Abstract
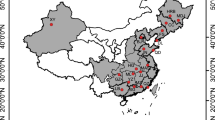
The camphor tree (Cinnamomum camphora) is an important natural resource in East Asia. From the early eighteenth to mid-twentieth century, C. camphora was widely cultivated to obtain camphor, a white crystalline substance used as a repellent, a component of medicine, and an ingredient in the production of smokeless gunpowder and celluloid. The vast utilization and cultivation of C. camphora have obscured its natural distribution, and the genetic structure of this species was likely affected by both natural factors and human activities in different areas and at different times. To estimate this process, we collected 817 samples from Japan, China, and Taiwan, including samples from trees estimated to be hundreds to thousands of years old. Population genetic analyses using 11 microsatellite markers detected the footprints of ancient genetic differentiation and recent human-mediated gene flow. The strong genetic differentiation between areas (Japan vs. China and Taiwan) and the decreased genetic diversity in Japan can be ascribed to the long-term geographical isolation during and after the glacial period. Within each area, the genetic composition of the cultivated or planted populations was often inconsistent with the surrounding natural and/or older specimens. This finding may be due to the regional transfer of plant materials with no regard to their genetic origin. We also detected rare but evident immigration from different areas; furthermore, these non-native genets seemed to locally hybridize with the native genets. We suggest that the intentional transfer and/or artificial hybridization between areas should be prohibited in order to preserve the genetic composition of C. camphora.




Similar content being viewed by others
References
Aoki K, Ueno S, Kamijo T, Setoguchi H, Murakami N, Kato M, Tsumura Y (2014) Genetic differentiation and genetic diversity of Castanopsis (Fagaceae), the dominant tree species in Japanese broadleaved evergreen forests, revealed by analysis of EST-associated microsatellites. PlosOne 9:e87429
Aoki K, Suzuki T, Hsu T-W, Murakami N (2004) Phylogeography of the component species of broad-leaved evergreen forests in Japan, based on chloroplast DNA variation. J Plant Res 117:77–94
Bagnoli F, Vendramin GG, Buonamici A, Doulis AG, Gonzálezmartínez SC, La Porta N, Magri D, Paddi P, Sebastiani F, Fineschi S (2009) Is Cupressus sempervirens native in Italy? An answer from genetic and palaeobotanical data. Mol Ecol 18:2276–2286
Baldoni L, Tosti N, Ricciolini C, Belaj A, Arcioni S, Pannelli G, Germana MA, Mulas M, Porceddu A (2006) Genetic structure of wild and cultivated olives in the Central Mediterranean Basin. Ann Bot 98:935–942
Barrett SCH, Husband BC (1990) The genetics of plant migration and colonization. In: Brown AHD, Clegg MT, Kahler AL, Weir BS (eds) Plant population genetics, breeding, and genetic resources. Sinauer Associations, Sunderland, pp 254–277
Brewer S, Cheddadi R, de Beaulieu JL, Reille M (2002) The spread of deciduous Quercus throughout Europe since the last glacial period. Forest Ecol Manag 156:27–48
Brownstein JM, Carpten JD, Smith JR (1996) Modulation of non-templated nucleotide addition by Taq DNA polymerase: primer modifications that facilitate genotyping. BioTechniques 20:1004–1010
Burban C, Petit RJ (2003) Phylogeography of maritime pine inferred with organelle markers having contrasted inheritance. Mol Ecol 12:1487–1495
Castillo-Cárdenas MF, Díaz-Gonzales F, Cerón-Souza I, Sanjur O, Toro-Perea N (2015) Jumping a geographic barrier: diversification of the mangrove species Pelliciera rhizophorae (Tetrameristaceae) across the Central American Isthmus. Tree Genet Genomes 11:822
Chen Z-H, Wu B, Li J-Y, Zhao J-G, Zhou X-Y, Zhang Y-K (2004) Germination of the seeds and growth of the seedlings of Cinnamomum camphora (L.) Presl. Plant Spec Biol 19:55–58
Cornuet JM, Santos F, Beaumont MA, Robert CP, Marin JM, Balding DJ, Guillemaud T, Estoup A (2008) Inferring population history with DIYABC: a user-friendly approach to approximate Bayesian computations. Bioinformatics 24:2713–2719
Cornuet JM, Ravigné V, Estoup A (2010) Inference on population history and model checking using DNA sequence and microsatellite data with the software DIYABC (v1.0). BMC Bioinformatics 11:401
Cornuet JM, Pudlo P, Veyssier J, Dehne-Garcia A, Gautier M, Leblois R, Marin J-M, Estoup A (2014) DIYABC v2.0: a software to make approximate Bayesian computation inferences about population history using single nucleotide polymorphism, DNA sequence and microsatellite data. Bioinformatics 30:1187–1189
Crawford GW (2011) Advances in understanding early agriculture in Japan. Curr Anthropol 52:S331–S345
Dakin EE, Avise JC (2004) Microsatellite null alleles in parentage analysis. Heredity 93:504–509
Earl DA, vonHoldt BM (2012) STRUCTURE HARVESTER: a website and program for visualizing STRUCTURE output and implementing the Evanno method. Conserv Genet Resour 4:359–361
Edmands S (2007) Between a rock and a hard place: evaluating the relative risks of inbreeding and outbreeding for conservation and management. Mol Ecol 16:463–475
Environment Agency of Japan (1991) Big trees in Japan. Printing Bureau of the Ministry of Finance. Tokyo, Japan
Evanno G, Regnaut S, Goudet J (2005) Detecting the number of clusters of individuals using the software STRUCTURE: a simulation study. Mol Ecol 14:2611–2620
Fang Z (1997) Effect of different slope positions on the growth and development of Cinnamomum camphora. Journal of Fujian Forestry Science and Technology 24:20–22 (In Chinese with English abstract)
Firth DJ (1981) Camphor laurel (Cinnamomum camphora)—a new weed in north-eastern New South Wales. Aust Weeds 1:26–28
Firth DJ (1980) History of introduction of Cinnamomum camphora (camphor laurel tree) to Australia. J Aust Inst Agric Sci 46:244–245
Frankham R, Ballou J, Eldridge MDB, Lacy RC, Ralls K, Dudash MR, Fenster CB (2011) Predicting the probability of outbreeding depression. Conserv Biol 25:465–475
Goudet J (1995) FSTAT (version 1.2): a computer program to calculate F-statistics. JHered 86:485–486
Hammer MF, Karafet TM, Park H, Omoto K, Harihara S, Stoneking M, Horai S (2006) Dual origins of the Japanese: common ground for hunter-gatherer and farmer Y chromosomes. J Hum Genet 51:47–58
Hampe A, Petit RJ (2007) Ever deeper phylogeographies: trees retain the genetic imprint of Tertiary plate tectonics. Mol Ecol 16:5113–5114
Haq BU, Hardenbol J, Vail PR (1987) Chronology of fluctuating sea levels since the Triassic. Science 235:1156–1167
Harrison SP, Yu G, Takahara H, Prentice IC (2001) Palaeovegetatio (Communications arising): Diversity of temperate plants in east Asia. Nature 413:129–130
Hattori T (1985) Synecological study on the lucidophyllous forest of Castanopsis-Persea type in Japan proper. Bull Kobe Geobot Soc 1:1–98
Hedrick PW (2005) A standardized genetic differentiation measure. Evolution 59:1633–1638
Iwaizumi MG, Tsuda Y, Ohtani M, Tsumura Y, Takahashi M (2013) Recent distribution changes affect geographic clines in genetic diversity and structure of Pinus densiflora natural populations in Japan. Forest Ecol Manag 304:407–416
Iwatsuki K, Boufford DE, Ohba H (2006) Flora of Japan IIa. Kodansha, Tokyo, Japan
Japan Forest Technical Association (1964) Illustrated important forest trees of Japan. Chikyu Shuppan, Tokyo, Japan
Kalinowski ST, Taper ML, Marshall TC (2007) Revising how the computer program CERVUS accommodates genotyping error increases success in paternity assignment. Mol Ecol 16:1099–1106
Kameyama Y (2012) Development of microsatellite markers for Cinnamomum camphora (Lauraceae). AmJ Bot 99:e1–e3
Kanowski J, Catterall CP, Neilan W (2008) Potential value of weedy regrowth for rainforest restoration. Ecol Manage Rest 9:88–99
Kimura M (2000) Paleography of the Ryukyu Islands. Tropics 10:5–24
Kitamura K, Matsui T, Kobayashi M, Saitou H, Namikawa K, Tsuda Y (2015) Decline in gene diversity and strong genetic drift in the northward-expanding marginal populations of Fagus crenata. Tree Genet Genomes 11:36
Kondo T, Crisp MD, Linde C, Bowman DMJS, Kawamura K, Kaneko S, Isagi Y (2012) Not an ancient relic: the endemic Livistona palms of arid central Australia could have been introduced by humans. Proc R Soc B 279:2652–2661
König AO, Ziegenhagen B, van Dam BC, Csaikl UM, Coart E, Degen B, Burg K, de Vries SMG, Petit RJ (2002) Chloroplast DNA variation of oaks in western Central Europe and genetic consequences of human influences. Forest Ecol Manag 156:147–166
Koskela J, Vinceti B, Dvorak W, Bush D, Dawson IK, Loo J, Kjaer ED, Navarro C, Padolina C, Bordács S, Jamnadass R, Graudal L, Ramamonjisoa L (2014) Utilization and transfer of forest genetic resources: a global review. Forest Ecol Manag 333:22–34
Koyama S (1992) Prehistoric Japanese populations: a subsistence-demographic approach. In: Hanihara K (ed) Japanese as a member of the Asian and Pacific populations. International Research Center for Japanese Studies, Kyoto, pp 187–197
Lambeck K, Chappell J (2001) Sea-level change through the last glacial cycle. Science 292:679–686
Lang P, Dane F, Kubisiak TL, Huang H (2007) Molecular evidence for an Asian origin and a unique westward migration of species in the genus Castanea via Europe to North America. Mol Phylogenet Evol 43:49–59
Leberg PL (2002) Estimating allelic richness: effects of sample size and bottlenecks. Mol Ecol 11:2445–2449
Ledig FT (1992) Human impacts on genetic diversity in forest ecosystems. Oikos 63:87–108
Lefèvre F (2004) Human impacts on forest genetic resources in the temperate zone: an updated review. Forest Ecol Manag 197:257–271
Lendvay B, Höhn M, Brodbeck S, Mîndrescu M, Gugerli F (2014) Genetic structure in Pinus cembra from the Carpathian Mountains inferred from nuclear and chloroplast microsatellites confirms post-glacial range contraction and identifies introduced individuals. Tree Genet Genomes 10:1419–1433
Magri D, Vendramin GG, Comps B, Dupanloup I, Geburek T, Gömöry D, Latałowa M, Litt T, Paule L, Roure JM, Tantau I, van der Knaap WO, Petit RJ, de Beaulieu J-L (2006) A new scenario for the Quaternary history of European beech populations: palaeobotanical evidence and genetic consequences. New Phytol 171:199–221
Magri D, Fineschi S, Bellarosa R, Buonamici A, Sebastiani F, Schirone B, Simeone MC, Vendramin GG (2007) The distribution of Quercus suber chloroplast haplotypes matches the palaeogeographical history of the western Mediterranean. Mol Ecol 16:5259–5266
Marshall TC, Slate J, Kruuk LEB, Pemberton JM (1998) Statistical confidence for likelihood-based paternity inference in natural populations. Mol Ecol 7:639–655
Martín MA, Mattioni C, Molina JR, Alvarez JB, Cherubini M, Herrera MA, Villani F, Martín LM (2012) Landscape genetic structure of chestnut (Castanea sativa Mill.) in Spain. Tree Genet Genomes 8:127–136
Matsuoka K, Miyoshi N (1998) Transition of an evergreen broad-leaved forest after the last glacial maximum. In: Yasuda Y, Miyoshi N (eds) Illustrated vegetation history of the Japanese archipelago. Asakura Publishing, Tokyo, pp 224–236 (In Japanese)
Mattioni C, Martin MA, Pollegioni P, Cherubini M, Villani F (2013) Microsatellite markers reveal a strong geographical structure in European populations of Castanea sativa (Fagaceae): evidence for multiple glacial refugia. Am JBot 100:951–961
Mattioni C, Cherubini M, Micheli E, Villani F, Bucci G (2008) Role of domestication in shaping Castanea sativa genetic variation in Europe. Tree Genet Genomes 4:563–574
McKay JK, Christian CE, Harrison S, Rice KJ (2005) “How local is local?”—a review of practical and conceptual issues in the genetics of restoration. Restor Ecol 13:432–440
Nakagawa S (1985) Growth of Cinnamomum camphora of Kajiya at Yugawara City, West Kanagawa. Bulletin of the Kanagawa Prefecture Forest Experiment Station 11:11–18 (In Japanese with English abstract)
Nakanishi H (1996) Plant species with northbound distribution in western Kyushu, Japan: definition, composition and origin. Acta Phytotax Geobot 47:113–124 (in Japanese with English abstract)
Neophytou C, Michiels H-G (2013) Upper Rhine Valley: a migration crossroads of middle European oaks. Forest Ecol Manag 304:89–98
Neophytou C, Gärtner SM, Vargas-Gaete R, Michiels H-G (2015) Genetic variation of Central European oaks: shaped by evolutionary factors and human intervention? Tree Genet Genomes 11:79
Nishio S, Iketani H, Fujii H, Yamamoto T, Terakami S, Takada N, Saito T (2014) Use of population structure and parentage analyses to elucidate the spread of native cultivars of Japanese chestnut. Tree Genet Genomes 10:1171–1180
Peakall R, Smouse PE (2006) GENEALEX 6: genetic analysis in Excel. Population genetic software for teaching and research. Mol Ecol Notes 6:288–295
Peakall R, Smouse PE, Huff DR (1995) Evolutionary implications of allozyme and RAPD variation in diploid populations of dioecious buffalograss Buchloë dactyloides. Mol Ecol 4:135–148
Petit RJ, Csaikl UM, Bordács S, Burg K, Coart E, Cottrell J, van Dam B, Deans JD, Dumolin-Lapègue S, Fineschi S, Finkeldey R, Gillies A, Glaz I, Goicoechea PG, Jensen JS, König AO, Lowe AJ, Madsen SF, Mátyás G, Munro RC, Olalde M, Pemonge M-H, Popescu F, Slade D, Tabbener H, Taurchini D, de Vries SGM, Ziegenhagen B, Kremer A (2002) Chloroplast DNA variation in European white oaks: phylogeography and patterns of diversity based on data from over 2600 populations. Forest Ecol Manag 156:5–26
Petit RJ (2004) Biological invasions at the gene level. Divers Distrib 10:159–165
Petit RJ, Mousadik AE, Pons O (1998) Identifying populations for conservation on the basis of genetic markers. Conserv Biol 12:844–855
Pollegioni P, Woeste K, Olimpieri I, Marandola D, Cannata F, Malvolti ME (2011) Long-term human impacts on genetic structure of Italian walnut inferred by SSR markers. Tree Genet Genomes 7:707–723
Pollegioni P, Woeste KE, Chiocchini F, Olimpieri I, Tortolano V, Clark J, Hemery GE, Mapelli S, Malvolti ME (2014) Landscape genetics of Persian walnut (Juglans regia L.) across its Asian range. Tree Genet Genomes 10:1027–1043
Pritchard JK, Stephens M, Donnelly P (2000) Inference of population structure using multilocus genotype data. Genetics 155:945–959
Qiu Y-X, Fu C-X, Comes HP (2011) Plant molecular phylogeography in China and adjacent regions: tracing the genetic imprints of Quaternary climate and environmental change in the world’s most diverse temperate flora. Mol Phylogenet Evol 59:225–244
Quintana J, Contreras A, Merino I, Vinuesa A, Orozco G, Ovalle F, Gomez L (2015) Genetic characterization of chestnut (Castanea sativa Mill.) orchards and traditional nut varieties in El Bierzo, a glacial refuge and major cultivation site in northwestern Spain. Tree Genet Genomes 11:826
Sakaguchi S, Qiu Y-X, Liu Y-H, Qi X-S, Kim S-H, Han J, Takeuchi Y, Worth JRP, Yamasaki M, Sakurai S, Isagi Y (2012) Climate oscillation during the Quaternary associated with landscape heterogeneity promoted allopatric lineage divergence of a temperate tree Kalopanax septemlobus (Araliaceae) in East Asia. Mol Ecol 21:3823–3838
Stewart CN Jr, Via LE (1993) A rapid CTAB DNA isolation technique useful for RAPD fingerprinting and other PCR applications. BioTechniques 14:748–750
Thomas E, Jalonen R, Loo J, Boshier D, Gallo L, Cavers S, Bordács S, Smith P, Bozzano M (2014) Genetic considerations in ecosystem restoration using native tree species. Forest EcolManag 333:66–75
Tsukada M (1974) Paleoecology. II. Synthesis. Kyoritsu Shuppan, Tokyo (In Japanese)
Yano K, Yano K (2010) Camphora tree. Hosei Univeristy Press, Tokyo, Japan (In Japanese)
Acknowledgments
We thank Kojiro Suzuki at the Tokyo University of Agriculture, Chao Chien-Ti at National Chung Hsing University, and Bangping Cai at the Siamen Botanical Garden for their substantial assistance with field sampling. This work was supported by JSPS KAKENHI Grant Number JP24710276.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Data archiving statement
All data are available at figshare: https://doi.org/10.6084/m9.figshare.4664002.v1.
Additional information
Communicated by Y. Tsumura
Rights and permissions
About this article
Cite this article
Kameyama, Y., Furumichi, J., Li, J. et al. Natural genetic differentiation and human-mediated gene flow: the spatiotemporal tendency observed in a long-lived Cinnamomum camphora (Lauraceae) tree. Tree Genetics & Genomes 13, 38 (2017). https://doi.org/10.1007/s11295-017-1119-y
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11295-017-1119-y