluster by oncogenic Ras initiates a tumor-promoting feed-forward pathway. Genes Dev 24(24):2754–2759. doi:10.1101/gad.1950610
Osaki M, Takeshita F, Sugimoto Y, Kosaka N, Yamamoto Y, Yoshioka Y, Kobayashi E, Yamada T, Kawai A, Inoue T, Ito H, Oshimura M, Ochiya T (2011) MicroRNA-143 regulates human osteosarcoma metastasis by regulating matrix metalloprotease-13 expression. Mol Ther 19(6):1123–1130. doi:10.1038/mt.2011.53
Miao Y, Zhang LF, Guo R, Liang S, Zhang M, Shi S, Shang-Guan CF, Liu MF, Li B (2016) (18)F-FDG PET/CT for monitoring the response of breast cancer to miR-143-based therapeutics by targeting tumor glycolysis. Mol Ther Nucleic Acids 5(8):e357. doi:10.1038/mtna.2016.72
Guo W, Qiu Z, Wang Z, Wang Q, Tan N, Chen T, Chen Z, Huang S, Gu J, Li J, Yao M, Zhao Y, He X (2015) MiR-199a-5p is negatively associated with malignancies and regulates glycolysis and lactate production by targeting hexokinase 2 in liver cancer. Hepatology 62(4):1132–1144. doi:10.1002/hep.27929
Teng Y, Zhang Y, Qu K, Yang X, Fu J, Chen W, Li X (2015) MicroRNA-29B (mir-29b) regulates the Warburg effect in ovarian cancer by targeting AKT2 and AKT3. Oncotarget 6(38):40799–40814. doi:10.18632/oncotarget.5695
Imamura K, Tanaka T (1972) Multimolecular forms of pyruvate kinase from rat and other mammalian tissues. I. Electrophoretic studies. J Biochem 71(6):1043–1051
Mazurek S (2011) Pyruvate kinase type M2: a key regulator of the metabolic budget system in tumor cells. Int J Biochem Cell Biol 43(7):969–980. doi:10.1016/j.biocel.2010.02.005
Luo W, Semenza GL (2012) Emerging roles of PKM2 in cell metabolism and cancer progression. Trends Endocrinol Metab 23(11):560–566. doi:10.1016/j.tem.2012.06.010
Gupta V, Bamezai RN (2010) Human pyruvate kinase M2: a multifunctional protein. Protein Sci 19(11):2031–2044. doi:10.1002/pro.505
Yang W, Xia Y, Ji H, Zheng Y, Liang J, Huang W, Gao X, Aldape K, Lu Z (2011) Nuclear PKM2 regulates beta-catenin transactivation upon EGFR activation. Nature 480(7375):118–122. doi:10.1038/nature10598
Yang W, Xia Y, Hawke D, Li X, Liang J, Xing D, Aldape K, Hunter T, Alfred Yung WK, Lu Z (2012) PKM2 phosphorylates histone H3 and promotes gene transcription and tumorigenesis. Cell 150(4):685–696. doi:10.1016/j.cell.2012.07.018
Taniguchi K, Sugito N, Kumazaki M, Shinohara H, Yamada N, Nakagawa Y, Ito Y, Otsuki Y, Uno B, Uchiyama K, Akao Y (2015) MicroRNA-124 inhibits cancer cell growth through PTB1/PKM1/PKM2 feedback cascade in colorectal cancer. Cancer Lett 363(1):17–27. doi:10.1016/j.canlet.2015.03.026
Xu Q, Zhang M, Tu J, Pang L, Cai W, Liu X (2015) MicroRNA-122 affects cell aggressiveness and apoptosis by targeting PKM2 in human hepatocellular carcinoma. Oncol Rep 34(4):2054–2064. doi:10.3892/or.2015.4175
Sun Y, Zhao X, Zhou Y, Hu Y (2012) miR-124, miR-137 and miR-340 regulate colorectal cancer growth via inhibition of the Warburg effect. Oncol Rep 28(4):1346–1352. doi:10.3892/or.2012.1958
Wong TS, Liu XB, Chung-Wai Ho A, Po-Wing Yuen A, Wai-Man Ng R, Ignace Wei W (2008) Identification of pyruvate kinase type M2 as potential oncoprotein in squamous cell carcinoma of tongue through microRNA profiling. Int J Cancer 123(2):251–257. doi:10.1002/ijc.23583
Liu AM, Xu Z, Shek FH, Wong KF, Lee NP, Poon RT, Chen J, Luk JM (2014) miR-122 targets pyruvate kinase M2 and affects metabolism of hepatocellular carcinoma. PLoS One 9(1):e86872. doi:10.1371/journal.pone.0086872
Tang R, Yang C, Ma X, Wang Y, Luo D, Huang C, Xu Z, Liu P, Yang L (2016) MiR-let-7a inhibits cell proliferation, migration, and invasion by down-regulating PKM2 in gastric cancer. Oncotarget. doi:10.18632/oncotarget.6821
Luan W, Wang Y, Chen X, Shi Y, Wang J, Zhang J, Qian J, Li R, Tao T, Wei W, Hu Q, Liu N, You Y (2015) PKM2 promotes glucose metabolism and cell growth in gliomas through a mechanism involving a let-7a/c-Myc/hnRNPA1 feedback loop. Oncotarget 6(15):13006–13018. doi:10.18632/oncotarget.3514
Kefas B, Comeau L, Erdle N, Montgomery E, Amos S, Purow B (2010) Pyruvate kinase M2 is a target of the tumor-suppressive microRNA-326 and regulates the survival of glioma cells. Neuro Oncol 12(11):1102–1112. doi:10.1093/neuonc/noq080
David CJ, Chen M, Assanah M, Canoll P, Manley JL (2010) HnRNP proteins controlled by c-Myc deregulate pyruvate kinase mRNA splicing in cancer. Nature 463(7279):364–368. doi:10.1038/nature08697
Taniguchi K, Sakai M, Sugito N, Kumazaki M, Shinohara H, Yamada N, Nakayama T, Ueda H, Nakagawa Y, Ito Y, Futamura M, Uno B, Otsuki Y, Yoshida K, Uchiyama K, Akao Y (2016) PTBP1-associated microRNA-1 and -133b suppress the Warburg effect in colorectal tumors. Oncotarget 7(14):18940–18952. doi:10.18632/oncotarget.8005
Sugiyama T, Taniguchi K, Matsuhashi N, Tajirika T, Futamura M, Takai T, Akao Y, Yoshida K (2016) MiR-133b inhibits growth of human gastric cancer cells by silencing pyruvate kinase muscle-splicer polypyrimidine tract-binding protein 1. Cancer Sci 107(12):1767–1775. doi:10.1111/cas.13091
Takai T, Yoshikawa Y, Inamoto T, Minami K, Taniguchi K, Sugito N, Kuranaga Y, Shinohara H, Kumazaki M, Tsujino T, Takahara K, Ito Y, Akao Y, Azuma H (2017) A novel combination RNAi toward Warburg effect by replacement with miR-145 and silencing of PTBP1 induces apoptotic cell death in bladder cancer cells. Int J Mol Sci 18(1). doi:10.3390/ijms18010179
Brooks GA, Dubouchaud H, Brown M, Sicurello JP, Butz CE (1999) Role of mitochondrial lactate dehydrogenase and lactate oxidation in the intracellular lactate shuttle. Proc Natl Acad Sci USA 96(3):1129–1134
Miao P, Sheng S, Sun X, Liu J, Huang G (2013) Lactate dehydrogenase A in cancer: a promising target for diagnosis and therapy. IUBMB Life 65(11):904–910. doi:10.1002/iub.1216
Kaller M, Liffers ST, Oeljeklaus S, Kuhlmann K, Roh S, Hoffmann R, Warscheid B, Hermeking H (2011) Genome-wide characterization of miR-34a induced changes in protein and mRNA expression by a combined pulsed SILAC and microarray analysis. Mol Cell Proteomics 10(8):M111 010462. doi:10.1074/mcp.M111.010462
Wang J, Wang H, Liu A, Fang C, Hao J, Wang Z (2015) Lactate dehydrogenase A negatively regulated by miRNAs promotes aerobic glycolysis and is increased in colorectal cancer. Oncotarget 6(23):19456–19468. doi:10.18632/oncotarget.3318
Xiao X, Huang X, Ye F, Chen B, Song C, Wen J, Zhang Z, Zheng G, Tang H, Xie X (2016) The miR-34a-LDHA axis regulates glucose metabolism and tumor growth in breast cancer. Sci Rep 6:21735. doi:10.1038/srep21735
Li X, Zhao H, Zhou X, Song L (2015) Inhibition of lactate dehydrogenase A by microRNA-34a resensitizes colon cancer cells to 5-fluorouracil. Mol Med Rep 11(1):577–582. doi:10.3892/mmr.2014.2726
Li X, Lu P, Li B, Yang R, Chu Y, Zhang Z, Wan H, Niu C, Wang C, Luo K (2016) Sensitization of hepatocellular carcinoma cells to irradiation by miR34a through targeting lactate dehydrogenase A. Mol Med Rep 13(4):3661–3667. doi:10.3892/mmr.2016.4974
Isozaki Y, Hoshino I, Nohata N, Kinoshita T, Akutsu Y, Hanari N, Mori M, Yoneyama Y, Akanuma N, Takeshita N, Maruyama T, Seki N, Nishino N, Yoshida M, Matsubara H (2012) Identification of novel molecular targets regulated by tumor suppressive miR-375 induced by histone acetylation in esophageal squamous cell carcinoma. Int J Oncol 41(3):985–994. doi:10.3892/ijo.2012.1537
Isozaki Y, Hoshino I, Akutsu Y, Hanari N, Mori M, Nishimori T, Murakami K, Akanuma N, Takeshita N, Maruyama T, Toyozumi T, Takahashi M, Suito H, Matsubara H (2015) Usefulness of microRNA375 as a prognostic and therapeutic tool in esophageal squamous cell carcinoma. Int J Oncol 46(3):1059–1066. doi:10.3892/ijo.2014.2789
Zhang S, Hulver MW, McMillan RP, Cline MA, Gilbert ER (2014) The pivotal role of pyruvate dehydrogenase kinases in metabolic flexibility. Nutr Metab (Lond) 11(1):10. doi:10.1186/1743-7075-11-10
Roche TE, Hiromasa Y (2007) Pyruvate dehydrogenase kinase regulatory mechanisms and inhibition in treating diabetes, heart ischemia, and cancer. Cell Mol Life Sci 64(7–8):830–849. doi:10.1007/s00018-007-6380-z
Hitosugi T, Fan J, Chung TW, Lythgoe K, Wang X, Xie J, Ge Q, Gu TL, Polakiewicz RD, Roesel JL, Chen GZ, Boggon TJ, Lonial S, Fu H, Khuri FR, Kang S, Chen J (2011) Tyrosine phosphorylation of mitochondrial pyruvate dehydrogenase kinase 1 is important for cancer metabolism. Mol Cell 44(6):864–877. doi:10.1016/j.molcel.2011.10.015
Sutendra G, Michelakis ED (2013) Pyruvate dehydrogenase kinase as a novel therapeutic target in oncology. Front Oncol 3:38. doi:10.3389/fonc.2013.00038
Kim HR, Roe JS, Lee JE, Cho EJ, Youn HD (2013) p53 regulates glucose metabolism by miR-34a. Biochem Biophys Res Commun 437(2):225–231. doi:10.1016/j.bbrc.2013.06.043
Ma X, Li C, Sun L, Huang D, Li T, He X, Wu G, Yang Z, Zhong X, Song L, Gao P, Zhang H (2014) Lin28/let-7 axis regulates aerobic glycolysis and cancer progression via PDK1. Nat Commun 5:5212. doi:10.1038/ncomms6212
Mazar J, Qi F, Lee B, Marchica J, Govindarajan S, Shelley J, Li JL, Ray A, Perera RJ (2016) MicroRNA 211 functions as a metabolic switch in human melanoma cells. Mol Cell Biol 36(7):1090–1108. doi:10.1128/MCB.00762-15
Tang H, Lee M, Sharpe O, Salamone L, Noonan EJ, Hoang CD, Levine S, Robinson WH, Shrager JB (2012) Oxidative stress-responsive microRNA-320 regulates glycolysis in diverse biological systems. FASEB J 26(11):4710–4721. doi:10.1096/fj.11-197467
Calin GA, Cimmino A, Fabbri M, Ferracin M, Wojcik SE, Shimizu M, Taccioli C, Zanesi N, Garzon R, Aqeilan RI, Alder H, Volinia S, Rassenti L, Liu X, Liu CG, Kipps TJ, Negrini M, Croce CM (2008) MiR-15a and miR-16-1 cluster functions in human leukemia. Proc Natl Acad Sci USA 105(13):5166–5171. doi:10.1073/pnas.0800121105
Seidler NW (2013) GAPDH and intermediary metabolism. Adv Exp Med Biol 985:37–59. doi:10.1007/978-94-007-4716-6_2
Sikand K, Singh J, Ebron JS, Shukla GC (2012) Housekeeping gene selection advisory: glyceraldehyde-3-phosphate dehydrogenase (GAPDH) and beta-actin are targets of miR-644a. PLoS One 7(10):e47510. doi:10.1371/journal.pone.0047510
Pinweha P, Rattanapornsompong K, Charoensawan V, Jitrapakdee S (2016) MicroRNAs and oncogenic transcriptional regulatory networks controlling metabolic reprogramming in cancers. Comput Struct Biotechnol J 14:223–233. doi:10.1016/j.csbj.2016.05.005
Jackstadt R, Hermeking H (2015) MicroRNAs as regulators and mediators of c-MYC function. Biochim Biophys Acta 1849(5):544–553. doi:10.1016/j.bbagrm.2014.04.003
Movafagh S, Crook S, Vo K (2015) Regulation of hypoxia-inducible factor-1a by reactive oxygen species: new developments in an old debate. J Cell Biochem 116(5):696–703. doi:10.1002/jcb.25074
Gauthier BR, Wollheim CB (2006) MicroRNAs: ‘ribo-regulators’ of glucose homeostasis. Nat Med 12(1):36–38. doi:10.1038/nm0106-36
Frost RJ, Olson EN (2011) Control of glucose homeostasis and insulin sensitivity by the Let-7 family of microRNAs. Proc Natl Acad Sci USA 108(52):21075–21080. doi:10.1073/pnas.1118922109
Zhu H, Shyh-Chang N, Segre AV, Shinoda G, Shah SP, Einhorn WS, Takeuchi A, Engreitz JM, Hagan JP, Kharas MG, Urbach A, Thornton JE, Triboulet R, Gregory RI, Consortium D, Investigators M, Altshuler D, Daley GQ (2011) The Lin28/let-7 axis regulates glucose metabolism. Cell 147 (1):81–94. doi:10.1016/j.cell.2011.08.033
Fu X, Dong B, Tian Y, Lefebvre P, Meng Z, Wang X, Pattou F, Han W, Wang X, Lou F, Jove R, Staels B, Moore DD, Huang W (2015) MicroRNA-26a regulates insulin sensitivity and metabolism of glucose and lipids. J Clin Invest 125(6):2497–2509. doi:10.1172/JCI75438
Jordan SD, Kruger M, Willmes DM, Redemann N, Wunderlich FT, Bronneke HS, Merkwirth C, Kashkar H, Olkkonen VM, Bottger T, Braun T, Seibler J, Bruning JC (2011) Obesity-induced overexpression of miRNA-143 inhibits insulin-stimulated AKT activation and impairs glucose metabolism. Nat Cell Biol 13(4):434–446. doi:10.1038/ncb2211
Dooley J, Garcia-Perez JE, Sreenivasan J, Schlenner SM, Vangoitsenhoven R, Papadopoulou AS, Tian L, Schonefeldt S, Serneels L, Deroose C, Staats KA, Van der Schueren B, De Strooper B, McGuinness OP, Mathieu C, Liston A (2016) The microRNA-29 family dictates the balance between homeostatic and pathological glucose handling in diabetes and obesity. Diabetes 65(1):53–61. doi:10.2337/db15-0770
Poy MN, Eliasson L, Krutzfeldt J, Kuwajima S, Ma X, Macdonald PE, Pfeffer S, Tuschl T, Rajewsky N, Rorsman P, Stoffel M (2004) A pancreatic islet-specific microRNA regulates insulin secretion. Nature 432(7014):226–230. doi:10.1038/nature03076
Poy MN, Hausser J, Trajkovski M, Braun M, Collins S, Rorsman P, Zavolan M, Stoffel M (2009) miR-375 maintains normal pancreatic alpha- and beta-cell mass. Proc Natl Acad Sci USA 106(14):5813–5818. doi:10.1073/pnas.0810550106
Sun X, Lin J, Zhang Y, Kang S, Belkin N, Wara AK, Icli B, Hamburg NM, Li D, Feinberg MW (2016) MicroRNA-181b improves glucose homeostasis and insulin sensitivity by regulating endothelial function in white adipose tissue. Circ Res. doi:10.1161/CIRCRESAHA.115.308166
Tripathi P, Kamarajan P, Somashekar BS, MacKinnon N, Chinnaiyan AM, Kapila YL, Rajendiran TM, Ramamoorthy A (2012) Delineating metabolic signatures of head and neck squamous cell carcinoma: phospholipase A2, a potential therapeutic target. Int J Biochem Cell Biol 44(11):1852–1861. doi:10.1016/j.biocel.2012.06.025
Wei S, Liu L, Zhang J, Bowers J, Gowda GA, Seeger H, Fehm T, Neubauer HJ, Vogel U, Clare SE, Raftery D (2013) Metabolomics approach for predicting response to neoadjuvant chemotherapy for breast cancer. Mol Oncol 7(3):297–307. doi:10.1016/j.molonc.2012.10.003
Zhang X, Xu L, Shen J, Cao B, Cheng T, Zhao T, Liu X, Zhang H (2013) Metabolic signatures of esophageal cancer: NMR-based metabolomics and UHPLC-based focused metabolomics of blood serum. Biochim Biophys Acta 1832(8):1207–1216. doi:10.1016/j.bbadis.2013.03.009
Wang Y, Zhang L, Chen WL, Wang JH, Li N, Li JM, Mi JQ, Zhang WN, Li Y, Wu SF, Jin J, Wang YG, Huang H, Chen Z, Chen SJ, Tang H (2013) Rapid diagnosis and prognosis of de novo acute myeloid leukemia by serum metabonomic analysis. J Proteome Res 12(10):4393–4401. doi:10.1021/pr400403p
Fong MY, Zhou W, Liu L, Alontaga AY, Chandra M, Ashby J, Chow A, O’Connor ST, Li S, Chin AR, Somlo G, Palomares M, Li Z, Tremblay JR, Tsuyada A, Sun G, Reid MA, Wu X, Swiderski P, Ren X, Shi Y, Kong M, Zhong W, Chen Y, Wang SE (2015) Breast-cancer-secreted miR-122 reprograms glucose metabolism in premetastatic niche to promote metastasis. Nat Cell Biol 17(2):183–194. doi:10.1038/ncb3094
Zhang A, Sun H, Wang P, Han Y, Wang X (2012) Modern analytical techniques in metabolomics analysis. Analyst 137(2):293–300. doi:10.1039/c1an15605e
Yonezawa K, Nishiumi S, Kitamoto-Matsuda J, Fujita T, Morimoto K, Yamashita D, Saito M, Otsuki N, Irino Y, Shinohara M, Yoshida M, Nibu K (2013) Serum and tissue metabolomics of head and neck cancer. Cancer Genomics Proteomics 10(5):233–238
Serizawa M, Kusuhara M, Zangiacomi V, Urakami K, Watanabe M, Takahashi T, Yamaguchi K, Yamamoto N, Koh Y (2014) Identification of metabolic signatures associated with erlotinib resistance of non-small cell lung cancer cells. Anticancer Res 34(6):2779–2787
Wen T, Gao L, Wen Z, Wu C, Tan CS, Toh WZ, Ong CN (2013) Exploratory investigation of plasma metabolomics in human lung adenocarcinoma. Mol Biosyst 9(9):2370–2378. doi:10.1039/c3mb70138g
Beyoglu D, Imbeaud S, Maurhofer O, Bioulac-Sage P, Zucman-Rossi J, Dufour JF, Idle JR (2013) Tissue metabolomics of hepatocellular carcinoma: tumor energy metabolism and the role of transcriptomic classification. Hepatology 58(1):229–238. doi:10.1002/hep.26350
Ramirez CM, Goedeke L, Rotllan N, Yoon JH, Cirera-Salinas D, Mattison JA, Suarez Y, de Cabo R, Gorospe M, Fernandez-Hernando C (2013) MicroRNA 33 regulates glucose metabolism. Mol Cell Biol 33(15):2891–2902. doi:10.1128/MCB.00016-13
Khan N, Islam MM, Mahmood S, Hossain GA, Chakraborty RK (2011) 18F-fluorodeoxyglucose uptake in tumor. Mymensingh Med J 20(2):332–342
Baldwin SA, Barros LF, Griffiths M (1995) Trafficking of glucose transporters—signals and mechanisms. Biosci Rep 15(6):419–426
Leung K (2004) [18F]Fluoro-2-deoxy-2-d-glucose. In: Molecular Imaging and Contrast Agent MICAD updated 12 Jan 2005
Pauwels EK, Ribeiro MJ, Stoot JH, McCready VR, Bourguignon M, Maziere B (1998) FDG accumulation and tumor biology. Nucl Med Biol 25(4):317–322
Speizer L, Haugland R, Kutchai H (1985) Asymmetric transport of a fluorescent glucose analogue by human erythrocytes. Biochim Biophys Acta 815(1):75–84
Yoshioka K, Saito M, Oh KB, Nemoto Y, Matsuoka H, Natsume M, Abe H (1996) Intracellular fate of 2-NBDG, a fluorescent probe for glucose uptake activity, in Escherichia coli cells. Biosci Biotechnol Biochem 60(11):1899–1901. doi:10.1271/bbb.60.1899
Di Chiro G, DeLaPaz RL, Brooks RA, Sokoloff L, Kornblith PL, Smith BH, Patronas NJ, Kufta CV, Kessler RM, Johnston GS, Manning RG, Wolf AP (1982) Glucose utilization of cerebral gliomas measured by [18F] fluorodeoxyglucose and positron emission tomography. Neurology 32(12):1323–1329
Mankoff DA, Eary JF, Link JM, Muzi M, Rajendran JG, Spence AM, Krohn KA (2007) Tumor-specific positron emission tomography imaging in patients: [18F] fluorodeoxyglucose and beyond. Clin Cancer Res 13(12):3460–3469. doi:10.1158/1078-0432.CCR-07-0074
Kang L, Xu X, Wang RF, Yan P, Zhang C (2013) Evaluation of the effect of microRNA on hepatocellular carcinoma proliferation and metastasis by FDG small-animal PET. J Nucl Med Meeting Abstr 54:1330
Zhang Y, Tang Y, Sun S, Wang Z, Wu W, Zhao X, Czajkowsky DM, Li Y, Tian J, Xu L, Wei W, Deng Y, Shi Q (2015) Single-cell codetection of metabolic activity, intracellular functional proteins, and genetic mutations from rare circulating tumor cells. Anal Chem 87(19):9761–9768. doi:10.1021/acs.analchem.5b01901
Agarwal P, Srivastava R, Srivastava AK, Ali S, Datta M (2013) miR-135a targets IRS2 and regulates insulin signaling and glucose uptake in the diabetic gastrocnemius skeletal muscle. Biochim Biophys Acta 1832(8):1294–1303. doi:10.1016/j.bbadis.2013.03.021
Etxeberria E, Gonzalez P, Tomlinson P, Pozueta-Romero J (2005) Existence of two parallel mechanisms for glucose uptake in heterotrophic plant cells. J Exp Bot 56(417):1905–1912. doi:10.1093/jxb/eri185
Sasaki A, Nagatomo K, Ono K, Yamamoto T, Otsuka Y, Teshima T, Yamada K (2016) Uptake of a fluorescent l-glucose derivative 2-NBDLG into three-dimensionally accumulating insulinoma cells in a phloretin-sensitive manner. Hum Cell 29(1):37–45. doi:10.1007/s13577-015-0125-3
Hu F, Chen Z, Zhang L, Shen Y, Wei L, Min W (2015) Vibrational imaging of glucose uptake activity in live cells and tissues by stimulated Raman scattering. Angew Chem Int Ed Engl 54(34):9821–9825. doi:10.1002/anie.201502543
Bar-Shalom R, Yefremov N, Guralnik L, Gaitini D, Frenkel A, Kuten A, Altman H, Keidar Z, Israel O (2003) Clinical performance of PET/CT in evaluation of cancer: additional value for diagnostic imaging and patient management. J Nucl Med 44(8):1200–1209
Mohammed A, Janakiram NB, Lightfoot S, Gali H, Vibhudutta A, Rao CV (2012) Early detection and prevention of pancreatic cancer: use of genetically engineered mouse models and advanced imaging technologies. Curr Med Chem 19(22):3701–3713
Wang Y, Black KC, Luehmann H, Li W, Zhang Y, Cai X, Wan D, Liu SY, Li M, Kim P, Li ZY, Wang LV, Liu Y, Xia Y (2013) Comparison study of gold nanohexapods, nanorods, and nanocages for photothermal cancer treatment. ACS Nano 7(3):2068–2077. doi:10.1021/nn304332s
Acknowledgements
We thank other members of the Liu laboratory for discussions and comments on the manuscript. This work was supported by Grants from the National Natural Science Foundation of China (31325008, 91640201, 91419307, and 31300656), Ministry of Science and Technology of China (2014CB943103 and 2014CB964802), Science and Technology Commission of Shanghai Municipality (13ZR1464300 and 16XD1404900), and Chinese Academy of Sciences (“Strategic Priority Research Program” Grant XDB19010202). We apologize to authors whose studies have not been cited in this manuscript due to space limitations.
Author contributions
All three authors were responsible for writing, reviewing, and revising the manuscript. Provenance: the authors were invited to submit this review manuscript.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflicts of interest.
Rights and permissions
About this article
Cite this article
Zhang, LF., Jiang, S. & Liu, MF. MicroRNA regulation and analytical methods in cancer cell metabolism. Cell. Mol. Life Sci. 74, 2929–2941 (2017). https://doi.org/10.1007/s00018-017-2508-y
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00018-017-2508-y